

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140850.3 + phase: 0 /pseudo
(650 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
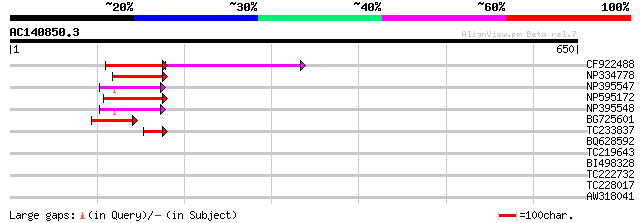
Score E
Sequences producing significant alignments: (bits) Value
CF922488 110 5e-28
NP334778 reverse transcriptase [Glycine max] 106 3e-23
NP395547 reverse transcriptase [Glycine max] 77 2e-14
NP595172 polyprotein [Glycine max] 76 5e-14
NP395548 reverse transcriptase [Glycine max] 67 3e-11
BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] -... 51 2e-06
TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 44 2e-04
BQ628592 35 0.14
TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotei... 33 0.40
BI498328 32 0.68
TC222732 similar to UP|Q9LM48 (Q9LM48) F2E2.18, partial (24%) 30 2.6
TC228017 29 5.7
AW318041 29 5.7
>CF922488
Length = 741
Score = 110 bits (275), Expect(2) = 5e-28
Identities = 53/70 (75%), Positives = 59/70 (83%)
Frame = +3
Query: 111 VPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVFSFMDGFSGYNLIKMSP 170
V K+DGKV MCVD+RDLN ASPKD FPLPHI+VLVDNT FSFMDGFSGYN IK++P
Sbjct: 3 VLKEDGKV*MCVDYRDLN*ASPKDKFPLPHINVLVDNTTSFSQFSFMDGFSGYNQIKIAP 182
Query: 171 EDREKTSFIT 180
ED EKT+FIT
Sbjct: 183 EDMEKTTFIT 212
Score = 32.7 bits (73), Expect(2) = 5e-28
Identities = 48/161 (29%), Positives = 69/161 (42%)
Frame = +1
Query: 179 ITHGILSATK*CRSA**MLVLPTKEE*LLCFMT*FPKKSKYMWTI*L*SQKMRSNMSNT* 238
+ +G SA + CR * ML T T* ++ + W * *+Q+ R N +
Sbjct: 208 LLYGEPSAIRLCRLG*RMLGQHTSGPWWHYSRT*CTRR*RSTWMT*S*NQERRRNTLSIC 387
Query: 239 QKCSKG*ESTSFD*TLTNVRSASDLENY*ASLSVKRALKSTLIKSVPSEKCQPRRPRNKS 298
+ C +T D* +V + E+ L + + + S + R+KS
Sbjct: 388 ESCLGDYVNTG*D*IPQSVCLR*NPESCSTLLIAREE*RWIRTR*K*SLRWPSHIQRSKS 567
Query: 299 EVSSGD*ITSPDSYLT*PQPAGRSSSYSGRISLLYGTMNAK 339
+VS G * TS DSY +* A S RISL GTM K
Sbjct: 568 KVSWGG*TTS*DSYHS*LPLASLFSYCCARISLSNGTMIVK 690
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 106 bits (265), Expect = 3e-23
Identities = 48/62 (77%), Positives = 55/62 (88%)
Frame = +3
Query: 119 RMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVFSFMDGFSGYNLIKMSPEDREKTSF 178
RMCVD+RDLN+ASPKDNFPLPHID+L+ N A +FSFMDGFSGYN IKM+PED EKT+F
Sbjct: 3 RMCVDYRDLNRASPKDNFPLPHIDILMANMASFALFSFMDGFSGYNQIKMAPEDMEKTTF 182
Query: 179 IT 180
IT
Sbjct: 183 IT 188
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 77.0 bits (188), Expect = 2e-14
Identities = 36/93 (38%), Positives = 55/93 (58%), Gaps = 18/93 (19%)
Frame = +1
Query: 104 WVANIVPVPKKDGKV------------------RMCVDFRDLNKASPKDNFPLPHIDVLV 145
WV+ + VPKK G RMC+D+R LN+A+ KD++PLP +D ++
Sbjct: 64 WVSPVQVVPKKGGMTVVKNDRNELIPTRRVTRWRMCIDYRKLNEATRKDHYPLPFMDQML 243
Query: 146 DNTAQSKVFSFMDGFSGYNLIKMSPEDREKTSF 178
A+ + F+DG+SGYN I + P+D+EKT+F
Sbjct: 244 KRLARQSFYRFLDGYSGYNQIAVDPQDQEKTAF 342
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 75.9 bits (185), Expect = 5e-14
Identities = 37/74 (50%), Positives = 50/74 (67%)
Frame = +1
Query: 108 IVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVFSFMDGFSGYNLIK 167
I+ V KKDG R C D+R LN + KD+FP+P +D L+D ++ FS +D SGY+ I
Sbjct: 1909 ILLVKKKDGSWRFCTDYRALNAITVKDSFPMPTVDELLDELHGAQYFSKLDLRSGYHQIL 2088
Query: 168 MSPEDREKTSFITH 181
+ PEDREKT+F TH
Sbjct: 2089 VQPEDREKTAFRTH 2130
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 66.6 bits (161), Expect = 3e-11
Identities = 31/93 (33%), Positives = 53/93 (56%), Gaps = 18/93 (19%)
Frame = +1
Query: 104 WVANIVPVPKKDGKV------------------RMCVDFRDLNKASPKDNFPLPHIDVLV 145
WV+ ++ V KK+G ++C+D+R LN+A+ KD+FPLP +D ++
Sbjct: 64 WVSPVLVVSKKEGMTVIRNEKNDLIPTRTVTSWKLCIDYRKLNEATRKDHFPLPFMDQML 243
Query: 146 DNTAQSKVFSFMDGFSGYNLIKMSPEDREKTSF 178
+ A + F+D + GYN I + P+D+EK +F
Sbjct: 244 ERLAGHAYYCFLDAYFGYNQIVVDPKDQEKMAF 342
>BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (1%)
Length = 285
Score = 50.8 bits (120), Expect = 2e-06
Identities = 23/52 (44%), Positives = 34/52 (65%)
Frame = -3
Query: 95 FLMTVEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVD 146
F+ + Y + ++V V K +GK R+C D+ DLN A PKD +PLP+ID + D
Sbjct: 163 FIRDINYST*LFSVVMVKKPNGKWRICTDYIDLN*ACPKDAYPLPNIDHMTD 8
>TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 402
Score = 43.9 bits (102), Expect = 2e-04
Identities = 20/27 (74%), Positives = 23/27 (85%)
Frame = +2
Query: 154 FSFMDGFSGYNLIKMSPEDREKTSFIT 180
FSFMDGFSGYN I M+ ED EKT+F+T
Sbjct: 2 FSFMDGFSGYNQI*MAREDVEKTTFVT 82
>BQ628592
Length = 423
Score = 34.7 bits (78), Expect = 0.14
Identities = 26/64 (40%), Positives = 33/64 (50%)
Frame = -2
Query: 338 AKKLLIASRITCCNHLSLSHPWKEGL*LCICQCLMNLWDAYLVNKMRLERKNMLSTI*AR 397
+KK ASRI C+ H +E L C C +LWDA + L ++N TI*AR
Sbjct: 374 SKKSNRASRIPRCS----CHL*QEDLFSCT*LC*TSLWDACWFSTTTLGKRNKPFTI*AR 207
Query: 398 SSPT 401
S PT
Sbjct: 206 SLPT 195
>TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotein (Fragment),
partial (8%)
Length = 1320
Score = 33.1 bits (74), Expect = 0.40
Identities = 12/18 (66%), Positives = 14/18 (77%)
Frame = +3
Query: 95 FLMTVEYPEWVANIVPVP 112
FL YP+WVANIVP+P
Sbjct: 1266 FLAVARYPKWVANIVPIP 1319
>BI498328
Length = 335
Score = 32.3 bits (72), Expect = 0.68
Identities = 22/53 (41%), Positives = 25/53 (46%)
Frame = +2
Query: 529 SSMVLSMLMVKESGQSLYPRRGITSLLPPGFCSNVQTIWPNTKRVSLGSMKQL 581
+SM ML E GQSLYPR L V TIWP+TK G + L
Sbjct: 161 ASMGHPMLWATE*GQSLYPRMISVFLSRLD*VLIVPTIWPSTKHAPSGFRRPL 319
>TC222732 similar to UP|Q9LM48 (Q9LM48) F2E2.18, partial (24%)
Length = 930
Score = 30.4 bits (67), Expect = 2.6
Identities = 19/47 (40%), Positives = 24/47 (50%)
Frame = +3
Query: 455 HAGKCSCPNMTLCSRLKKQSKVAFLPIILPTNLLMITNQLSLISPMK 501
H + P MT+ S LKKQ V +IL LLMI L ++ MK
Sbjct: 657 HCSLLTFPLMTMMSTLKKQHLVHMKRLILLITLLMIMILLVMVGLMK 797
>TC228017
Length = 752
Score = 29.3 bits (64), Expect = 5.7
Identities = 16/46 (34%), Positives = 24/46 (51%), Gaps = 3/46 (6%)
Frame = +1
Query: 79 WLSRLRVRFKSRLMRVFLMTVE-YPEW--VANIVPVPKKDGKVRMC 121
W +L + +K + RV ++ YP W V N+V PKK K+ C
Sbjct: 445 WEYKLLIGWKQLVPRVLKFSISCYPTWFLVVNVVS*PKK*KKINFC 582
>AW318041
Length = 310
Score = 29.3 bits (64), Expect = 5.7
Identities = 9/25 (36%), Positives = 16/25 (64%)
Frame = +1
Query: 351 NHLSLSHPWKEGL*LCICQCLMNLW 375
N+++ +HPW G C C ++N+W
Sbjct: 166 NNVASTHPWALG*ETCCCYIMLNVW 240
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.347 0.151 0.514
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,396,944
Number of Sequences: 63676
Number of extensions: 476905
Number of successful extensions: 4257
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 3065
Number of HSP's successfully gapped in prelim test: 125
Number of HSP's that attempted gapping in prelim test: 1153
Number of HSP's gapped (non-prelim): 3246
length of query: 650
length of database: 12,639,632
effective HSP length: 103
effective length of query: 547
effective length of database: 6,081,004
effective search space: 3326309188
effective search space used: 3326309188
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC140850.3


