

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141111.3 + phase: 0 /pseudo
(64 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
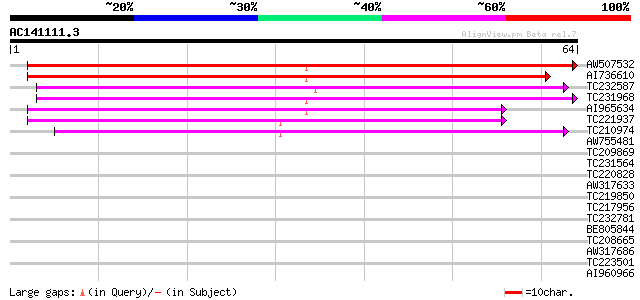
Score E
Sequences producing significant alignments: (bits) Value
AW507532 48 9e-07
AI736610 44 2e-05
TC232587 similar to UP|Q9LME0 (Q9LME0) F14D16.2, partial (17%) 41 1e-04
TC231968 40 3e-04
AI965634 39 4e-04
TC221937 similar to UP|Q8VZE4 (Q8VZE4) AT4g01570/T15B16_21, part... 39 4e-04
TC210974 similar to UP|Q6K9W7 (Q6K9W7) Pentatricopeptide (PPR) r... 39 7e-04
AW755481 similar to GP|2244996|emb salt-inducible protein homolo... 37 0.002
TC209869 similar to UP|Q9LG03 (Q9LG03) F20N2.6, partial (33%) 36 0.004
TC231564 similar to GB|AAP31962.1|30387593|BT006618 At3g06430 {A... 35 0.010
TC220828 similar to UP|Q9FLD8 (Q9FLD8) Gb|AAD55291.1, partial (37%) 33 0.022
AW317633 similar to PIR|T46163|T46 nodulin / glutamate-ammonia l... 33 0.038
TC219850 weakly similar to UP|Q9FKI5 (Q9FKI5) Emb|CAB69839.1, pa... 33 0.038
TC217956 similar to UP|O65793 (O65793) Leaf protein, partial (30%) 32 0.049
TC232781 similar to UP|Q9ZU27 (Q9ZU27) F5F19.2 protein, partial ... 32 0.084
BE805844 31 0.14
TC208665 weakly similar to UP|O65567 (O65567) Puative protein, p... 30 0.19
AW317686 30 0.19
TC223501 29 0.55
AI960966 similar to PIR|G85360|G8 puative protein [imported] - A... 28 0.71
>AW507532
Length = 396
Score = 48.1 bits (113), Expect = 9e-07
Identities = 24/63 (38%), Positives = 40/63 (63%), Gaps = 1/63 (1%)
Frame = +3
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+V YNT+I GY + +D AE +F+ + GR I P+ S+ +M++G+ AG + A +E
Sbjct: 153 VVTYNTLINGYFRFKKVDEAEKLFVEMKGRDIVPNVISFTTMLKGYVAAGRIDDALKVFE 332
Query: 62 ELK 64
E+K
Sbjct: 333 EMK 341
>AI736610
Length = 411
Score = 43.5 bits (101), Expect = 2e-05
Identities = 22/60 (36%), Positives = 37/60 (61%), Gaps = 1/60 (1%)
Frame = +2
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+V YNT+I GY + +D AE +F+ + GR I P+ S+ +M++G+ AG + A +E
Sbjct: 230 VVTYNTLINGYFRFKKVDEAEKLFVEMKGRDIVPNVISFTTMLKGYVAAGRIDDALKVFE 409
>TC232587 similar to UP|Q9LME0 (Q9LME0) F14D16.2, partial (17%)
Length = 516
Score = 41.2 bits (95), Expect = 1e-04
Identities = 20/61 (32%), Positives = 35/61 (56%), Gaps = 1/61 (1%)
Frame = +3
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTLGGRI-EPDETSYRSMIEGWGRAGNYEKARWYYEE 62
V Y+ ++ G ++ AE VF+ + + PDE Y +++ WG+AGN EKA +Y+
Sbjct: 279 VTYSIVMEALGHCGYLEEAESVFVEMQQKNWVPDEPVYGLLVDLWGKAGNVEKASEWYQA 458
Query: 63 L 63
+
Sbjct: 459 M 461
Score = 26.9 bits (58), Expect = 2.1
Identities = 11/31 (35%), Positives = 17/31 (54%)
Frame = +3
Query: 33 IEPDETSYRSMIEGWGRAGNYEKARWYYEEL 63
+ PD +Y +I G+AGN A W + E+
Sbjct: 54 LSPDTFTYSVIINCLGKAGNLAAAHWLFCEM 146
>TC231968
Length = 610
Score = 39.7 bits (91), Expect = 3e-04
Identities = 23/62 (37%), Positives = 35/62 (56%), Gaps = 1/62 (1%)
Frame = +1
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
V YNT I G KA D F + + +EPD +Y ++IE +GR+GN E++ + E
Sbjct: 73 VSYNTXINGLRKAGRXDMCFVYFKEMTEKGVEPDLLTYTAIIEIFGRSGNVEESLKCFRE 252
Query: 63 LK 64
+K
Sbjct: 253 MK 258
Score = 26.2 bits (56), Expect = 3.5
Identities = 18/63 (28%), Positives = 31/63 (48%), Gaps = 2/63 (3%)
Frame = +1
Query: 3 IVVYNTMITGYGKASNMDGAEGVF--LTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYY 60
++ Y +I +G++ N++ + F + L G + P YRS+I + G E A
Sbjct: 175 LLTYTAIIEIFGRSGNVEESLKCFREMKLKG-VLPSIYIYRSLIHNLNKTGKVELATELL 351
Query: 61 EEL 63
EEL
Sbjct: 352 EEL 360
>AI965634
Length = 415
Score = 39.3 bits (90), Expect = 4e-04
Identities = 20/55 (36%), Positives = 31/55 (56%), Gaps = 1/55 (1%)
Frame = +2
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKA 56
+V YNT+I GY K N+ GA + + R I P + +Y +I+ + R + EKA
Sbjct: 143 LVTYNTLIAGYSKVENLAGALDLVKEMEERCIAPSKVTYTILIDAFARLNHTEKA 307
>TC221937 similar to UP|Q8VZE4 (Q8VZE4) AT4g01570/T15B16_21, partial (22%)
Length = 801
Score = 39.3 bits (90), Expect = 4e-04
Identities = 21/55 (38%), Positives = 31/55 (56%), Gaps = 1/55 (1%)
Frame = +2
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKA 56
IV+YNT+I GKAS +D +F + I PD +Y ++IE +AG + A
Sbjct: 275 IVMYNTLINALGKASRIDEVNKLFEQMRSSGINPDVVTYNTLIEVHSKAGRLKDA 439
>TC210974 similar to UP|Q6K9W7 (Q6K9W7) Pentatricopeptide (PPR)
repeat-containing protein-like, partial (3%)
Length = 637
Score = 38.5 bits (88), Expect = 7e-04
Identities = 20/59 (33%), Positives = 33/59 (55%), Gaps = 1/59 (1%)
Frame = +2
Query: 6 YNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKARWYYEEL 63
Y TM++ Y A +M+GAE F L EP+ +Y ++I+G+ + + E YEE+
Sbjct: 155 YTTMLSAYINADDMEGAEKFFKRLIQDGFEPNVVTYGTLIKGYAKINDLEMVMKKYEEM 331
>AW755481 similar to GP|2244996|emb salt-inducible protein homolog
{Arabidopsis thaliana}, partial (17%)
Length = 435
Score = 37.0 bits (84), Expect = 0.002
Identities = 22/64 (34%), Positives = 36/64 (55%), Gaps = 3/64 (4%)
Frame = +2
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTLGGRIEP---DETSYRSMIEGWGRAGNYEKARWYY 60
V Y+ MI YG+A N+D A ++ R E D ++ ++I+ +G AGNY+ Y
Sbjct: 68 VTYSAMIDAYGRAGNIDMALRLYDR--ARTEKWRLDTVTFSTLIKMYGLAGNYDGCLNVY 241
Query: 61 EELK 64
+E+K
Sbjct: 242 QEMK 253
Score = 33.9 bits (76), Expect = 0.017
Identities = 13/31 (41%), Positives = 21/31 (66%)
Frame = +2
Query: 34 EPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
EPD+ +Y +MI+ +GRAGN + A Y+ +
Sbjct: 56 EPDDVTYSAMIDAYGRAGNIDMALRLYDRAR 148
Score = 29.3 bits (64), Expect = 0.42
Identities = 14/54 (25%), Positives = 29/54 (52%), Gaps = 1/54 (1%)
Frame = +2
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEK 55
+V+YNT++ G+A A+ ++ + P+ +Y S++ +GR G Y +
Sbjct: 275 MVIYNTLLDAMGRAKRPWQAKSIYTEMTNNGFSPNWVTYASLLRAYGR-GRYSE 433
>TC209869 similar to UP|Q9LG03 (Q9LG03) F20N2.6, partial (33%)
Length = 1085
Score = 35.8 bits (81), Expect = 0.004
Identities = 19/62 (30%), Positives = 32/62 (50%), Gaps = 1/62 (1%)
Frame = +1
Query: 3 IVVYNTMITGYGKASNMDGAEGVF-LTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
++ + T+I G +A N+D + F + PD +Y MI G+ AG EKA Y+
Sbjct: 181 VLHFTTLIDGLSRAGNLDACKYFFDEMIKNECRPDVVAYTVMITGYVVAGEIEKALEMYQ 360
Query: 62 EL 63
++
Sbjct: 361 DM 366
Score = 30.0 bits (66), Expect = 0.25
Identities = 10/31 (32%), Positives = 21/31 (67%)
Frame = +1
Query: 33 IEPDETSYRSMIEGWGRAGNYEKARWYYEEL 63
IEP + ++I+G RAGN + +++++E+
Sbjct: 169 IEPTVLHFTTLIDGLSRAGNLDACKYFFDEM 261
>TC231564 similar to GB|AAP31962.1|30387593|BT006618 At3g06430 {Arabidopsis
thaliana;} , partial (40%)
Length = 797
Score = 34.7 bits (78), Expect = 0.010
Identities = 20/63 (31%), Positives = 35/63 (54%), Gaps = 1/63 (1%)
Frame = +1
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
I YN +++ KA ++ E VF + + +PD+T+Y MIE + + G +K +Y E
Sbjct: 391 ITFYNAVLSACAKAEDLMEMERVFKRMKDSQCQPDDTTYTIMIEAYRKEGMNDKI-YYLE 567
Query: 62 ELK 64
+ K
Sbjct: 568 QEK 576
>TC220828 similar to UP|Q9FLD8 (Q9FLD8) Gb|AAD55291.1, partial (37%)
Length = 872
Score = 33.5 bits (75), Expect = 0.022
Identities = 21/62 (33%), Positives = 33/62 (52%), Gaps = 1/62 (1%)
Frame = +3
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
+ Y+T+I+ + KA +D A +F L + DE Y++MI + RAG A+ E
Sbjct: 69 ITYSTIISIWEKAGKLDRAAILFQKLRSSGVRIDEVLYQTMIVAYERAGLVAHAKRLLHE 248
Query: 63 LK 64
LK
Sbjct: 249 LK 254
Score = 27.7 bits (60), Expect = 1.2
Identities = 18/59 (30%), Positives = 27/59 (45%)
Frame = +3
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
V+Y TMI Y +A + A+ + L PD + I RAG E+A W + +
Sbjct: 174 VLYQTMIVAYERAGLVAHAKRLLHELK---RPDNIPRDTAIGILARAGRIEEATWVFRQ 341
>AW317633 similar to PIR|T46163|T46 nodulin / glutamate-ammonia ligase-like
protein - Arabidopsis thaliana, partial (26%)
Length = 484
Score = 32.7 bits (73), Expect = 0.038
Identities = 16/54 (29%), Positives = 31/54 (56%), Gaps = 1/54 (1%)
Frame = +1
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGG-RIEPDETSYRSMIEGWGRAGNYEK 55
IV YNT+I +GKA ++ + FL + ++P+ +Y S++ + + G +K
Sbjct: 28 IVTYNTVIEVFGKAGEIEKMDQHFLKMKHLGVKPNSITYCSLVSAYSKVGCIDK 189
>TC219850 weakly similar to UP|Q9FKI5 (Q9FKI5) Emb|CAB69839.1, partial (53%)
Length = 1153
Score = 32.7 bits (73), Expect = 0.038
Identities = 18/64 (28%), Positives = 33/64 (51%), Gaps = 1/64 (1%)
Frame = +2
Query: 2 CIVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYY 60
C+ Y+TMI YG+ + A + + R +P+ Y S+I+ GR N ++ +
Sbjct: 440 CVYAYSTMIVMYGRTGRVRSAMKLVAKMKERGCKPNVWIYNSLIDMHGRDKNLKQLEKLW 619
Query: 61 EELK 64
+E+K
Sbjct: 620 KEMK 631
>TC217956 similar to UP|O65793 (O65793) Leaf protein, partial (30%)
Length = 1224
Score = 32.3 bits (72), Expect = 0.049
Identities = 17/55 (30%), Positives = 30/55 (53%), Gaps = 1/55 (1%)
Frame = +1
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKA 56
+V ++T++ + A M+ E +F + IEPD +Y + +G+ RAG KA
Sbjct: 313 VVTFSTIMNAWSSAGLMENCEEIFNDMVKAGIEPDIHAYSILAKGYVRAGQPRKA 477
>TC232781 similar to UP|Q9ZU27 (Q9ZU27) F5F19.2 protein, partial (3%)
Length = 1118
Score = 31.6 bits (70), Expect = 0.084
Identities = 18/62 (29%), Positives = 32/62 (51%), Gaps = 1/62 (1%)
Frame = +2
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYE 61
I YN +I YG+A ++ E +F L + ++PD ++ S I + + Y K +E
Sbjct: 548 ISTYNILINRYGQAGFIERMEDLFQLLPSKGLKPDVVTWTSRIGAYSKKKLYLKCLEIFE 727
Query: 62 EL 63
E+
Sbjct: 728 EM 733
>BE805844
Length = 429
Score = 30.8 bits (68), Expect = 0.14
Identities = 17/60 (28%), Positives = 32/60 (53%), Gaps = 1/60 (1%)
Frame = +3
Query: 6 YNTMITGYGKASNMDGAEGVF-LTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
YN MI +GK + +D A F L I PD ++ ++I+ ++G ++ A + E++
Sbjct: 111 YNVMIDTFGKYNCLDHAMATFERMLSEGIPPDIVTWNTLIDCHCKSGRHDMAEELFSEMQ 290
>TC208665 weakly similar to UP|O65567 (O65567) Puative protein, partial (36%)
Length = 1204
Score = 30.4 bits (67), Expect = 0.19
Identities = 15/63 (23%), Positives = 29/63 (45%), Gaps = 1/63 (1%)
Frame = +1
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLG-GRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
++ YNT+I YGK + + + +Y SM++ +G+ G E R +
Sbjct: 805 VITYNTIIAAYGKNKDFNNMSSTVQKMEFDGFSVSLEAYNSMLDAYGKDGQMETFRSVLQ 984
Query: 62 ELK 64
++K
Sbjct: 985 KMK 993
>AW317686
Length = 447
Score = 30.4 bits (67), Expect = 0.19
Identities = 13/50 (26%), Positives = 29/50 (58%), Gaps = 1/50 (2%)
Frame = +1
Query: 3 IVVYNTMITGYGKASNMDGAEGVF-LTLGGRIEPDETSYRSMIEGWGRAG 51
+V +NT+ITG+ + + + + VF + ++EP+ ++ ++ G AG
Sbjct: 34 VVTWNTLITGFAQNGHGEESLAVFRRMIEAKVEPNHVTFLGVLSGCNHAG 183
>TC223501
Length = 790
Score = 28.9 bits (63), Expect = 0.55
Identities = 17/60 (28%), Positives = 31/60 (51%), Gaps = 1/60 (1%)
Frame = +2
Query: 6 YNTMITGYGKASNMDGAEGVFLTLGG-RIEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
+N++I + KA ++ A F + + EP++ +Y SMI G+ A Y + E+K
Sbjct: 56 WNSIIHAFCKAGRLEDARRTFRRMMFLQFEPNDQTYLSMINGYVLAEKYFLVLMLWNEVK 235
>AI960966 similar to PIR|G85360|G8 puative protein [imported] - Arabidopsis
thaliana, partial (13%)
Length = 440
Score = 28.5 bits (62), Expect = 0.71
Identities = 23/68 (33%), Positives = 31/68 (44%), Gaps = 5/68 (7%)
Frame = +1
Query: 2 CIVVYNTMITG-----YGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKA 56
C VV N G Y K N++ AE F + G E++Y SMI + R YEKA
Sbjct: 61 CGVVPNVATIGMLMGLYRKGWNLEEAEFAFSRMRGFRIVCESAYSSMITIYTRLRLYEKA 240
Query: 57 RWYYEELK 64
E ++
Sbjct: 241 EGVIELMR 264
Score = 26.2 bits (56), Expect = 3.5
Identities = 11/14 (78%), Positives = 12/14 (85%)
Frame = +1
Query: 3 IVVYNTMITGYGKA 16
IV +NTMITG GKA
Sbjct: 391 IVAFNTMITGXGKA 432
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.137 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,610,136
Number of Sequences: 63676
Number of extensions: 21148
Number of successful extensions: 166
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 155
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 164
length of query: 64
length of database: 12,639,632
effective HSP length: 40
effective length of query: 24
effective length of database: 10,092,592
effective search space: 242222208
effective search space used: 242222208
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 52 (24.6 bits)
Medicago: description of AC141111.3


