

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140670.15 - phase: 0
(682 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
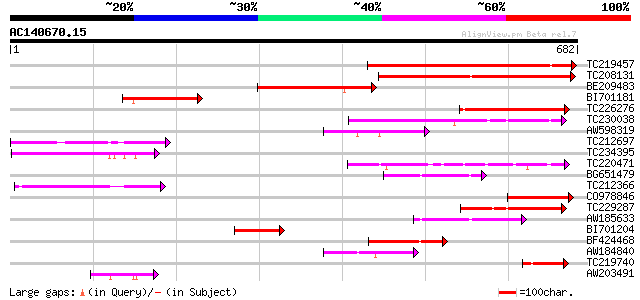
Score E
Sequences producing significant alignments: (bits) Value
TC219457 similar to GB|AAN46891.1|24111445|BT001092 At2g05250/F5... 451 e-127
TC208131 weakly similar to UP|Q6K2F0 (Q6K2F0) Heat shock protein... 282 4e-76
BE209483 165 7e-41
BI701181 similar to GP|15983799|gb| At2g05250/F5G3.15 {Arabidops... 156 2e-38
TC226276 weakly similar to UP|Q6K2F0 (Q6K2F0) Heat shock protein... 154 2e-37
TC230038 127 1e-29
AW598319 102 4e-22
TC212697 weakly similar to UP|O13633 (O13633) HLJ1 PROTEIN (SPBC... 102 4e-22
TC234395 similar to UP|Q9ZU99 (Q9ZU99) Expressed protein (At2g01... 99 6e-21
TC220471 98 1e-20
BG651479 95 1e-19
TC212366 similar to UP|Q9ZU99 (Q9ZU99) Expressed protein (At2g01... 91 2e-18
CO978846 88 1e-17
TC229287 similar to UP|Q8RYF9 (Q8RYF9) Heat shock protein-like, ... 86 4e-17
AW185633 82 1e-15
BI701204 82 1e-15
BF424468 80 2e-15
AW184840 75 1e-13
TC219740 similar to UP|Q6K2F0 (Q6K2F0) Heat shock protein-like, ... 60 3e-09
AW203491 similar to GP|9757996|dbj| DnaJ protein-like {Arabidops... 60 4e-09
>TC219457 similar to GB|AAN46891.1|24111445|BT001092 At2g05250/F5G3.15
{Arabidopsis thaliana;} , partial (31%)
Length = 1054
Score = 451 bits (1161), Expect = e-127
Identities = 214/251 (85%), Positives = 230/251 (91%)
Frame = +3
Query: 431 VSDNQLEHRKTVPVTITVPDPDFHDFDKDRSEPCFKPKQIWALYDEEDGMPRLYCLIREV 490
VS QLE+ KT P++ITVPD DFHDFDKDRSE CF+PKQIWALYDEEDGMPRLYC+IREV
Sbjct: 3 VSGLQLENGKTGPISITVPDSDFHDFDKDRSEECFRPKQIWALYDEEDGMPRLYCMIREV 182
Query: 491 VSVNPFKINISYLSSKTDSEFGPVNWLVSGFTKSCGNFRAMTSDVVDQVNIFSHVLSRVK 550
VSVNPFKI+ISYLSSKTDSEFG VNWL SGFTKSCGNFRA SD VDQVNIFSHVLS+ K
Sbjct: 183 VSVNPFKIHISYLSSKTDSEFGSVNWLDSGFTKSCGNFRAFNSDAVDQVNIFSHVLSKEK 362
Query: 551 AGRGGCVRIYPKCGDVWAVYRNWSTDWNRSTPDEVRHQYDMVEVLDDYSEELGLCVSPLI 610
AGRGGCVRIYP+ GD+WAVYRNWS DWNRSTPDEVRHQY+MVEVLDDYSEELG+CVSPLI
Sbjct: 363 AGRGGCVRIYPRSGDIWAVYRNWSPDWNRSTPDEVRHQYEMVEVLDDYSEELGVCVSPLI 542
Query: 611 KLDGFKTVYKRNADKSAIRYIPRREMLRFSHQVPSWLLKGEEASNLPDKCWDLDPAATPD 670
KL GFKTVY+ N DKS I++IPRREMLRFSHQVPSWLLKG EASNLP++CWDLDPAATPD
Sbjct: 543 KLAGFKTVYQSNTDKSTIKWIPRREMLRFSHQVPSWLLKG-EASNLPERCWDLDPAATPD 719
Query: 671 ELLHAAIEANA 681
ELLHAA E NA
Sbjct: 720 ELLHAATEPNA 752
>TC208131 weakly similar to UP|Q6K2F0 (Q6K2F0) Heat shock protein-like,
partial (32%)
Length = 1371
Score = 282 bits (721), Expect = 4e-76
Identities = 131/237 (55%), Positives = 175/237 (73%)
Frame = +3
Query: 444 VTITVPDPDFHDFDKDRSEPCFKPKQIWALYDEEDGMPRLYCLIREVVSVNPFKINISYL 503
VTI VPDPDFH+FD DR E F Q+WA YD++DGMPR Y I +V+S+ PFK+ IS+L
Sbjct: 117 VTINVPDPDFHNFDLDRDENSFAEDQVWAAYDDDDGMPRYYAKIHKVISMKPFKMRISWL 296
Query: 504 SSKTDSEFGPVNWLVSGFTKSCGNFRAMTSDVVDQVNIFSHVLSRVKAGRGGCVRIYPKC 563
+S+++SE GP++W+ SGF K+CG+FR ++ + +N FSH + K RG VRI+P
Sbjct: 297 NSRSNSELGPIDWVGSGFYKTCGDFRTGKHEITESLNSFSHKVRWTKGTRG-VVRIFPGK 473
Query: 564 GDVWAVYRNWSTDWNRSTPDEVRHQYDMVEVLDDYSEELGLCVSPLIKLDGFKTVYKRNA 623
G+VWA+YRNWS DWN TPDEV H+YDMVEVL+D+ EE G+ V+PL+K+ GF+TV++R+
Sbjct: 474 GEVWALYRNWSPDWNEHTPDEVIHKYDMVEVLEDFDEEQGILVTPLVKVAGFRTVFQRHM 653
Query: 624 DKSAIRYIPRREMLRFSHQVPSWLLKGEEASNLPDKCWDLDPAATPDELLHAAIEAN 680
D R I + EM +FSHQVP++LL G+EA N P C +LDPAATP +LL A EAN
Sbjct: 654 DCDQERRILKEEMFQFSHQVPNYLLTGQEADNAPKGCRELDPAATPLDLLQIATEAN 824
>BE209483
Length = 447
Score = 165 bits (417), Expect = 7e-41
Identities = 96/148 (64%), Positives = 114/148 (76%), Gaps = 5/148 (3%)
Frame = +1
Query: 299 SSDVSFSRNEPQEVKPSRPEKKRKVL-GASLRNVHEGKGSKCASELALANGNGSVGHGQR 357
+SDV FS +EPQE K SRP+KK+KV+ GAS RN ++ KGSK ASE +ANGN S+GHGQ+
Sbjct: 4 NSDVPFSCSEPQEDKLSRPDKKQKVVVGASFRNGYDEKGSKRASESIVANGNDSMGHGQK 183
Query: 358 ISSTSEIPTKQYSMAPAFDARKLLIEKARTEIRKKLEEMKLASE----TAAAVIEGKKSQ 413
S T E+ TKQ SMAPAFDARKLLIEKAR EIRKKLEEM+L+SE AAA+ E +KSQ
Sbjct: 184 PSCTVEVQTKQCSMAPAFDARKLLIEKARKEIRKKLEEMRLSSEAAATAAAALNEKEKSQ 363
Query: 414 ADVGQVKGDICTKTALNVSDNQLEHRKT 441
A+VGQVK + C K A VS QLE+ KT
Sbjct: 364 AEVGQVKRETCRKAAPIVSGLQLENGKT 447
>BI701181 similar to GP|15983799|gb| At2g05250/F5G3.15 {Arabidopsis
thaliana}, partial (7%)
Length = 341
Score = 156 bits (395), Expect = 2e-38
Identities = 74/112 (66%), Positives = 82/112 (73%), Gaps = 16/112 (14%)
Frame = +1
Query: 136 NHKGLSSVHASG--------------GNQDTFWTICTSCKVQYEYLRKYVNKKLSCKNCR 181
N LS HA+G G DTFWTICTSCKVQYEYLRKYVNK+LSCKNCR
Sbjct: 4 NQTNLSPAHATGAAGYNKCSNLSTPCGGLDTFWTICTSCKVQYEYLRKYVNKRLSCKNCR 183
Query: 182 GIFIALET--APANGSFPYSPWSYGSSSGYGSHSYDGVTYVPTNGAYFNGNG 231
G F+A+ET APANGSFPY PWSY + +GYGSHS+DGV YVPT+ YFNGNG
Sbjct: 184 GTFVAVETGAAPANGSFPYCPWSYVAGNGYGSHSFDGVAYVPTSAPYFNGNG 339
>TC226276 weakly similar to UP|Q6K2F0 (Q6K2F0) Heat shock protein-like,
partial (19%)
Length = 992
Score = 154 bits (388), Expect = 2e-37
Identities = 74/132 (56%), Positives = 94/132 (71%)
Frame = +3
Query: 542 FSHVLSRVKAGRGGCVRIYPKCGDVWAVYRNWSTDWNRSTPDEVRHQYDMVEVLDDYSEE 601
FSH + R + G G + IYP+ GDVWA+YRNWS DWN T DEV H++D+VEVL+D+ E
Sbjct: 6 FSHKV-RWRTGAEGAICIYPRKGDVWAIYRNWSPDWNELTADEVIHKFDVVEVLEDFIEG 182
Query: 602 LGLCVSPLIKLDGFKTVYKRNADKSAIRYIPRREMLRFSHQVPSWLLKGEEASNLPDKCW 661
G+ V PL+K+ GF+TV+ + D IR IPR EM RFSHQ+PS++L G+EA P C
Sbjct: 183 HGIDVIPLVKVAGFRTVFHHHLDPKEIRIIPREEMFRFSHQIPSYVLTGQEAPEAPKGCR 362
Query: 662 DLDPAATPDELL 673
LDPAATP ELL
Sbjct: 363 VLDPAATPFELL 398
>TC230038
Length = 1229
Score = 127 bits (320), Expect = 1e-29
Identities = 89/271 (32%), Positives = 142/271 (51%), Gaps = 9/271 (3%)
Frame = +3
Query: 408 EGKKSQADVGQVKGDICT--KTALNVSDNQLEHRKTVPVTITVPDPDFHDFDKDRSEPCF 465
EG S V + ++ T K +++ SDN ++ P I VPD F DFD R+ F
Sbjct: 165 EGDASIPKVNLERSNLATENKDSVDDSDNCCAPPESSPEAINVPDTQFFDFDGGRALEKF 344
Query: 466 KPKQIWALYDEEDGMPRLYCLIREVVSVNPFKINISYLSSKTDSEFGPVNWLVSGFTKSC 525
+ QIWA Y +EDG+P+ Y I+++ + ++++ +L+ E + W SC
Sbjct: 345 QIGQIWAFYSDEDGLPKYYGQIKKIETSPDLELHVYWLTCCWLPE-NTIKWEDKDILISC 521
Query: 526 GNFRAMTS----DVVDQVNIFSHVLSRVKAGRGGCVRIYPKCGDVWAVYRNWSTDWNRST 581
G F+ + V + SH + G+ I+P+ GDVWA+YR W+ N+
Sbjct: 522 GRFKVNETHDFLSVYSTTSCVSHQVHADAVGKNKNYAIFPRKGDVWALYRKWT---NKMK 692
Query: 582 PDEVRH-QYDMVEVLDDYSEELGLCVSPLIKLDGFKTVY--KRNADKSAIRYIPRREMLR 638
E+ + +YD+VEV+++ +L + V L + G+ +V+ K N S IPR+E+LR
Sbjct: 693 CFEMENCEYDIVEVVEE--TDLFINVLVLEFVSGYTSVFRGKSNEGSSVNLRIPRKELLR 866
Query: 639 FSHQVPSWLLKGEEASNLPDKCWDLDPAATP 669
FSHQ+P++ L EE NL W+LDP A P
Sbjct: 867 FSHQIPAFKLT-EEHGNLKG-FWELDPGALP 953
>AW598319
Length = 440
Score = 102 bits (255), Expect = 4e-22
Identities = 55/141 (39%), Positives = 83/141 (58%), Gaps = 13/141 (9%)
Frame = +3
Query: 378 RKLLIEKARTEIRKKLEEMKLASETAAAVIEGKKSQADV---GQVKGDICTKTALNVSDN 434
+ +L+EKAR EI KKL+E K AS + + + K + ++ G+ + K V D+
Sbjct: 21 KNILVEKARKEIVKKLDEWK-ASSASNNLDKSKNTDTEIREKGKEREGNVVKPGAQVVDS 197
Query: 435 QLEHRKTVP----------VTITVPDPDFHDFDKDRSEPCFKPKQIWALYDEEDGMPRLY 484
+ ++K +++ VPDPDFHDFD DR+E F Q+WA YD +DGMPR Y
Sbjct: 198 ETVNKKCFSADLEPELPGSLSMNVPDPDFHDFDGDRTENAFGENQVWAAYDNDDGMPRYY 377
Query: 485 CLIREVVSVNPFKINISYLSS 505
CLI +V+S NP + IS+L++
Sbjct: 378 CLIHDVISKNPLNMRISWLNA 440
>TC212697 weakly similar to UP|O13633 (O13633) HLJ1 PROTEIN (SPBC17A3.05c
protein) (Pi041 protein), partial (10%)
Length = 741
Score = 102 bits (255), Expect = 4e-22
Identities = 61/194 (31%), Positives = 99/194 (50%), Gaps = 1/194 (0%)
Frame = -3
Query: 1 MEAKKEEALKAIENAEKRF-SHRDFVGAKSYALKAKTLCPGLEGISQLVTTFEVYIASQV 59
M + EAL+ AE +F + + A YA +A LCP L G+ + V V A
Sbjct: 601 MAEAESEALRLKAMAESKFKASNNAKSALKYANRAHRLCPHLAGVPETVAALSVLAAP-- 428
Query: 60 TCNGELDWYSIMGLNPSTNIEAVKKQYKKMAGLLHPDNNKCVGADGAFHLVSEAWARLSG 119
DWY +G P + +++QYKK+A LLHPD N V ++ AF L+ EA+ LS
Sbjct: 427 ------DWYRALGAEPFASSSVIRRQYKKLALLLHPDKNPHVASEEAFKLLGEAFRFLS- 269
Query: 120 SYDMKRNAQLGAGNGVNHKGLSSVHASGGNQDTFWTICTSCKVQYEYLRKYVNKKLSCKN 179
D R + A + + A+ +TFWT C++C++ +++ R+Y+ ++L C +
Sbjct: 268 --DRNRRREYDA------ELRRKIEAAESESETFWTACSTCRLLHQFERRYLGQELVCPS 113
Query: 180 CRGIFIALETAPAN 193
C F A+E ++
Sbjct: 112 CEKGFRAVEAVQSD 71
>TC234395 similar to UP|Q9ZU99 (Q9ZU99) Expressed protein
(At2g01710/T8O11.12), partial (48%)
Length = 760
Score = 99.0 bits (245), Expect = 6e-21
Identities = 68/219 (31%), Positives = 102/219 (46%), Gaps = 41/219 (18%)
Frame = +3
Query: 3 AKKEEALKAIENAEKRFSHRDFVGAKSYALKAKTLCPGLEGISQLVTTFEVYIASQVTCN 62
A + EA + + AEK +RD VG++ +A A+ P LEG Q++ +V +A+ N
Sbjct: 42 ATRAEAERLLGIAEKLLQNRDLVGSREFAFLAQETEPLLEGSDQILAIVDVLLAADKRVN 221
Query: 63 GELDWYSIMGLNP-STNIEAVKKQYKKMAGLLHPDNNKCVGADGAFHLVSEAWARLS--- 118
DWY+++ ++ S +++ +KKQY+++A LLHPD ++ AD AF LV++AWA LS
Sbjct: 222 NHPDWYAVLQVDRRSDDLDLIKKQYRRLALLLHPDKSRFHFADHAFQLVADAWALLSDPI 401
Query: 119 --GSYDMK-----------------------RNAQLGAGNGVN--------HKGLSSVHA 145
YD + R G G G +HA
Sbjct: 402 KKSVYDKELSFFSRVDLSVPGWVQQQEKLPVRRTGPGPGPGPGPTAGRNSAASAREDIHA 581
Query: 146 SGGNQ----DTFWTICTSCKVQYEYLRKYVNKKLSCKNC 180
++ TFWT C C YEY R Y L C+NC
Sbjct: 582 DENSRRRRSSTFWTACPYCYRLYEYPRVYEGCCLRCQNC 698
>TC220471
Length = 962
Score = 98.2 bits (243), Expect = 1e-20
Identities = 77/277 (27%), Positives = 142/277 (50%), Gaps = 10/277 (3%)
Frame = +3
Query: 407 IEGKKSQADVGQ--VKGDICTKTALNVSDNQLEHRKTVPVTITVPDP---DFHDFDKDRS 461
++ +KS D+ + +GD TA +DN + K V V+ +P + F K++S
Sbjct: 126 LKARKSPRDLSKKNAQGDAGEWTAGKKTDNHSSNSKNVKVS-NIPQSVGASCYGFKKEKS 302
Query: 462 EPCFKPKQIWALYDEEDGMPRLYCLIREVVSVNPFKINISYLSSKTDSEFGPVNWLVSGF 521
E F+ QIWA+Y + D MP Y IR + F++ + L P N L
Sbjct: 303 EEMFQCGQIWAIYGDRDHMPDTYAQIRMIECTPNFRLQVYML-----EPCPPPNDLKR-- 461
Query: 522 TKSCGNFRAMTSDV-VDQVNIFSHVLSRVKAGRGGCVRIYPKCGDVWAVYRNWSTDWNRS 580
T SCG F + + + ++ FSH L + + IYP+ G++WA+Y++ ++ ++
Sbjct: 462 TISCGTFSVKEAKLRMLSLSAFSHQL-KAELVANNRYEIYPRKGEIWALYKD--QNYEQT 632
Query: 581 TPDEVRHQYDMVEVLDDYSEELGLCVSPLIKLDGFKTVYKR---NADKSAIRYIPRREML 637
+ ++ R + +VEVL D ++ + + V L+ +T++K K+ + I R+E+
Sbjct: 633 SSNQGRGECHIVEVLADNNKSIQVVV--LVPHGNSQTIFKAPRIQRSKTGVIEILRKEVG 806
Query: 638 RFSHQVPSWLLKGEEASNLPDK-CWDLDPAATPDELL 673
RFSHQ+P++ + + N+ + CW+LDP++ P +
Sbjct: 807 RFSHQIPAF----QHSDNVHLRGCWELDPSSVPGSFI 905
>BG651479
Length = 392
Score = 94.7 bits (234), Expect = 1e-19
Identities = 44/124 (35%), Positives = 72/124 (57%)
Frame = +1
Query: 450 DPDFHDFDKDRSEPCFKPKQIWALYDEEDGMPRLYCLIREVVSVNPFKINISYLSSKTDS 509
D +F DFDK +++ CF QIWA+YD +GMPR Y LIR+V+S F++ I + D
Sbjct: 19 DAEFSDFDKGKNKECFTAGQIWAIYDTSEGMPRFYALIRKVLSPG-FRLQIIWFEPHPDC 195
Query: 510 EFGPVNWLVSGFTKSCGNFRAMTSDVVDQVNIFSHVLSRVKAGRGGCVRIYPKCGDVWAV 569
+ +NW+ +CG ++ D+ + +FSH + K R ++YP+ G+ WA+
Sbjct: 196 K-DEINWVNEEMPVACGKYKLSDIDITEDHLMFSHPVLCEKISR-NTFKVYPRKGETWAL 369
Query: 570 YRNW 573
++NW
Sbjct: 370 FKNW 381
>TC212366 similar to UP|Q9ZU99 (Q9ZU99) Expressed protein
(At2g01710/T8O11.12), partial (50%)
Length = 701
Score = 90.9 bits (224), Expect = 2e-18
Identities = 63/184 (34%), Positives = 91/184 (49%), Gaps = 3/184 (1%)
Frame = +1
Query: 7 EALKAIENAEKRFSHRDFVGAKSYALKAKTLCPGLEGISQLVTTFEVYIASQVTC-NGEL 65
E L AI EK RD ++ +A+ A+ P LEG Q++ EV +A++ N L
Sbjct: 4 ERLLAI--GEKLLQSRDLSSSRDFAILAQEAEPLLEGSDQILAIVEVLLAAEKPITNDHL 177
Query: 66 DWYSIMGLNPST-NIEAVKKQYKKMAGLLHPDNNKCVGADGAFHLVSEAWARLSGSYDMK 124
DWY+I+ ++ + +++ +KKQY+++ LLHPD N AD AF LVS+AWA LS
Sbjct: 178 DWYAILQVDRTCQDLDLIKKQYRRLGLLLHPDKNPFSLADHAFKLVSDAWAVLSDPV--- 348
Query: 125 RNAQLGAGNGVNHKGLSSVHASGGNQ-DTFWTICTSCKVQYEYLRKYVNKKLSCKNCRGI 183
K + +G Q ++FWT C C YEY L C+NC
Sbjct: 349 ------------QKAIYDRDVAGSVQPESFWTACPYCYFLYEYPAVCEGCCLRCQNCERS 492
Query: 184 FIAL 187
F L
Sbjct: 493 FHGL 504
>CO978846
Length = 823
Score = 87.8 bits (216), Expect = 1e-17
Identities = 42/79 (53%), Positives = 54/79 (68%)
Frame = -3
Query: 600 EELGLCVSPLIKLDGFKTVYKRNADKSAIRYIPRREMLRFSHQVPSWLLKGEEASNLPDK 659
EE G+ ++PL+K+ GFKTV+++NAD ++ I + EM RFSHQVPS L G E N P
Sbjct: 821 EEKGVNIAPLVKVSGFKTVFRQNADPRKVKNISKAEMFRFSHQVPSHWLTGVEGHNAPKG 642
Query: 660 CWDLDPAATPDELLHAAIE 678
C +LDPAATP ELL E
Sbjct: 641 CLELDPAATPMELLQVLAE 585
>TC229287 similar to UP|Q8RYF9 (Q8RYF9) Heat shock protein-like, partial (4%)
Length = 817
Score = 86.3 bits (212), Expect = 4e-17
Identities = 49/129 (37%), Positives = 79/129 (60%), Gaps = 2/129 (1%)
Frame = +2
Query: 543 SHVLSRVKAGRGGCVRIYPKCGDVWAVYRNWSTDWNRSTPDEVRHQYDMVEVLDDYSEEL 602
SH + + G+ I+P+ G++WA+YRNW+T RS D + +YD+VEV+ + ++L
Sbjct: 161 SHQVQVINDGKKKEYEIFPRKGEIWALYRNWTTKIKRS--DLLNLEYDIVEVVGE--QDL 328
Query: 603 GLCVSPLIKLDGFKTVYKR--NADKSAIRYIPRREMLRFSHQVPSWLLKGEEASNLPDKC 660
+ V PL + G+ +V+KR NA + I +++LRFSHQ+P++ L E+ NL
Sbjct: 329 WMDVLPLELVSGYNSVFKRKSNAGSARATKIYWKDLLRFSHQIPAFELTEEQDGNLRG-F 505
Query: 661 WDLDPAATP 669
W+LDP A P
Sbjct: 506 WELDPGAVP 532
>AW185633
Length = 443
Score = 81.6 bits (200), Expect = 1e-15
Identities = 43/137 (31%), Positives = 74/137 (53%), Gaps = 1/137 (0%)
Frame = +1
Query: 486 LIREVVSVNPFKINISYLSSKTDSEFGPVNWLVSGFTKSCGNFRAMTSDVVDQVNIFSHV 545
+IR+V S FK+ I++ D + V+W+ +CG + +D + +FSH+
Sbjct: 4 VIRKVFSPG-FKLRITWFEPDPDEQ-DQVHWVEEELPIACGKHKLGITDTTEDRLMFSHL 177
Query: 546 LSRVKAGRGGCV-RIYPKCGDVWAVYRNWSTDWNRSTPDEVRHQYDMVEVLDDYSEELGL 604
+ K GR C ++YP+ G+ WA+++NW W+ + ++ VE+L DY E +G+
Sbjct: 178 IVCEKIGR--CTYKVYPRKGETWALFKNWDIKWHMDAESHREYDFEFVEILSDYVEGVGV 351
Query: 605 CVSPLIKLDGFKTVYKR 621
VS L KL GF ++ R
Sbjct: 352 VVSYLAKLKGFVCLFSR 402
>BI701204
Length = 422
Score = 81.6 bits (200), Expect = 1e-15
Identities = 41/61 (67%), Positives = 50/61 (81%), Gaps = 1/61 (1%)
Frame = +2
Query: 271 NGKAKRGRPKVKSGADRKHCLTETVVNISSDVSFSRNEPQEVKPSRPEKKRK-VLGASLR 329
NG KRGRPKVKSGAD++H + ET+VN +SDV FS +EPQE K SRP+KK+K V+GAS R
Sbjct: 5 NGNVKRGRPKVKSGADKRHHMVETMVNTNSDVPFSCSEPQEDKLSRPDKKQKVVVGASFR 184
Query: 330 N 330
N
Sbjct: 185 N 187
>BF424468
Length = 394
Score = 80.5 bits (197), Expect = 2e-15
Identities = 38/97 (39%), Positives = 59/97 (60%), Gaps = 2/97 (2%)
Frame = +1
Query: 432 SDNQLEHRKTVPVT--ITVPDPDFHDFDKDRSEPCFKPKQIWALYDEEDGMPRLYCLIRE 489
+DN + P T I PDPDF DF++D++E CF Q+WA++D D MPR Y L+++
Sbjct: 91 ADNCSPLKSNFPPTSEICCPDPDFSDFERDKAEDCFAVNQLWAIFDNTDSMPRFYALVKK 270
Query: 490 VVSVNPFKINISYLSSKTDSEFGPVNWLVSGFTKSCG 526
V S PFK+ I++L +D + G ++W + +CG
Sbjct: 271 VYS--PFKLRITWLEPDSDDQ-GEIDWHEASLPVACG 372
>AW184840
Length = 465
Score = 74.7 bits (182), Expect = 1e-13
Identities = 46/138 (33%), Positives = 70/138 (50%), Gaps = 24/138 (17%)
Frame = +1
Query: 378 RKLLIEKARTEIRKKLEEMKLASETAAAVIEGKKSQADVGQVKGDICTKTALNVSDNQLE 437
+ LL+EKAR EI KL +++ + A+ E +V + KG+ C++ + + + +E
Sbjct: 55 KNLLMEKARKEISNKLRQVQSNAVDKTAMKENGNDFQEVSE-KGEKCSRNSEMCAQDNIE 231
Query: 438 H-----------------------RKTVPVT-ITVPDPDFHDFDKDRSEPCFKPKQIWAL 473
RK + T + V PDFHDF KDR+E F Q+WA+
Sbjct: 232 KSEDRKSGSRAIKPFAGSTIAKVSRKFLETTPVDVLYPDFHDFCKDRTEGSFGENQVWAV 411
Query: 474 YDEEDGMPRLYCLIREVV 491
YD +DGMPR Y LIR ++
Sbjct: 412 YDNDDGMPRCYVLIRRII 465
>TC219740 similar to UP|Q6K2F0 (Q6K2F0) Heat shock protein-like, partial (6%)
Length = 594
Score = 60.1 bits (144), Expect = 3e-09
Identities = 30/56 (53%), Positives = 40/56 (70%)
Frame = +1
Query: 617 TVYKRNADKSAIRYIPRREMLRFSHQVPSWLLKGEEASNLPDKCWDLDPAATPDEL 672
TV+ R++ R IP+ E+ RFSHQVP++LL G+EA N P C +LDPAATP +L
Sbjct: 1 TVFHRHSHDQG-RKIPKVEIFRFSHQVPNYLLTGQEAHNAPKGCRELDPAATPLDL 165
>AW203491 similar to GP|9757996|dbj| DnaJ protein-like {Arabidopsis
thaliana}, partial (7%)
Length = 367
Score = 59.7 bits (143), Expect = 4e-09
Identities = 42/121 (34%), Positives = 50/121 (40%), Gaps = 39/121 (32%)
Frame = +1
Query: 98 NKCVGADGAFHLVSEAWARLSG-----SYDMKRNAQLGAGNGVNHKGLSSVHASG----- 147
+K GA+GAF LVSEAW+ LS +Y+ R + N N G S S
Sbjct: 4 SKSPGAEGAFKLVSEAWSLLSDKVKRLAYNQNRRLEGFQHNAPNLVGTQSKAPSSNGYKK 183
Query: 148 -----------GNQD------------------TFWTICTSCKVQYEYLRKYVNKKLSCK 178
GN D TFWTIC CK YEYLR Y+N+ L C
Sbjct: 184 HNKNATSSIRTGNNDARAHPHPPSIPPPHTNVGTFWTICNKCKTHYEYLRTYLNQTLLCP 363
Query: 179 N 179
N
Sbjct: 364 N 366
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.132 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,443,267
Number of Sequences: 63676
Number of extensions: 472895
Number of successful extensions: 1761
Number of sequences better than 10.0: 115
Number of HSP's better than 10.0 without gapping: 1710
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1739
length of query: 682
length of database: 12,639,632
effective HSP length: 104
effective length of query: 578
effective length of database: 6,017,328
effective search space: 3478015584
effective search space used: 3478015584
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 62 (28.5 bits)
Medicago: description of AC140670.15


