

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140721.10 + phase: 0 /pseudo
(329 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
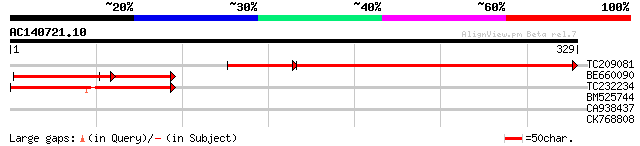
TC209081 similar to UP|Q8LD35 (Q8LD35) Nuclear matrix protein 1,... 295 5e-93
BE660090 homologue to GP|15077144|gb nuclear matrix protein 1 {L... 106 5e-33
TC232234 similar to UP|Q93XE2 (Q93XE2) Nuclear matrix protein 1,... 92 4e-19
BM525744 30 1.1
CA938437 28 7.4
CK768808 28 7.4
>TC209081 similar to UP|Q8LD35 (Q8LD35) Nuclear matrix protein 1, partial
(65%)
Length = 876
Score = 295 bits (756), Expect(2) = 5e-93
Identities = 148/163 (90%), Positives = 157/163 (95%)
Frame = +1
Query: 167 IP*VAELESKLTEQSKILLNLQQKVDDLASKHAYNPDEEYTEVESQLRAHLESFLETART 226
+P V+ELE K +EQSKILLNLQQKVDDLASKHAYNPDEEYTEVESQLRAHLESFLETART
Sbjct: 169 LPDVSELELKFSEQSKILLNLQQKVDDLASKHAYNPDEEYTEVESQLRAHLESFLETART 348
Query: 227 FNMIYTKEIRPWTHMMEVPQLHGFGPAANRLLEAYKMLLKFLGNLQNLRDSHAALAFGSS 286
FN+IYTKEIRPWTHMMEVPQLHGFGPAANRLLEAYKMLLKFLGNL+NLRDSHAALAFGSS
Sbjct: 349 FNLIYTKEIRPWTHMMEVPQLHGFGPAANRLLEAYKMLLKFLGNLRNLRDSHAALAFGSS 528
Query: 287 ETSGGPSSVSRIISECESEMTVINRDLGILSASIAREHGEKMS 329
ETS GPSSV+RIISECES +TV+NRDLGILSASIARE GEKM+
Sbjct: 529 ETSDGPSSVTRIISECESALTVLNRDLGILSASIAREQGEKMN 657
Score = 63.9 bits (154), Expect(2) = 5e-93
Identities = 33/41 (80%), Positives = 35/41 (84%)
Frame = +2
Query: 127 VLTSR*PRTYT**ILLQKNRHKYFLKNVNCFPQMFRFSPYI 167
VLTSR*PRT ** LQK++ KYFLKNVNCF QMFRFSPYI
Sbjct: 44 VLTSR*PRTSN**TPLQKSKLKYFLKNVNCFLQMFRFSPYI 166
>BE660090 homologue to GP|15077144|gb nuclear matrix protein 1 {Lycopersicon
esculentum}, partial (33%)
Length = 540
Score = 106 bits (264), Expect(2) = 5e-33
Identities = 52/59 (88%), Positives = 54/59 (91%)
Frame = +3
Query: 3 SKQMEVIQKKLGMLNYPRANASAQSLLFAGMERYALFEWLFFRLLGDKSPFSQQNLQGD 61
S+QME I+KKL LNYPRANA AQSLLFAGMERYAL EWLFFRLLGDKSPFSQQNLQGD
Sbjct: 153 SRQMEEIRKKLADLNYPRANAPAQSLLFAGMERYALLEWLFFRLLGDKSPFSQQNLQGD 329
Score = 52.8 bits (125), Expect(2) = 5e-33
Identities = 28/44 (63%), Positives = 31/44 (69%)
Frame = +2
Query: 53 FSQQNLQGDALDRDEETARIQYLAEIAKFLGITTTVDTDAIQTH 96
FS + +G + RIQYLAEIAKFLGITTTVDTDAIQ H
Sbjct: 305 FSTKPTRGWPMIVTRRLVRIQYLAEIAKFLGITTTVDTDAIQGH 436
>TC232234 similar to UP|Q93XE2 (Q93XE2) Nuclear matrix protein 1, partial
(21%)
Length = 580
Score = 91.7 bits (226), Expect = 4e-19
Identities = 56/99 (56%), Positives = 63/99 (63%), Gaps = 3/99 (3%)
Frame = +3
Query: 1 MASKQMEVIQKKLGMLNYPRANASAQSLLFAGMERYALFEWLF---FRLLGDKSPFSQQN 57
MAS+QME I+KKL LNYPRANA AQSLLFAGME F FRL+ PF +
Sbjct: 261 MASRQMEEIRKKLADLNYPRANAPAQSLLFAGMETLCPSXXAFLSAFRLI--XXPFLHKT 434
Query: 58 LQGDALDRDEETARIQYLAEIAKFLGITTTVDTDAIQTH 96
+G A+ DEET I YL +IAKFLG TTTV T A H
Sbjct: 435 XKGMAMIVDEETGXIPYLXKIAKFLGFTTTVXTXAXPGH 551
>BM525744
Length = 411
Score = 30.4 bits (67), Expect = 1.1
Identities = 14/53 (26%), Positives = 25/53 (46%)
Frame = +3
Query: 170 VAELESKLTEQSKILLNLQQKVDDLASKHAYNPDEEYTEVESQLRAHLESFLE 222
V ++ K+ E ILL L+ +D +H YN + Y + R ++ L+
Sbjct: 9 VIDMTKKMVEPRNILLTLKDHNNDTTIRHIYNARQAYRSSQKGPRTEMQHLLK 167
>CA938437
Length = 422
Score = 27.7 bits (60), Expect = 7.4
Identities = 10/36 (27%), Positives = 22/36 (60%)
Frame = +2
Query: 33 MERYALFEWLFFRLLGDKSPFSQQNLQGDALDRDEE 68
+E + F+W+FFR+ + + ++ +Q + R+EE
Sbjct: 17 LEMFRRFQWVFFRVENEWNKITRSGVQLTEIPREEE 124
>CK768808
Length = 641
Score = 27.7 bits (60), Expect = 7.4
Identities = 12/38 (31%), Positives = 22/38 (57%)
Frame = +1
Query: 170 VAELESKLTEQSKILLNLQQKVDDLASKHAYNPDEEYT 207
+ +L++ + + ++LLN+QQK+D L N E T
Sbjct: 430 ICQLKNSVQLRDQVLLNMQQKLDSLCELVVNNSKEHST 543
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.332 0.143 0.430
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,755,253
Number of Sequences: 63676
Number of extensions: 141845
Number of successful extensions: 728
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 725
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 727
length of query: 329
length of database: 12,639,632
effective HSP length: 98
effective length of query: 231
effective length of database: 6,399,384
effective search space: 1478257704
effective search space used: 1478257704
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 59 (27.3 bits)
Medicago: description of AC140721.10


