

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140916.4 - phase: 0
(250 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
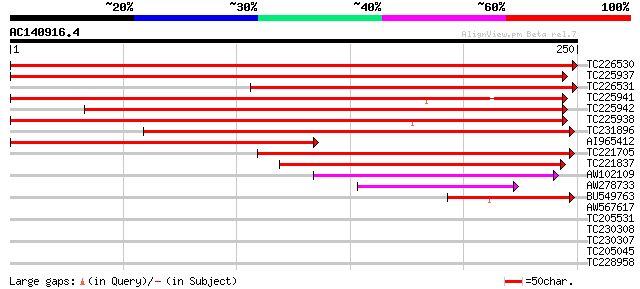
Score E
Sequences producing significant alignments: (bits) Value
TC226530 similar to UP|Q8VYX0 (Q8VYX0) IAA-amino acid conjugate ... 442 e-125
TC225937 similar to UP|O81642 (O81642) IAA-Ala hydrolase (IAR3) ... 256 9e-69
TC226531 weakly similar to UP|O81702 (O81702) Gr1-protein, parti... 252 1e-67
TC225941 similar to UP|O81642 (O81642) IAA-Ala hydrolase (IAR3) ... 226 1e-59
TC225942 similar to UP|O81642 (O81642) IAA-Ala hydrolase (IAR3) ... 217 5e-57
TC225938 similar to UP|Q9SWX9 (Q9SWX9) Auxin conjugate hydrolase... 213 5e-56
TC231896 similar to UP|Q84XG9 (Q84XG9) IAA-amino acid hydrolase,... 197 3e-51
AI965412 162 1e-40
TC221705 weakly similar to UP|O81641 (O81641) IAA-amino acid hyd... 140 4e-34
TC221837 weakly similar to UP|O65840 (O65840) IAA amidohydrolase... 126 1e-29
AW102109 87 7e-18
AW278733 weakly similar to PIR|F96556|F965 IAA-Ala hydrolase (IA... 57 1e-08
BU549763 similar to GP|27948556|gb| IAA-amino acid hydrolase {Or... 52 3e-07
AW567617 31 0.61
TC205531 weakly similar to UP|O04559 (O04559) T7N9.16, partial (... 28 3.0
TC230308 28 3.0
TC230307 28 3.9
TC205045 similar to UP|P93166 (P93166) SCOF-1, partial (45%) 27 6.7
TC228958 similar to UP|MCCB_ARATH (Q9LDD8) Methylcrotonyl-CoA ca... 27 8.8
>TC226530 similar to UP|Q8VYX0 (Q8VYX0) IAA-amino acid conjugate
hydrolase-like protein (Fragment), partial (79%)
Length = 1480
Score = 442 bits (1137), Expect = e-125
Identities = 216/250 (86%), Positives = 233/250 (92%)
Frame = +1
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
M+QDGALEDVEAIFA HVSHEHPTG+IGSRPGPLLAGCGFFRAVISGK+ AANP S D
Sbjct: 727 MMQDGALEDVEAIFAAHVSHEHPTGIIGSRPGPLLAGCGFFRAVISGKKGLAANPHRSVD 906
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
PVLAASAAVIS+QGIVSRE+NPLDSQVVSVTSFNGGN+ DMIPDSVV+ GTFRAFSNTSF
Sbjct: 907 PVLAASAAVISLQGIVSREANPLDSQVVSVTSFNGGNNLDMIPDSVVLLGTFRAFSNTSF 1086
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
YQLLERIEQVIV+QASVY C AEVDFFEKEYTIYPPTVND++MYEHVKKVSIDLLG KNF
Sbjct: 1087YQLLERIEQVIVEQASVYRCLAEVDFFEKEYTIYPPTVNDNRMYEHVKKVSIDLLGHKNF 1266
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
RVVPPMMGAED+SFYS+V+PS FFYIG+RNETLGSTHTGHSP+F IDED LPIGAA HA+
Sbjct: 1267RVVPPMMGAEDFSFYSEVVPSGFFYIGVRNETLGSTHTGHSPYFMIDEDVLPIGAAAHAS 1446
Query: 241 IAERYLNEHG 250
A+RYL EHG
Sbjct: 1447YAKRYLIEHG 1476
>TC225937 similar to UP|O81642 (O81642) IAA-Ala hydrolase (IAR3)
(At1g51760/F19C24_4), partial (89%)
Length = 1569
Score = 256 bits (653), Expect = 9e-69
Identities = 123/246 (50%), Positives = 174/246 (70%)
Frame = +1
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
++ G LE++ AIF +H++ +P G + SR GP+ AG GFF A I+G+ AA P++S D
Sbjct: 604 ILDAGVLENISAIFGLHIAPTYPIGEVASRSGPIFAGSGFFEATINGRGGHAAIPQHSID 783
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
P+LAAS ++S+Q IVSRE++PLDSQVV+V F GG + ++IP SV IGGTFRAFS SF
Sbjct: 784 PILAASNVIVSLQHIVSREADPLDSQVVTVGKFQGGGAFNVIPHSVAIGGTFRAFSKESF 963
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
QL +RIEQVI QA+V C A V+F + E +PPTVN+ ++E+ K V+ LLG N
Sbjct: 964 MQLRQRIEQVITGQAAVQRCNATVNFLDDEKPFFPPTVNNGDLHEYFKSVAGSLLGVNNV 1143
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
+ + P+MG+ED++FY +V P FF +G+ N ++ + HSP+F I+EDALP GAA+HA+
Sbjct: 1144KDMQPLMGSEDFAFYQEVFPGYFFLLGMENVSIEHLESPHSPYFKINEDALPYGAALHAS 1323
Query: 241 IAERYL 246
+A YL
Sbjct: 1324LASSYL 1341
>TC226531 weakly similar to UP|O81702 (O81702) Gr1-protein, partial (23%)
Length = 756
Score = 252 bits (644), Expect = 1e-67
Identities = 120/144 (83%), Positives = 130/144 (89%)
Frame = +1
Query: 107 VIGGTFRAFSNTSFYQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEH 166
V+ GTFRA SFYQLLERIEQVIV+Q SVY C AEVDFFEKEYTIYPPTVND++MYEH
Sbjct: 1 VLLGTFRAXXTXSFYQLLERIEQVIVEQTSVYRCLAEVDFFEKEYTIYPPTVNDNRMYEH 180
Query: 167 VKKVSIDLLGQKNFRVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTI 226
VKKVSIDLLG KNFRVVPPMMGAED+SFYS+V+PS FFYIG+RNETLGSTHTGHSP+F I
Sbjct: 181 VKKVSIDLLGHKNFRVVPPMMGAEDFSFYSEVVPSGFFYIGVRNETLGSTHTGHSPYFMI 360
Query: 227 DEDALPIGAAVHATIAERYLNEHG 250
DED LPIGAA HA+IAERYL EHG
Sbjct: 361 DEDVLPIGAAAHASIAERYLIEHG 432
>TC225941 similar to UP|O81642 (O81642) IAA-Ala hydrolase (IAR3)
(At1g51760/F19C24_4), partial (54%)
Length = 1052
Score = 226 bits (575), Expect = 1e-59
Identities = 117/247 (47%), Positives = 170/247 (68%), Gaps = 1/247 (0%)
Frame = +3
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
+I GAL++V AIF +HV E G + SR GP+LAG G F A ISGK AA P++S D
Sbjct: 42 IIDSGALDNVTAIFGLHVVPELRVGEVASRSGPVLAGSGIFEAKISGKGGHAAIPQHSID 221
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
P+LAAS +IS+Q +VSRE++PL+ QVV+V+ F GG + ++IPD V IGGTFRAFS +
Sbjct: 222 PLLAASNVIISLQHLVSREADPLEPQVVTVSKFQGGAAFNVIPDYVTIGGTFRAFSGETL 401
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
L +RIEQVI+ QA+V C A V+FF++E +YPPTVN ++++ V+ +L+G N
Sbjct: 402 QHLKQRIEQVIIGQAAVQRCNASVNFFDEEKPLYPPTVNHGELHKLFLDVAGNLIGINNV 581
Query: 181 RV-VPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHA 239
+ P MG+ED++FY +VIP +F +G+++ + HSP+ I+E+ LP GA++HA
Sbjct: 582 IIDESPSMGSEDFAFYQEVIPGYYFMLGVKSSP-EPNQSLHSPYLKINENGLPYGASLHA 758
Query: 240 TIAERYL 246
++A YL
Sbjct: 759 SLAANYL 779
>TC225942 similar to UP|O81642 (O81642) IAA-Ala hydrolase (IAR3)
(At1g51760/F19C24_4), partial (48%)
Length = 685
Score = 217 bits (552), Expect = 5e-57
Identities = 108/213 (50%), Positives = 148/213 (68%)
Frame = +2
Query: 34 LLAGCGFFRAVISGKRASAANPRNSADPVLAASAAVISIQGIVSRESNPLDSQVVSVTSF 93
+ AG GFF A I+G+ AA P++S DP+LAAS ++S+Q IVSRE +PLDSQVV+V F
Sbjct: 14 IFAGSGFFEATINGRGGHAAIPQHSIDPILAASNVIVSLQHIVSREVDPLDSQVVTVGKF 193
Query: 94 NGGNSHDMIPDSVVIGGTFRAFSNTSFYQLLERIEQVIVQQASVYSCFAEVDFFEKEYTI 153
GG + ++IPDSV IGGTFRAFS SF QL +RIEQVI QA+V C A V+F + E
Sbjct: 194 QGGGAFNVIPDSVTIGGTFRAFSKESFMQLRQRIEQVITGQAAVQRCNATVNFLDDEKPF 373
Query: 154 YPPTVNDDQMYEHVKKVSIDLLGQKNFRVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETL 213
PPTVN+ ++ + + V+ LLG N + + P+MG+ED++FY +V P FF +G+ N +
Sbjct: 374 SPPTVNNGDLHGYFESVAGSLLGVNNVKEMQPLMGSEDFAFYQEVFPGYFFLLGMDNASN 553
Query: 214 GSTHTGHSPHFTIDEDALPIGAAVHATIAERYL 246
+ HSP+F I+EDALP GAA+H ++A YL
Sbjct: 554 EHLESPHSPYFKINEDALPYGAALHVSLASSYL 652
>TC225938 similar to UP|Q9SWX9 (Q9SWX9) Auxin conjugate hydrolase (ILL5),
partial (85%)
Length = 1727
Score = 213 bits (543), Expect = 5e-56
Identities = 112/247 (45%), Positives = 162/247 (65%), Gaps = 1/247 (0%)
Frame = +1
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
++ GAL++V AIF +HV+ + P G + SR GPL AG G F A+I GK AA P+ S D
Sbjct: 568 ILDAGALDNVTAIFGLHVTPDIPVGEVASRCGPLSAGSGVFEAIIRGKGGHAALPQLSID 747
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
PV+AA+ +IS+Q +VSRE++PLD QV+++ GG++ ++IPD V IGGTFRAFS
Sbjct: 748 PVMAATNVIISLQNLVSREADPLDPQVLTIAKLQGGDAFNVIPDYVTIGGTFRAFSRERL 927
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLG-QKN 179
L +RIEQVI+ QA+V C A V+F ++E +YPPTVN+ +++ V+ +LLG K
Sbjct: 928 EHLKQRIEQVIIGQAAVQRCNATVNFLDEENPLYPPTVNNGDLHKFFVDVAGNLLGINKV 1107
Query: 180 FRVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHA 239
+ M AED++FY + IP +F +G+ + HSP+ I+ED LP GAA+HA
Sbjct: 1108DTNMEQDMAAEDFAFYQEFIPGYYFTLGMEIASSEPVAPLHSPYLVINEDGLPYGAALHA 1287
Query: 240 TIAERYL 246
++A YL
Sbjct: 1288SLATGYL 1308
>TC231896 similar to UP|Q84XG9 (Q84XG9) IAA-amino acid hydrolase, partial
(36%)
Length = 730
Score = 197 bits (502), Expect = 3e-51
Identities = 95/190 (50%), Positives = 133/190 (70%)
Frame = +3
Query: 60 DPVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTS 119
DP+ +IS+Q +V RE++ DSQVV+V F GGN+ ++IPDSV IGGTFRAFS S
Sbjct: 9 DPIXXTXNVIISLQHLVXREADXXDSQVVTVGKFQGGNAFNVIPDSVTIGGTFRAFSKES 188
Query: 120 FYQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKN 179
F QL +RIEQV++ QA+V C A V+FFE E +P T+N++ ++EH V+++LLG
Sbjct: 189 FQQLRQRIEQVVIAQAAVLRCNATVNFFEGEKPFFPATINNNDLHEHFGTVAVNLLGINK 368
Query: 180 FRVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHA 239
+PP+MGAED+SFY +V+P F +IGI+N + HSP+F I+ED LP GAA+HA
Sbjct: 369 VNDMPPLMGAEDFSFYQEVMPGYFAFIGIQNPSHEKLEQVHSPYFKINEDVLPYGAALHA 548
Query: 240 TIAERYLNEH 249
++A YL +H
Sbjct: 549 SLAVSYLLKH 578
>AI965412
Length = 411
Score = 162 bits (410), Expect = 1e-40
Identities = 81/136 (59%), Positives = 103/136 (75%)
Frame = +3
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
++ GALE+V AIF +HV+ P G + SR GPLLAG GFF A+ISGK AA P+ S D
Sbjct: 3 ILDAGALENVAAIFGLHVTPNFPIGEVASRSGPLLAGSGFFEAIISGKGGHAAIPQQSID 182
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
P+LA S +IS+Q +VSRE++PLDSQVV+V F GGN+ ++IPDSV IGGTFRAFS SF
Sbjct: 183 PILATSNVIISLQHLVSREADPLDSQVVTVGKFQGGNAFNVIPDSVTIGGTFRAFSKESF 362
Query: 121 YQLLERIEQVIVQQAS 136
QL +RIEQV++ QA+
Sbjct: 363 QQLRQRIEQVVIAQAA 410
>TC221705 weakly similar to UP|O81641 (O81641) IAA-amino acid hydrolase
homolog ILL3, partial (24%)
Length = 550
Score = 140 bits (354), Expect = 4e-34
Identities = 65/140 (46%), Positives = 93/140 (66%)
Frame = +3
Query: 110 GTFRAFSNTSFYQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKK 169
GT R+ +N Y +R++++I QA+V+ C A VDF E+ +T YP VND+ ++ HV++
Sbjct: 3 GTLRSLTNEGMYHFRQRLKEIIEGQAAVHRCNAYVDFKEEYFTPYPAVVNDNNLHLHVER 182
Query: 170 VSIDLLGQKNFRVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDED 229
V LLG N +M ED++F+ QVIP F IGIRN+ +G+ H+ HSP F +DE+
Sbjct: 183 VGQILLGPDNVHAAKKVMAGEDFAFFQQVIPGVLFSIGIRNDKVGAIHSPHSPFFFLDEE 362
Query: 230 ALPIGAAVHATIAERYLNEH 249
LPIGA++H IAE YLNEH
Sbjct: 363 VLPIGASLHTAIAELYLNEH 422
>TC221837 weakly similar to UP|O65840 (O65840) IAA amidohydrolase (Fragment),
partial (84%)
Length = 450
Score = 126 bits (316), Expect = 1e-29
Identities = 59/126 (46%), Positives = 82/126 (64%)
Frame = +1
Query: 120 FYQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKN 179
FY L +RIE+VI QA V+ C EV+F E+ PPT ND ++Y+ ++VS ++G+ N
Sbjct: 1 FYGLRKRIEEVIKGQAEVHRCSGEVEFCGNEHPTIPPTTNDVRIYQLARQVSSKIVGEDN 180
Query: 180 FRVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHA 239
+ P G+ED++FY + +P +F +G RNE GS H HSP+F IDED LPIGAA+HA
Sbjct: 181 IELAPLFTGSEDFAFYLEKVPGSFVLVGTRNEKSGSIHPAHSPYFFIDEDVLPIGAALHA 360
Query: 240 TIAERY 245
A Y
Sbjct: 361 AFALSY 378
>AW102109
Length = 329
Score = 87.0 bits (214), Expect = 7e-18
Identities = 43/108 (39%), Positives = 63/108 (57%)
Frame = +2
Query: 135 ASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNFRVVPPMMGAEDYSF 194
A ++ CF +V+F E+ +P T D ++Y+ + VS + G+ N + + G EDY+F
Sbjct: 5 AELHMCFGKVEFCGDEHPTFPATTTDVRIYQLARPVSSGIGGEDNIELPLAVTGREDYTF 184
Query: 195 YSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHATIA 242
Y + +P AF RN+ GS H HS +F IDEDALPI AA+ A A
Sbjct: 185 YLKEVPGAFVLGDTRNQKSGSIHAAHSSYFIIDEDALPIEAALTAACA 328
>AW278733 weakly similar to PIR|F96556|F965 IAA-Ala hydrolase (IAR3)
[imported] - Arabidopsis thaliana, partial (13%)
Length = 250
Score = 56.6 bits (135), Expect = 1e-08
Identities = 24/71 (33%), Positives = 42/71 (58%)
Frame = +2
Query: 154 YPPTVNDDQMYEHVKKVSIDLLGQKNFRVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETL 213
+PPT ND +++ V+ +LG N + + P+MG+ED +FY +V P F+ +G+ N +
Sbjct: 38 FPPTDNDGDLHQCCYSVAGSVLGVNNVKDMAPLMGSEDSAFYLEVFPGYFYLLGMDNVFI 217
Query: 214 GSTHTGHSPHF 224
+ H P+F
Sbjct: 218 ETLEFSHDPYF 250
>BU549763 similar to GP|27948556|gb| IAA-amino acid hydrolase {Oryza sativa
(indica cultivar-group)}, partial (7%)
Length = 324
Score = 52.0 bits (123), Expect = 3e-07
Identities = 24/57 (42%), Positives = 37/57 (64%), Gaps = 1/57 (1%)
Frame = -3
Query: 194 FYSQVIPSAFFYIGIRN-ETLGSTHTGHSPHFTIDEDALPIGAAVHATIAERYLNEH 249
FY +VIP +F +G++N + HSP+ I+ED LP GAA+HA++A YL ++
Sbjct: 322 FYQEVIPGYYFTLGMKNASSFEPVAPLHSPYLVINEDGLPYGAALHASLATGYLTKY 152
>AW567617
Length = 340
Score = 30.8 bits (68), Expect = 0.61
Identities = 11/30 (36%), Positives = 19/30 (62%)
Frame = +2
Query: 5 GALEDVEAIFAVHVSHEHPTGMIGSRPGPL 34
G LE++ AIF +H++ +P + S GP+
Sbjct: 251 GVLENISAIFRLHIAPTYPISEVASMSGPI 340
>TC205531 weakly similar to UP|O04559 (O04559) T7N9.16, partial (42%)
Length = 1377
Score = 28.5 bits (62), Expect = 3.0
Identities = 14/28 (50%), Positives = 15/28 (53%)
Frame = -3
Query: 93 FNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
FNG H MIP S V FR S+T F
Sbjct: 847 FNGLRLHTMIPSSPVSSSNFRVNSSTPF 764
>TC230308
Length = 624
Score = 28.5 bits (62), Expect = 3.0
Identities = 19/75 (25%), Positives = 35/75 (46%), Gaps = 5/75 (6%)
Frame = +2
Query: 146 FFEKEYTIYP-----PTVNDDQMYEHVKKVSIDLLGQKNFRVVPPMMGAEDYSFYSQVIP 200
+FE TIY N D +++ ++ V+ + QKNFR+ G +++ F +
Sbjct: 143 YFENGKTIYAVVQELKNTNQDMLFDAIQNVTGSTINQKNFRI---WKGNKNFQFAWSCLC 313
Query: 201 SAFFYIGIRNETLGS 215
+ + I+N T S
Sbjct: 314 FSCPCLDIQNNTNSS 358
>TC230307
Length = 392
Score = 28.1 bits (61), Expect = 3.9
Identities = 13/42 (30%), Positives = 22/42 (51%), Gaps = 5/42 (11%)
Frame = +2
Query: 146 FFEKEYTIYP-----PTVNDDQMYEHVKKVSIDLLGQKNFRV 182
+FE TIY N D +++ ++ V+ + QKNFR+
Sbjct: 14 YFENGKTIYAVVQELKNTNQDMLFDAIQNVTGSTINQKNFRI 139
>TC205045 similar to UP|P93166 (P93166) SCOF-1, partial (45%)
Length = 1150
Score = 27.3 bits (59), Expect = 6.7
Identities = 18/80 (22%), Positives = 34/80 (42%)
Frame = -1
Query: 23 PTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSADPVLAASAAVISIQGIVSRESNP 82
PT +G + G G V G + P AA+++ QG+ RE+
Sbjct: 718 PTERVGGGGSAAVGGSGAAAVVSVGVAVTVVVPA-------VVVAALVATQGLTGRETLV 560
Query: 83 LDSQVVSVTSFNGGNSHDMI 102
D +V ++ +GG+ + ++
Sbjct: 559 TDGTLVHPSAADGGSRNAVV 500
>TC228958 similar to UP|MCCB_ARATH (Q9LDD8) Methylcrotonyl-CoA carboxylase
beta chain, mitochondrial precursor
(3-Methylcrotonyl-CoA carboxylase 2) (MCCase beta
subunit) (3-methylcrotonyl-CoA:carbon dioxide ligase
beta subunit) , partial (63%)
Length = 1226
Score = 26.9 bits (58), Expect = 8.8
Identities = 11/41 (26%), Positives = 21/41 (50%)
Frame = -2
Query: 202 AFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHATIA 242
AF Y+G +L +TH +S + +++ IG H ++
Sbjct: 922 AFLYLGQHTSSLSTTHYRYSSIWPKEQEVGTIGTTTHCIVS 800
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.135 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,795,477
Number of Sequences: 63676
Number of extensions: 120182
Number of successful extensions: 498
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 492
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 494
length of query: 250
length of database: 12,639,632
effective HSP length: 95
effective length of query: 155
effective length of database: 6,590,412
effective search space: 1021513860
effective search space used: 1021513860
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC140916.4


