

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140913.9 - phase: 2 /pseudo
(273 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
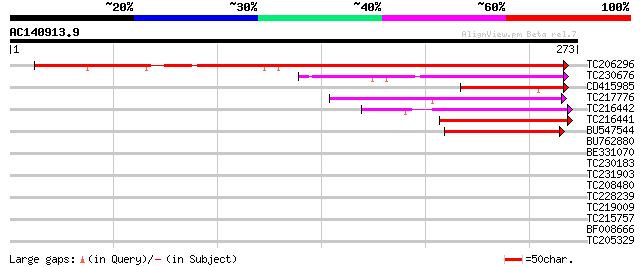
Score E
Sequences producing significant alignments: (bits) Value
TC206296 similar to UP|Q94AX5 (Q94AX5) At1g64150/F22C12_10, part... 288 1e-78
TC230676 weakly similar to UP|Q94AX5 (Q94AX5) At1g64150/F22C12_1... 96 2e-20
CD415985 similar to GP|15010676|gb| At1g64150/F22C12_10 {Arabido... 93 1e-19
TC217776 similar to UP|Q93Y38 (Q93Y38) Transmembrane protein FT2... 71 6e-13
TC216442 similar to GB|AAD49978.1|5734713|F24J5 Is a member of P... 61 5e-10
TC216441 similar to GB|AAD49978.1|5734713|F24J5 Is a member of P... 60 1e-09
BU547544 similar to GP|15450794|gb| transmembrane protein FT27/P... 55 3e-08
BU762880 39 0.003
BE331070 similar to GP|5734713|gb|A Is a member of PF|01169 Unch... 30 1.2
TC230183 homologue to UP|GLHR_ANTEL (P35409) Probable glycoprote... 28 4.4
TC231903 similar to UP|Q9MA20 (Q9MA20) T5E21.13, partial (9%) 28 4.4
TC208480 28 4.4
TC228239 weakly similar to UP|Q6IVC4 (Q6IVC4) Zinc finger protei... 28 5.8
TC219009 27 7.6
TC215757 27 7.6
BF008666 27 9.9
TC205329 similar to UP|Q9LW57 (Q9LW57) Arabidopsis thaliana geno... 27 9.9
>TC206296 similar to UP|Q94AX5 (Q94AX5) At1g64150/F22C12_10, partial (60%)
Length = 1372
Score = 288 bits (738), Expect = 1e-78
Identities = 174/272 (63%), Positives = 199/272 (72%), Gaps = 15/272 (5%)
Frame = +3
Query: 13 SEDDSVAENSDSQCQLATQNVSSD-----DASMTVLKFMLFSAFFALQDAFPAVAAS-DF 66
+ S+A S+ + LA VSS ++S +L ML AF LQ ++PA+AAS DF
Sbjct: 231 NSSQSLAHISNCKFPLARPLVSSSHPASTNSSTKLLNSMLLFAFLTLQPSYPALAASSDF 410
Query: 67 ATGLNSIPIFGDVGDLSTGFASYHRHFC*YFSLNWETRLFSLQHC*QLEIQPVLF--SLG 124
+T IFGD+GDLSTGFAS +FS + F V+F + G
Sbjct: 411 ST------IFGDIGDLSTGFAS--AFLLIFFSELGDKTFFIAALLAARNSAGVVFIGTFG 566
Query: 125 HLAH-------LRRTFHYVDELLPFRFGETDLPIDDIAAVCLLVYFGVSTLLDASSSDSQ 177
LA L RTFHYVDE+LPFRFGETDLPIDDIAAVCLLVYFGVSTLLDASSSD Q
Sbjct: 567 ALAAMTLISVVLGRTFHYVDEILPFRFGETDLPIDDIAAVCLLVYFGVSTLLDASSSDGQ 746
Query: 178 KSDDEQKEAELAVSDFSGDGAGILAAASTIVSTFLLVFVAEWGDKSFFSTIALAAASSPL 237
KSD+EQKEAELAVS+FSG+GAGIL+AAST+ STFLLVFVAEWGDKSFFSTIALAAASSPL
Sbjct: 747 KSDEEQKEAELAVSEFSGNGAGILSAASTVASTFLLVFVAEWGDKSFFSTIALAAASSPL 926
Query: 238 GVIAGSLAGHGVATLIAVLGGSLLGTFLSEKV 269
GVIAG+LAGHGVATL+AVLGGSLLGT+LSEKV
Sbjct: 927 GVIAGALAGHGVATLLAVLGGSLLGTYLSEKV 1022
>TC230676 weakly similar to UP|Q94AX5 (Q94AX5) At1g64150/F22C12_10, partial
(19%)
Length = 900
Score = 95.5 bits (236), Expect = 2e-20
Identities = 60/138 (43%), Positives = 83/138 (59%), Gaps = 8/138 (5%)
Frame = +3
Query: 140 LPFRFGETDLPIDDIAAVCLLVYFGVSTLLDASS--SDSQKSD------DEQKEAELAVS 191
+P +F +T LPI + AAV LL++FG+ + DA SD K D DE EAE V
Sbjct: 72 VPAQF-QTTLPIGEYAAVTLLLFFGLKAIKDAWDLPSDVVKGDNSSPELDELAEAEELVK 248
Query: 192 DFSGDGAGILAAASTIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVAT 251
+ + + +F LVF AEWGD+S +TIAL AA SP GV +G++AGH +AT
Sbjct: 249 EKVS--TRLSNPLEIVWKSFSLVFFAEWGDRSMLATIALGAAQSPWGVASGAIAGHLLAT 422
Query: 252 LIAVLGGSLLGTFLSEKV 269
IA+LGG+ L ++SEK+
Sbjct: 423 TIAILGGAFLANYISEKL 476
>CD415985 similar to GP|15010676|gb| At1g64150/F22C12_10 {Arabidopsis
thaliana}, partial (19%)
Length = 496
Score = 93.2 bits (230), Expect = 1e-19
Identities = 49/57 (85%), Positives = 52/57 (90%), Gaps = 5/57 (8%)
Frame = -1
Query: 218 EWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLI-----AVLGGSLLGTFLSEKV 269
EWGDKSFFSTIALAAASSPLGVIAG+LAGHGVATL+ AVLGGSLLGT+LSEKV
Sbjct: 496 EWGDKSFFSTIALAAASSPLGVIAGALAGHGVATLVTLFSLAVLGGSLLGTYLSEKV 326
>TC217776 similar to UP|Q93Y38 (Q93Y38) Transmembrane protein FT27/PFT27-like
(At5g36290), partial (76%)
Length = 1436
Score = 70.9 bits (172), Expect = 6e-13
Identities = 40/120 (33%), Positives = 64/120 (53%), Gaps = 6/120 (5%)
Frame = +2
Query: 155 AAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELAVSDFSGDGAGILA------AASTIV 208
AA L +FG+ L A S KS +++ E+ G G + +
Sbjct: 734 AATVLYAFFGLRLLYIAWRSSDSKSSQKKEMEEVEEKLDGGQGKTSVRRFFSRFCTPIFL 913
Query: 209 STFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEK 268
+F+L F+AEWGD+S +TIALA + +GV G+ GH + T +AV+GGS+L + +S++
Sbjct: 914 ESFILTFLAEWGDRSQIATIALATHKNAIGVAVGATIGHTICTSLAVVGGSMLASKISQR 1093
>TC216442 similar to GB|AAD49978.1|5734713|F24J5 Is a member of PF|01169
Uncharacterized (transmembrane domain) protein family.
{Arabidopsis thaliana;} , partial (96%)
Length = 1268
Score = 61.2 bits (147), Expect = 5e-10
Identities = 34/103 (33%), Positives = 55/103 (53%), Gaps = 1/103 (0%)
Frame = +2
Query: 170 DASSSDSQKSDDEQKEAELA-VSDFSGDGAGILAAASTIVSTFLLVFVAEWGDKSFFSTI 228
+ ++ +S K DD K+ + + +S F + + F + F EWGDKS +TI
Sbjct: 551 NGATKNSNKDDDATKKHKRSFLSQFF---------SPIFLQAFSITFFGEWGDKSQLATI 703
Query: 229 ALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKVTS 271
LAA +P GV+ G + G + T AV+GG L + +SEK+ +
Sbjct: 704 GLAADENPFGVVLGGILGQALCTAAAVVGGKSLASQISEKIVA 832
Score = 30.4 bits (67), Expect = 0.90
Identities = 21/61 (34%), Positives = 33/61 (53%), Gaps = 4/61 (6%)
Frame = +2
Query: 202 AAASTIVSTFL----LVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLG 257
A S+IV F + ++E GDK+FF+ LA V++G L+ V T+++VL
Sbjct: 203 ANMSSIVQGFTKSLAMTILSEIGDKTFFAAAILAMRHPRRLVLSGCLSALIVMTILSVLV 382
Query: 258 G 258
G
Sbjct: 383 G 385
>TC216441 similar to GB|AAD49978.1|5734713|F24J5 Is a member of PF|01169
Uncharacterized (transmembrane domain) protein family.
{Arabidopsis thaliana;} , partial (43%)
Length = 572
Score = 59.7 bits (143), Expect = 1e-09
Identities = 27/64 (42%), Positives = 39/64 (60%)
Frame = +1
Query: 208 VSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSE 267
+ F + F EWGDKS +TI LAA +P GV+ G + G + T AV+GG L + +SE
Sbjct: 85 LQAFSITFFGEWGDKSQLATIGLAADENPFGVVLGGILGQALCTSAAVVGGKSLASQISE 264
Query: 268 KVTS 271
K+ +
Sbjct: 265 KIVA 276
>BU547544 similar to GP|15450794|gb| transmembrane protein FT27/PFT27-like
{Arabidopsis thaliana}, partial (28%)
Length = 411
Score = 55.1 bits (131), Expect = 3e-08
Identities = 26/58 (44%), Positives = 39/58 (66%)
Frame = -1
Query: 210 TFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSE 267
+F+L F+AEWGD+S +TIALA + V GS G + T +AV+GGS+L + +S+
Sbjct: 408 SFILTFLAEWGDRSQIATIALATHKNAXXVAVGSTIGPTICTSLAVVGGSMLASKISQ 235
>BU762880
Length = 423
Score = 38.5 bits (88), Expect = 0.003
Identities = 17/42 (40%), Positives = 26/42 (61%)
Frame = -1
Query: 230 LAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKVTS 271
LAA +P GV+ G + G + T AV+GG L + +SEK+ +
Sbjct: 423 LAADENPFGVVLGGILGQALCTAAAVVGGKSLASQISEKIVA 298
>BE331070 similar to GP|5734713|gb|A Is a member of PF|01169 Uncharacterized
(transmembrane domain) protein family., partial (40%)
Length = 445
Score = 30.0 bits (66), Expect = 1.2
Identities = 16/46 (34%), Positives = 27/46 (57%)
Frame = +2
Query: 213 LVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGG 258
+ ++E GDK+FF+ LA V++G L+ V T+++VL G
Sbjct: 32 MTILSEIGDKTFFAAAILAMRHPRRLVLSGCLSALMVMTILSVLVG 169
>TC230183 homologue to UP|GLHR_ANTEL (P35409) Probable glycoprotein hormone
G-protein coupled receptor precursor, partial (6%)
Length = 878
Score = 28.1 bits (61), Expect = 4.4
Identities = 22/61 (36%), Positives = 30/61 (49%)
Frame = +2
Query: 200 ILAAASTIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGS 259
++A + +V++ LV VA + +AAAS L V AGSLA A L V S
Sbjct: 188 VVAPLALVVASLALVLVASLA-------LVVAAASLALVVAAGSLALVAAALLALVAAAS 346
Query: 260 L 260
L
Sbjct: 347 L 349
>TC231903 similar to UP|Q9MA20 (Q9MA20) T5E21.13, partial (9%)
Length = 723
Score = 28.1 bits (61), Expect = 4.4
Identities = 17/38 (44%), Positives = 22/38 (57%)
Frame = -1
Query: 200 ILAAASTIVSTFLLVFVAEWGDKSFFSTIALAAASSPL 237
+ +AS STFL W +KS FS A AAA++PL
Sbjct: 606 VYCSASGGSSTFLTGL--NWAEKSGFSAAAAAAATTPL 499
>TC208480
Length = 1576
Score = 28.1 bits (61), Expect = 4.4
Identities = 17/43 (39%), Positives = 21/43 (48%)
Frame = -1
Query: 105 LFSLQHC*QLEIQPVLFSLGHLAHLRRTFHYVDELLPFRFGET 147
LFSL +C L P FSL L R F ++ +L FR T
Sbjct: 568 LFSLCNCAILVSSPTSFSLKELNSFSRRFIFLSKLSAFRLKTT 440
>TC228239 weakly similar to UP|Q6IVC4 (Q6IVC4) Zinc finger protein
(Zinc-finger transcription factor), partial (32%)
Length = 1244
Score = 27.7 bits (60), Expect = 5.8
Identities = 15/54 (27%), Positives = 27/54 (49%)
Frame = -3
Query: 150 PIDDIAAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELAVSDFSGDGAGILAA 203
P D + + + +LLD + S + SDDE A + + + +G GA ++A
Sbjct: 345 PTDPVDVLIEDILDATDSLLDPAPSSGRNSDDEDDGARVPIEEPTG*GATGISA 184
>TC219009
Length = 1139
Score = 27.3 bits (59), Expect = 7.6
Identities = 21/60 (35%), Positives = 28/60 (46%), Gaps = 1/60 (1%)
Frame = +3
Query: 93 FC*YFSLNWETRLFSLQHC*QLEIQPVLFSLGHLAHLRRT-FHYVDELLPFRFGETDLPI 151
F *+FS+ WE L L C +QP++ L H T FH + L F F E + I
Sbjct: 945 FF*FFSICWEFALSMLLPCLH-SLQPIMGCLIHDCFCIET*FHVLSYLGRFTFSEIHVSI 1121
>TC215757
Length = 777
Score = 27.3 bits (59), Expect = 7.6
Identities = 12/28 (42%), Positives = 17/28 (59%)
Frame = +1
Query: 100 NWETRLFSLQHC*QLEIQPVLFSLGHLA 127
NWE+ S+QH I P+LFS H++
Sbjct: 10 NWESEALSMQH--SFTISPILFSSLHVS 87
>BF008666
Length = 433
Score = 26.9 bits (58), Expect = 9.9
Identities = 18/46 (39%), Positives = 21/46 (45%)
Frame = -3
Query: 4 DLSKGFVRASEDDSVAENSDSQCQLATQNVSSDDASMTVLKFMLFS 49
DLS+GF +DDSV N D DD+ TVL L S
Sbjct: 293 DLSEGF*THDDDDSVDSNDD------------DDSHSTVLWMQLLS 192
>TC205329 similar to UP|Q9LW57 (Q9LW57) Arabidopsis thaliana genomic DNA,
chromosome 3, P1 clone: MLM24, partial (62%)
Length = 1036
Score = 26.9 bits (58), Expect = 9.9
Identities = 16/45 (35%), Positives = 22/45 (48%)
Frame = +2
Query: 145 GETDLPIDDIAAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELA 189
G+T I + L G++ L AS D +K+DD KE E A
Sbjct: 209 GDTSDSISSLKLNLLSAVSGLNRGLAASEDDLRKADDAAKELEAA 343
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.138 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,178,895
Number of Sequences: 63676
Number of extensions: 130304
Number of successful extensions: 927
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 913
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 922
length of query: 273
length of database: 12,639,632
effective HSP length: 96
effective length of query: 177
effective length of database: 6,526,736
effective search space: 1155232272
effective search space used: 1155232272
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC140913.9


