

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140850.4 - phase: 0 /pseudo
(1010 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
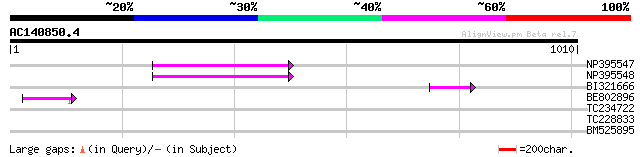
NP395547 reverse transcriptase [Glycine max] 102 8e-22
NP395548 reverse transcriptase [Glycine max] 94 4e-19
BI321666 60 5e-09
BE802896 56 9e-08
TC234722 similar to UP|Q6WAY9 (Q6WAY9) Pol (Fragment), partial (... 37 0.033
TC228833 similar to UP|Q9ZPI4 (Q9ZPI4) Molybdopterin synthase su... 33 0.82
BM525895 30 4.1
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 102 bits (254), Expect = 8e-22
Identities = 90/251 (35%), Positives = 118/251 (46%)
Frame = +2
Query: 255 RRWLS*LKSA*SIQSRIVHG*ARCK*CRRRVV*R*SKLRRMSSSLQEPSSGGGCVLITED 314
+R+ S + S Q +I G R K +++ *+ K+ MS LQE S G CVLI
Sbjct: 8 KRYSSY*RQGSSTQFQIAPGLVRFKLFQKKEG*QW*KMIEMS*FLQEESPDGECVLIIGS 187
Query: 315 LTKLQGKIISHCLSWTKCLNAFPVKLIIVSLMATRAITKSQSIRRTMRRQLLHVL*ESLH 374
K Q K I+H SW KCL VS T+ + Q I R ++QLLHVL L
Sbjct: 188 SMKPQEKTITHFPSWIKCLRDLQGNPSTVSWTDTQVTIRLQWILRIKKKQLLHVLSVFLL 367
Query: 375 IEECCLVYPMRLQHSNDVCKRYSPT*LRRVLRYSWMIFLFLDHPMMFA*TTWTPC*SGVK 434
I C VY M L DV ++ T R VL+ WMIF L H + A* C + VK
Sbjct: 368 IAACRSVYVMPLLLFRDV*WQFLMTW*RNVLKSLWMIFRSLVHLLEIA*QI*RKCYNVVK 547
Query: 435 KPILCLIGKNVI*W*PRALFLVTRFHQEALKLTRERLM*SESSHHG*MSMGSVAFLVMPS 494
I CL GKNV W + L T+ +E L+ ++ M + H M F VM
Sbjct: 548 NLIWCLTGKNVTLWYKKVLC*DTKSLKEELRWLKKN*MLLINFHPQLM*KAYTVFWVMLD 727
Query: 495 FTGDLSKTSQR 505
F GD +TS +
Sbjct: 728 FIGDS*RTSPK 760
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 93.6 bits (231), Expect = 4e-19
Identities = 87/252 (34%), Positives = 114/252 (44%)
Frame = +2
Query: 254 ARRWLS*LKSA*SIQSRIVHG*ARCK*CRRRVV*R*SKLRRMSSSLQEPSSGGGCVLITE 313
ARR+ S + S S G*A+ CRR+ *+ +++RM+ E S G IT
Sbjct: 5 ARRF*SF*RLGLSTPSPTALG*AQSWWCRRKRA*QSFEMKRMT*YQHELSLVGNYASITA 184
Query: 314 DLTKLQGKIISHCLSWTKCLNAFPVKLIIVSLMATRAITKSQSIRRTMRRQLLHVL*ESL 373
TK QGK IS SW +C LII S M T I + R RR H L SL
Sbjct: 185 SSTKPQGKTISLYPSWIRCWRDLQDTLIIASWMHTLDIIRLL*TPRIRRRWPSHALLVSL 364
Query: 374 HIEECCLVYPMRLQHSNDVCKRYSPT*LRRVLRYSWMIFLFLDHPMMFA*TTWTPC*SGV 433
I+ L M L HS C + * R+ +YSWM F +L * +W
Sbjct: 365 PIDGFHLGCAMHLPHSKCACWPFLQI*WRKASKYSWMTFQYLCPH*KVV*RSWRWYYKDA 544
Query: 434 KKPILCLIGKNVI*W*PRALFLVTRFHQEALKLTRERLM*SESSHHG*MSMGSVAFLVMP 493
K IG++V W +A +F E L+ T++RLM +S HH M S A P
Sbjct: 545 WKQT*Y*IGRSVTSWFEKA*S*AIKFRPEELR*TKQRLMSLKSCHHHQMLKASGAS*DKP 724
Query: 494 SFTGDLSKTSQR 505
T D S+TSQ+
Sbjct: 725 GSTEDSSRTSQK 760
>BI321666
Length = 430
Score = 60.1 bits (144), Expect = 5e-09
Identities = 37/83 (44%), Positives = 45/83 (53%)
Frame = +3
Query: 748 LQKYSNPVSIGQNFSKMLTLIASFVMNVKEPDQCQSGMKCLYKVFLKWKCLIVGVLILLA 807
LQ Y N I FSKM + S V+N KE + GM C F +W+ L +G LILL
Sbjct: 6 LQMYFNLDFICHPFSKMHMFMLSHVINAKELEVFPRGMNCHCIPFWRWRSLTIGALILLV 185
Query: 808 PSHHRTQMNTSW*P*IMSQSGWK 830
P H + MNT * *IM SGW+
Sbjct: 186 PFLHHSPMNTF****IM*ASGWR 254
>BE802896
Length = 416
Score = 55.8 bits (133), Expect = 9e-08
Identities = 42/99 (42%), Positives = 57/99 (57%), Gaps = 4/99 (4%)
Frame = +3
Query: 24 AWSV*TTILEPTRYERNFSNAYTIASNSFLVVA*LICASFKTLLA*CIAWKILFLR*PST 83
AWS+*T + P++ E NFS A T+A++SF VV * A+ K L * I L + * T
Sbjct: 3 AWSI*TIMRVPSK*E*NFSRAKTMANSSFSVVV*FAWAASKVLEE*YITLGNLSIF*ART 182
Query: 84 TPTT*SLASHINSEFKFQFGA----VIIGAEVNRSPTHL 118
TP * ASHI+S+ Q GA V+I A + +S +L
Sbjct: 183 TPNA*FDASHISSKGAVQSGAWMMGVVISALLRQSKAYL 299
Score = 37.4 bits (85), Expect = 0.033
Identities = 44/136 (32%), Positives = 62/136 (45%)
Frame = -3
Query: 489 FLVMPSFTGDLSKTSQR*QSH*EICSTKISLFSLIMIV*MLLKL*RSG*PPHQ*SQLQIG 548
FLVM TG L + ++* H C + +L++ LL + * P S+ G
Sbjct: 411 FLVMQDSTGAL*EILEK*PFHYPTCCKRR*SLTLMINANRLLIASKEH*LPPPSSRHPTG 232
Query: 549 T*TLN*CVMLVIMLLVSC*VNARTKFSMLYIMRVKF*MKNKLIMLPLKKSCSQ*YMHWKN 608
L+ CVM IM + YI+ + *M K I+L L+KS *++ KN
Sbjct: 231 QPLLSLCVMHQIMHWGLSLLRKLINCLG*YIILLGL*MLPKRIILLLRKSY*P*FLLLKN 52
Query: 609 FVHTS*VPKLLFTLTM 624
F+ V LLF LTM
Sbjct: 51 FILICLVLALLFILTM 4
>TC234722 similar to UP|Q6WAY9 (Q6WAY9) Pol (Fragment), partial (32%)
Length = 482
Score = 37.4 bits (85), Expect = 0.033
Identities = 25/71 (35%), Positives = 32/71 (44%)
Frame = -1
Query: 891 ILKPTVKPRSQTEKLKELWRK*SQHLGRIGRRSLMTPCGLIG*HTSLL*VTHHFKWFTTS 950
I K K + Q ++ E R H +IG S M GL G H+ L * HHF W
Sbjct: 230 IPKQMAKQKFQIKR*SEYLRILLCHQEKIGL*SWMMLSGLTGLHSRLP*ACHHFNWSMVK 51
Query: 951 HVIYQ*ISNTK 961
H Y+ +TK
Sbjct: 50 HATYRWSWSTK 18
>TC228833 similar to UP|Q9ZPI4 (Q9ZPI4) Molybdopterin synthase sulphurylase
(Fragment), partial (94%)
Length = 1414
Score = 32.7 bits (73), Expect = 0.82
Identities = 20/71 (28%), Positives = 35/71 (49%), Gaps = 4/71 (5%)
Frame = +2
Query: 315 LTKLQGKIISH----CLSWTKCLNAFPVKLIIVSLMATRAITKSQSIRRTMRRQLLHVL* 370
L +LQ ++SH CLSW CL+ F + + L + + + K Q + + L ++
Sbjct: 596 LLRLQLLLVSHSQEGCLSWMHCLDGFVLSKLEEGLCSVKLVEKMQHLPNSNSENL--IMR 769
Query: 371 ESLHIEECCLV 381
SL + CL+
Sbjct: 770 SSLRLPCVCLL 802
>BM525895
Length = 428
Score = 30.4 bits (67), Expect = 4.1
Identities = 30/84 (35%), Positives = 43/84 (50%), Gaps = 1/84 (1%)
Frame = +3
Query: 124 RKRRN*KNA*TSWKP*RRFLQRKQKWRS*KRKKRLKNQIMN*KCCRHILIMCS-LKKTAL 182
RKR+N*K A K R QR++K R KR+++L+NQ M R + S LK + +
Sbjct: 24 RKRKN*KKAKRRTKKQRN--QRRKKRR--KRREKLRNQKMTKNLRRKLRRQKSCLKCSCI 191
Query: 183 NKSLLATLYQAMKKTS*LKFLSKI 206
K +LA K + K S+I
Sbjct: 192 VKGVLARFVAPSKDSQGSKIYSQI 263
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.358 0.155 0.550
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 47,939,457
Number of Sequences: 63676
Number of extensions: 700114
Number of successful extensions: 7037
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 3836
Number of HSP's successfully gapped in prelim test: 278
Number of HSP's that attempted gapping in prelim test: 3093
Number of HSP's gapped (non-prelim): 4322
length of query: 1010
length of database: 12,639,632
effective HSP length: 107
effective length of query: 903
effective length of database: 5,826,300
effective search space: 5261148900
effective search space used: 5261148900
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 64 (29.3 bits)
Medicago: description of AC140850.4


