

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140670.14 + phase: 0
(359 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
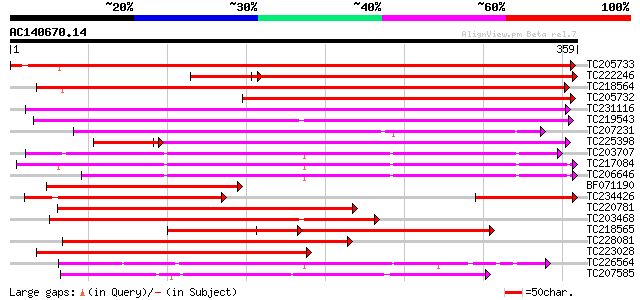
Score E
Sequences producing significant alignments: (bits) Value
TC205733 548 e-156
TC222246 373 e-122
TC218564 419 e-117
TC205732 333 5e-92
TC231116 similar to UP|Q9FFC6 (Q9FFC6) GDSL-motif lipase/hydrola... 293 8e-80
TC219543 weakly similar to UP|Q94CH8 (Q94CH8) Family II lipase E... 286 1e-77
TC207231 similar to UP|Q94CH6 (Q94CH6) Family II lipase EXL3, pa... 253 1e-67
TC225398 similar to UP|O04320 (O04320) Proline-rich protein APG ... 215 7e-66
TC203707 226 2e-59
TC217084 222 2e-58
TC206646 201 6e-52
BF071190 199 2e-51
TC234426 191 6e-49
TC220781 weakly similar to UP|Q94CH6 (Q94CH6) Family II lipase E... 184 5e-47
TC203468 182 2e-46
TC218565 168 4e-42
TC228081 similar to UP|Q9FFC6 (Q9FFC6) GDSL-motif lipase/hydrola... 166 2e-41
TC223028 weakly similar to UP|Q94CH7 (Q94CH7) Family II lipase E... 166 2e-41
TC226564 159 2e-39
TC207585 similar to UP|Q9AYM7 (Q9AYM7) CPRD47 protein, partial (... 152 3e-37
>TC205733
Length = 1377
Score = 548 bits (1413), Expect = e-156
Identities = 265/360 (73%), Positives = 305/360 (84%), Gaps = 2/360 (0%)
Frame = +2
Query: 1 MKGGQRKNLLYTNNILCILLLQRLDLSLVK--SAKVPAIIVFGDSSVDAGNNNFISTVAR 58
M+ GQRK T +LC ++ L LSLV SAKV A+IVFGDSSVDAGNNNFI T+AR
Sbjct: 56 MEEGQRKT---TTLLLCSHIVVLLLLSLVAETSAKVSAVIVFGDSSVDAGNNNFIPTIAR 226
Query: 59 SNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFAS 118
SNFQPYGRDF GGK TGRF NGRI TDFISE+FG+KPY+PAYLDP +NIS FA+GV+FAS
Sbjct: 227 SNFQPYGRDFEGGKATGRFCNGRIPTDFISESFGLKPYVPAYLDPKYNISDFASGVTFAS 406
Query: 119 AATGYDNATSDVLSVIPLWKQLEYYKEYQKKLGAYLGEKKAKETITKALYIISLGTNDFL 178
AATGYDNATSDVLSVIPLWKQLEYYK YQK L AYLGE KAK+TI +AL+++SLGTNDFL
Sbjct: 407 AATGYDNATSDVLSVIPLWKQLEYYKGYQKNLSAYLGESKAKDTIAEALHLMSLGTNDFL 586
Query: 179 ENYYTIPGRASQYTPSEYQNFLAGIAQNFIHKLYDLGAKKISLGGLPPMGCLPLERTTNF 238
ENYYT+PGRASQ+TP +YQNFLAGIA+NFI LY LGA+K+SLGGLPPMGCLPLERTT+
Sbjct: 587 ENYYTMPGRASQFTPQQYQNFLAGIAENFIRSLYGLGARKVSLGGLPPMGCLPLERTTSI 766
Query: 239 AGGNDCVSNYNNIALEFNGKLNKLTTKLKKDLPGIRLVFSNPYDVLLGVVKKPGQYGFQV 298
AGGNDCV+ YNNIALEFN +L LT KL ++LPG++LVFSNPY ++L ++K+P YGF+
Sbjct: 767 AGGNDCVARYNNIALEFNNRLKNLTIKLNQELPGLKLVFSNPYYIMLSIIKRPQLYGFES 946
Query: 299 ASMACCATGMFEMGYACSRASLFSCMDASRYVFWDSFHPTEKTNGIVANYLVKNALAQFL 358
S+ACCATGMFEMGYACSR +FSC DAS+YVFWDSFHPTE TN IVA Y+V L QFL
Sbjct: 947 TSVACCATGMFEMGYACSRGQMFSCTDASKYVFWDSFHPTEMTNSIVAKYVVLRVLYQFL 1126
>TC222246
Length = 781
Score = 373 bits (957), Expect(2) = e-122
Identities = 175/206 (84%), Positives = 191/206 (91%)
Frame = +2
Query: 154 LGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIAQNFIHKLYD 213
L + +A ET+ KAL+IISLGTNDFLENY+ IPGRASQYTP EYQNFLAGIA+NFI+KLY
Sbjct: 125 LEKSRANETVAKALHIISLGTNDFLENYFAIPGRASQYTPREYQNFLAGIAENFIYKLYG 304
Query: 214 LGAKKISLGGLPPMGCLPLERTTNFAGGNDCVSNYNNIALEFNGKLNKLTTKLKKDLPGI 273
LGA+KISLGGLPPMGCLPLERTTNF GGN+CVSNYNNIALEFN L+KLTTKLKKDLPGI
Sbjct: 305 LGARKISLGGLPPMGCLPLERTTNFVGGNECVSNYNNIALEFNDNLSKLTTKLKKDLPGI 484
Query: 274 RLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDASRYVFWD 333
RLVFSNPYD+LL ++K+P QYGFQV SMACCATGMFEMGYACSRAS FSC+DASRYVFWD
Sbjct: 485 RLVFSNPYDILLQIIKRPAQYGFQVTSMACCATGMFEMGYACSRASSFSCIDASRYVFWD 664
Query: 334 SFHPTEKTNGIVANYLVKNALAQFLH 359
SFHPTEKTNGI+A YLVKNALAQFLH
Sbjct: 665 SFHPTEKTNGIIAKYLVKNALAQFLH 742
Score = 84.3 bits (207), Expect(2) = e-122
Identities = 41/46 (89%), Positives = 41/46 (89%)
Frame = +1
Query: 115 SFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKLGAYLGEKKAK 160
SFASAATGYDNATSDVLSVIPLWKQLEYYK YQKKL YLGEK K
Sbjct: 7 SFASAATGYDNATSDVLSVIPLWKQLEYYKGYQKKLSVYLGEK*GK 144
>TC218564
Length = 1398
Score = 419 bits (1076), Expect = e-117
Identities = 197/340 (57%), Positives = 255/340 (74%), Gaps = 3/340 (0%)
Frame = +1
Query: 18 ILLLQRLDLSLVKSA---KVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPT 74
+LL Q L +++ S VPA+IVFGDSSVD+GNNN I+TV +SNF+PYGRDF G +PT
Sbjct: 28 LLLTQSLLVAVTTSEAKNNVPAVIVFGDSSVDSGNNNVIATVLKSNFKPYGRDFEGDRPT 207
Query: 75 GRFSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVI 134
GRF NGR+ DFI+EAFGIK +PAYLDP++ I FATGV FASA TGYDNATS VL+VI
Sbjct: 208 GRFCNGRVPPDFIAEAFGIKRTVPAYLDPAYTIQDFATGVCFASAGTGYDNATSAVLNVI 387
Query: 135 PLWKQLEYYKEYQKKLGAYLGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPS 194
PLWK++EYYKEYQ KL +LG +KA + I++ALY++SLGTNDFLENYY P R +T S
Sbjct: 388 PLWKEIEYYKEYQAKLRTHLGVEKANKIISEALYLMSLGTNDFLENYYVFPTRRLHFTVS 567
Query: 195 EYQNFLAGIAQNFIHKLYDLGAKKISLGGLPPMGCLPLERTTNFAGGNDCVSNYNNIALE 254
+YQ+FL IA+NF+ +LY LG +K+S+ GL P+GCLPLER TN G + C YN++AL
Sbjct: 568 QYQDFLLRIAENFVRELYALGVRKLSITGLGPVGCLPLERATNILGDHGCNQEYNDVALS 747
Query: 255 FNGKLNKLTTKLKKDLPGIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYA 314
FN KL + TKL ++LP ++ + +N Y ++ ++ KP YGF+V ACC+TG FEM Y
Sbjct: 748 FNRKLENVITKLNRELPRLKALSANAYSIVNDIITKPSTYGFEVVEKACCSTGTFEMSYL 927
Query: 315 CSRASLFSCMDASRYVFWDSFHPTEKTNGIVANYLVKNAL 354
CS + +C DA +YVFWD+FHPTEKTN IV++YL+ L
Sbjct: 928 CSDKNPLTCTDAEKYVFWDAFHPTEKTNRIVSSYLIPKLL 1047
>TC205732
Length = 974
Score = 333 bits (855), Expect = 5e-92
Identities = 154/211 (72%), Positives = 178/211 (83%)
Frame = +3
Query: 148 KKLGAYLGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIAQNF 207
K L AYLGE KAKET+ +AL+++SLGTNDFLENYYT+PGRASQYTP +YQ FLAGIA+NF
Sbjct: 3 KNLSAYLGESKAKETVAEALHLMSLGTNDFLENYYTMPGRASQYTPQQYQIFLAGIAENF 182
Query: 208 IHKLYDLGAKKISLGGLPPMGCLPLERTTNFAGGNDCVSNYNNIALEFNGKLNKLTTKLK 267
I LY LGA+KISLGGLPPMGCLPLERTTN GGNDCV+ YNNIALEFN KL LT KL
Sbjct: 183 IRSLYGLGARKISLGGLPPMGCLPLERTTNIVGGNDCVARYNNIALEFNDKLKNLTIKLN 362
Query: 268 KDLPGIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDAS 327
++LPG++LVFSNPY ++L ++K+P YGF+ S+ACCATGMFEMGYACSR +FSC DAS
Sbjct: 363 QELPGLKLVFSNPYYIMLNIIKRPQLYGFESTSVACCATGMFEMGYACSRGQMFSCTDAS 542
Query: 328 RYVFWDSFHPTEKTNGIVANYLVKNALAQFL 358
+YVFWDSFHPTE TN IVA Y+V L QFL
Sbjct: 543 KYVFWDSFHPTEMTNSIVAKYVVLRVLYQFL 635
>TC231116 similar to UP|Q9FFC6 (Q9FFC6) GDSL-motif lipase/hydrolase-like
protein, partial (94%)
Length = 1541
Score = 293 bits (750), Expect = 8e-80
Identities = 153/345 (44%), Positives = 206/345 (59%)
Frame = +1
Query: 11 YTNNILCILLLQRLDLSLVKSAKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLG 70
Y+ + L LL L VPAI FGDS VD GNNN T+ ++NF PYGRDF
Sbjct: 370 YSRSFLASFLLAVLLNVTNGQPLVPAIFTFGDSIVDVGNNNHQLTIVKANFPPYGRDFEN 549
Query: 71 GKPTGRFSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDV 130
PTGRF NG++ATDFI++ G Y PAYL+ G +FASA++GY TS +
Sbjct: 550 HFPTGRFCNGKLATDFIADILGFTSYQPAYLNLKTKGKNLLNGANFASASSGYFELTSKL 729
Query: 131 LSVIPLWKQLEYYKEYQKKLGAYLGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQ 190
S IPL KQLEYYKE Q KL G+ A I+ A+Y+IS GT+DF++NYY P
Sbjct: 730 YSSIPLSKQLEYYKECQTKLVEAAGQSSASSIISDAIYLISAGTSDFVQNYYINPLLNKL 909
Query: 191 YTPSEYQNFLAGIAQNFIHKLYDLGAKKISLGGLPPMGCLPLERTTNFAGGNDCVSNYNN 250
YT ++ + L NFI LY LGA++I + LPP+GCLP T A N+CV++ N+
Sbjct: 910 YTTDQFSDTLLRCYSNFIQSLYALGARRIGVTSLPPIGCLPAVITLFGAHINECVTSLNS 1089
Query: 251 IALEFNGKLNKLTTKLKKDLPGIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFE 310
A+ FN KLN + LK LPG+ LV + Y L + KP + GF A ACC TG+ E
Sbjct: 1090DAINFNEKLNTTSQNLKNMLPGLNLVVFDIYQPLYDLATKPSENGFFEARKACCGTGLIE 1269
Query: 311 MGYACSRASLFSCMDASRYVFWDSFHPTEKTNGIVANYLVKNALA 355
+ C++ S+ +C +AS YVFWD FHP+E N ++A+ L+ + ++
Sbjct: 1270VSILCNKKSIGTCANASEYVFWDGFHPSEAANKVLADELITSGIS 1404
>TC219543 weakly similar to UP|Q94CH8 (Q94CH8) Family II lipase EXL1, partial
(49%)
Length = 1085
Score = 286 bits (732), Expect = 1e-77
Identities = 141/343 (41%), Positives = 204/343 (59%), Gaps = 1/343 (0%)
Frame = +2
Query: 16 LCILLLQRLDLSLVKSAKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGG-KPT 74
+ I+ L +SL +PA+IVFGDS VD GNNN+I+T+A+ NF PYGRDF GG +PT
Sbjct: 62 IVIISLHVSSVSLPNYESIPAVIVFGDSIVDTGNNNYITTIAKCNFLPYGRDFGGGNQPT 241
Query: 75 GRFSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVI 134
GRFSNG +D I+ FG+K +P YLDP TGVSFAS A+GYD TS + S +
Sbjct: 242 GRFSNGLTPSDIIAAKFGVKELLPPYLDPKLQPQDLLTGVSFASGASGYDPLTSKIASAL 421
Query: 135 PLWKQLEYYKEYQKKLGAYLGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPS 194
L QL+ ++EY+ K+ +GE + I+K++YI+ G+ND Y+ R +Y
Sbjct: 422 SLSDQLDTFREYKNKIMEIVGENRTATIISKSIYILCTGSNDITNTYFV---RGGEYDIQ 592
Query: 195 EYQNFLAGIAQNFIHKLYDLGAKKISLGGLPPMGCLPLERTTNFAGGNDCVSNYNNIALE 254
Y + +A A NF+ +LY LGA++I + GLP +GC+P +RT + C N A+
Sbjct: 593 AYTDLMASQATNFLQELYGLGARRIGVVGLPVLGCVPSQRTLHGGIFRACSDFENEAAVL 772
Query: 255 FNGKLNKLTTKLKKDLPGIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYA 314
FN KL+ LKK R V+ + Y+ +L +++ P +YGF+V CC TG E+G
Sbjct: 773 FNSKLSSQMDALKKQFQEARFVYLDLYNPVLNLIQNPAKYGFEVMDQGCCGTGKLEVGPL 952
Query: 315 CSRASLFSCMDASRYVFWDSFHPTEKTNGIVANYLVKNALAQF 357
C+ +L C + S Y+FWDSFHPTE +V ++ + + F
Sbjct: 953 CNHFTLLICSNTSNYIFWDSFHPTEAAYNVVCTQVLDHKIKDF 1081
>TC207231 similar to UP|Q94CH6 (Q94CH6) Family II lipase EXL3, partial (70%)
Length = 1216
Score = 253 bits (645), Expect = 1e-67
Identities = 126/301 (41%), Positives = 182/301 (59%), Gaps = 2/301 (0%)
Frame = +3
Query: 41 GDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFGIKPYIPAY 100
GDS VD GNNN + T+ + +F PYG+DF GG PTGRF NG+I +D + E GIK +PAY
Sbjct: 3 GDSIVDPGNNNKVKTLVKCDFPPYGKDFEGGIPTGRFCNGKIPSDLLVEELGIKELLPAY 182
Query: 101 LDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKLGAYLGEKKAK 160
LDP+ S TGV FAS A+GYD T + SVI + +QL+ +KEY KL +GE + K
Sbjct: 183 LDPNLKPSDLVTGVCFASGASGYDPLTPKIASVISMSEQLDMFKEYIGKLKHIVGEDRTK 362
Query: 161 ETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIAQNFIHKLYDLGAKKIS 220
+ + +++ G++D Y+ R QY Y + + A NF+ +LY LGA++I
Sbjct: 363 FILANSFFLVVAGSDDIANTYFIARVRQLQYDIPAYTDLMLHSASNFVKELYGLGARRIG 542
Query: 221 LGGLPPMGCLPLERTTNFAGG--NDCVSNYNNIALEFNGKLNKLTTKLKKDLPGIRLVFS 278
+ PP+GC+P +RT AGG +C YN A FN KL++ LK +LP R+V+
Sbjct: 543 VLSAPPIGCVPSQRT--LAGGFQRECAEEYNYAAKLFNSKLSRELDALKHNLPNSRIVYI 716
Query: 279 NPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDASRYVFWDSFHPT 338
+ Y+ L+ ++ ++G++V CC TG E+ C+ +C DAS+YVFWDS+HPT
Sbjct: 717 DVYNPLMDIIVNYQRHGYKVVDRGCCGTGKLEVAVLCNPLGA-TCPDASQYVFWDSYHPT 893
Query: 339 E 339
E
Sbjct: 894 E 896
>TC225398 similar to UP|O04320 (O04320) Proline-rich protein APG isolog,
partial (93%)
Length = 1118
Score = 215 bits (547), Expect(2) = 7e-66
Identities = 107/265 (40%), Positives = 154/265 (57%), Gaps = 1/265 (0%)
Frame = +3
Query: 92 GIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKLG 151
G + Y PAYL P + G +FASAA+GYD + + IPL +QL Y+KEYQ KL
Sbjct: 117 GFQTYAPAYLSPQASGKNLLIGANFASAASGYDENAATLNHAIPLSQQLSYFKEYQGKLA 296
Query: 152 AYLGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIAQNFIHKL 211
G KKA I ALY++S G++DF++NYY P Y+P +Y ++L G +F+ L
Sbjct: 297 KVAGSKKAASIIKDALYVLSAGSSDFVQNYYVNPWINKVYSPDQYSSYLVGEFSSFVKDL 476
Query: 212 YDLGAKKISLGGLPPMGCLPLERTTNFAGGNDCVSNYNNIALEFNGKLNKLTTKLKKDLP 271
Y LGA+++ + LPP+GCLP RT N CVS N A FN KLN L+K LP
Sbjct: 477 YGLGARRLGVTSLPPLGCLPAARTIFGFHENGCVSRINTDAQGFNKKLNSAAASLQKQLP 656
Query: 272 GIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFE-MGYACSRASLFSCMDASRYV 330
G+++ + Y L +V+ P + GF A+ CC TG E C+ S +C +A++YV
Sbjct: 657 GLKIAIFDIYKPLYDLVQSPSKSGFVEANRGCCGTGTVETTSLLCNSKSPGTCSNATQYV 836
Query: 331 FWDSFHPTEKTNGIVANYLVKNALA 355
FWDS HP++ N ++A+ L+ ++
Sbjct: 837 FWDSVHPSQAANQVLADALILQGIS 911
Score = 53.9 bits (128), Expect(2) = 7e-66
Identities = 20/44 (45%), Positives = 32/44 (72%)
Frame = +1
Query: 54 STVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFGIKPYI 97
S ++++ PYGRDF+ +PTGRF NG++ATDF ++ G +P +
Sbjct: 4 SAPVKADYPPYGRDFVNHQPTGRFCNGKLATDFTADTLGFRPML 135
>TC203707
Length = 1394
Score = 226 bits (575), Expect = 2e-59
Identities = 121/343 (35%), Positives = 195/343 (56%), Gaps = 3/343 (0%)
Frame = +1
Query: 11 YTNNILCILLLQRLDLSLVKSAKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLG 70
Y N +L ++L + L ++ + A VFGDS VD GNNNF++T AR++ PYG D+
Sbjct: 40 YINIVLLGIVLVVVRLKGAEAQR--AFFVFGDSLVDNGNNNFLATTARADAPPYGIDYPT 213
Query: 71 GKPTGRFSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATS-D 129
G+PTGRFSNG DFIS++ G + +P YLDP + + G +FASA G N T
Sbjct: 214 GRPTGRFSNGYNIPDFISQSLGAESTLP-YLDPELDGERLLVGANFASAGIGILNDTGIQ 390
Query: 130 VLSVIPLWKQLEYYKEYQKKLGAYLGEKKAKETITKALYIISLGTNDFLENYYTIP--GR 187
+++I +++QLEY++EYQ+++ +G ++ + I AL +I+LG NDF+ NYY +P R
Sbjct: 391 FVNIIRIYRQLEYWEEYQQRVSGLIGPEQTERLINGALVLITLGGNDFVNNYYLVPYSAR 570
Query: 188 ASQYTPSEYQNFLAGIAQNFIHKLYDLGAKKISLGGLPPMGCLPLERTTNFAGGNDCVSN 247
+ QY +Y ++ + + +LY++GA+++ + G P+GC+P E G DC +
Sbjct: 571 SRQYNLPDYVKYIISEYKKVLRRLYEIGARRVLVTGTGPLGCVPAELAQRSTNG-DCSAE 747
Query: 248 YNNIALEFNGKLNKLTTKLKKDLPGIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATG 307
A FN +L ++ +L ++ V N + + + P +YGF + +ACC G
Sbjct: 748 LQQAAALFNPQLVQIIRQLNSEIGSNVFVGVNTQQMHIDFISNPQRYGFVTSKVACCGQG 927
Query: 308 MFEMGYACSRASLFSCMDASRYVFWDSFHPTEKTNGIVANYLV 350
+ C+ AS C + Y FWD FHPTE+ N I+ ++
Sbjct: 928 PYNGLGLCTPASNL-CPNRDSYAFWDPFHPTERANRIIVQQIL 1053
>TC217084
Length = 1310
Score = 222 bits (566), Expect = 2e-58
Identities = 124/360 (34%), Positives = 197/360 (54%), Gaps = 5/360 (1%)
Frame = +3
Query: 5 QRKNLLYTNNILCILLLQRLDLSLV--KSAKVPAIIVFGDSSVDAGNNNFISTVARSNFQ 62
Q L ++ LC+ LL L +V ++ A VFGDS VD GNNNF++T AR++
Sbjct: 66 QNNKSLVSSKFLCLCLLITLISIIVAPQAEAARAFFVFGDSLVDNGNNNFLATTARADSY 245
Query: 63 PYGRDFLGGKPTGRFSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATG 122
PYG D + +GRFSNG D ISE G +P +P YL P N + G +FASA G
Sbjct: 246 PYGIDSASHRASGRFSNGLNMPDLISEKIGSEPTLP-YLSPQLNGERLLVGANFASAGIG 422
Query: 123 YDNATS-DVLSVIPLWKQLEYYKEYQKKLGAYLGEKKAKETITKALYIISLGTNDFLENY 181
N T +++I + +QL Y+K+YQ+++ A +GE++ + + KAL +I+LG NDF+ NY
Sbjct: 423 ILNDTGIQFINIIRITEQLAYFKQYQQRVSALIGEEQTRNLVNKALVLITLGGNDFVNNY 602
Query: 182 YTIP--GRASQYTPSEYQNFLAGIAQNFIHKLYDLGAKKISLGGLPPMGCLPLERTTNFA 239
Y +P R+ +Y +Y FL + + LY+LGA+++ + G P+GC+P E +
Sbjct: 603 YLVPFSARSREYALPDYVVFLISEYRKILANLYELGARRVLVTGTGPLGCVPAELAMHSQ 782
Query: 240 GGNDCVSNYNNIALEFNGKLNKLTTKLKKDLPGIRLVFSNPYDVLLGVVKKPGQYGFQVA 299
G +C + FN +L +L +L + + +N + + L V P YGF +
Sbjct: 783 NG-ECATELQRAVNLFNPQLVQLLHELNTQIGSDVFISANAFTMHLDFVSNPQAYGFVTS 959
Query: 300 SMACCATGMFEMGYACSRASLFSCMDASRYVFWDSFHPTEKTNGIVANYLVKNALAQFLH 359
+ACC G + C+ AS C + Y FWD FHP+E+ N ++ + + + +++H
Sbjct: 960 KVACCGQGAYNGIGLCTPASNL-CPNRDLYAFWDPFHPSERANRLIVDKFMTGS-TEYMH 1133
>TC206646
Length = 1178
Score = 201 bits (510), Expect = 6e-52
Identities = 111/317 (35%), Positives = 174/317 (54%), Gaps = 3/317 (0%)
Frame = +1
Query: 46 DAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFGIKPYIPAYLDPSF 105
D+GNN+F+ T AR++ PYG D+ +PTGRFSNG D IS G+KP +P YL P
Sbjct: 7 DSGNNDFLVTTARADAPPYGIDYPTHRPTGRFSNGLNIPDLISLELGLKPTLP-YLSPLL 183
Query: 106 NISQFATGVSFASAATGYDNATS-DVLSVIPLWKQLEYYKEYQKKLGAYLGEKKAKETIT 164
+ G +FASA G N T L++I + KQL+ + EYQ++L ++G + +
Sbjct: 184 VGEKLLIGANFASAGIGILNDTGIQFLNIIHIQKQLKLFHEYQERLSLHIGAEGTRNLXN 363
Query: 165 KALYIISLGTNDFLENYYTIP--GRASQYTPSEYQNFLAGIAQNFIHKLYDLGAKKISLG 222
+AL +I+LG NDF+ NYY +P R+ Q++ +Y +L + + +LYDLGA+++ +
Sbjct: 364 RALVLITLGGNDFVNNYYLVPYSARSRQFSLPDYVRYLISEYRKVLRRLYDLGARRVLVT 543
Query: 223 GLPPMGCLPLERTTNFAGGNDCVSNYNNIALEFNGKLNKLTTKLKKDLPGIRLVFSNPYD 282
G PMGC+P E T G DC A FN +L ++ L ++L + +N
Sbjct: 544 GTGPMGCVPAELATRSRTG-DCDVELQRAASLFNPQLVQMLNGLNQELGADVFIAANAQR 720
Query: 283 VLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDASRYVFWDSFHPTEKTN 342
+ + V P YGF + +ACC G + C+ S C + Y FWD FHP+EK +
Sbjct: 721 MHMDFVSNPRAYGFVTSKIACCGQGPYNGVGLCTPTSNL-CPNRDLYAFWDPFHPSEKAS 897
Query: 343 GIVANYLVKNALAQFLH 359
I+ +++ +++H
Sbjct: 898 RIIVQQILRGT-TEYMH 945
>BF071190
Length = 379
Score = 199 bits (505), Expect = 2e-51
Identities = 95/124 (76%), Positives = 106/124 (84%)
Frame = +2
Query: 24 LDLSLVKSAKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIA 83
L L AKVP++IVF DSSVDAGNNN I T+ARSN+QPYGRDF GGK TGR+ NGRI
Sbjct: 8 LSLKAETIAKVPSMIVFCDSSVDAGNNNIIHTIARSNYQPYGRDFEGGKATGRYCNGRIP 187
Query: 84 TDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYY 143
TDFISE+FG+KPY+P YLDP +NIS FA+GVSFASAATGYDNATSDVL+VIPLW QLEYY
Sbjct: 188 TDFISESFGLKPYVPTYLDPKYNISDFASGVSFASAATGYDNATSDVLTVIPLWMQLEYY 367
Query: 144 KEYQ 147
K YQ
Sbjct: 368 KGYQ 379
>TC234426
Length = 652
Score = 191 bits (484), Expect = 6e-49
Identities = 94/129 (72%), Positives = 109/129 (83%), Gaps = 1/129 (0%)
Frame = +1
Query: 10 LYTNNILCILLLQRLDLSLVKSAKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFL 69
++++ I C+ L L + AKVPA+I FGDSSVDAGNNN+I+TVARSNFQPYGRDF+
Sbjct: 19 MHSSIIFCMFFLPWLSMV---GAKVPAMIAFGDSSVDAGNNNYIATVARSNFQPYGRDFV 189
Query: 70 GGKPTGRFSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSD 129
GGKPTGRFSNGRIATDF+S+AFGIKPY+P YLDP+ NIS FATGVSFASAATGYDNATSD
Sbjct: 190 GGKPTGRFSNGRIATDFLSQAFGIKPYVPPYLDPNHNISHFATGVSFASAATGYDNATSD 369
Query: 130 VLSV-IPLW 137
VL +P W
Sbjct: 370 VLGFRLPQW 396
Score = 127 bits (318), Expect = 1e-29
Identities = 58/64 (90%), Positives = 60/64 (93%)
Frame = +3
Query: 296 FQVASMACCATGMFEMGYACSRASLFSCMDASRYVFWDSFHPTEKTNGIVANYLVKNALA 355
FQV SMACCATGMFEMGYACSRAS FSC+DASRYV WDSFHPTEKTNGI+A YLVKNALA
Sbjct: 378 FQVTSMACCATGMFEMGYACSRASSFSCIDASRYVXWDSFHPTEKTNGIIAKYLVKNALA 557
Query: 356 QFLH 359
QFLH
Sbjct: 558 QFLH 569
>TC220781 weakly similar to UP|Q94CH6 (Q94CH6) Family II lipase EXL3, partial
(40%)
Length = 701
Score = 184 bits (467), Expect = 5e-47
Identities = 85/190 (44%), Positives = 122/190 (63%)
Frame = +3
Query: 31 SAKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEA 90
++ VPA++ FGDS VD GNNN I T+ + NF PYG+DF GG PTGRF NG+I +D I+E
Sbjct: 132 ASSVPAVLAFGDSIVDPGNNNNIKTLIKCNFPPYGKDFQGGNPTGRFCNGKIPSDLIAEQ 311
Query: 91 FGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKL 150
GIK Y+PAYLDP+ S TGV FAS A+GYD T + SV+ L QL+ ++EY KL
Sbjct: 312 LGIKEYLPAYLDPNLKSSDLVTGVCFASGASGYDPLTPKITSVLSLSTQLDMFREYIGKL 491
Query: 151 GAYLGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIAQNFIHK 210
+GE + ++ +LY++ G++D Y+ R QY Y + + A NF+ +
Sbjct: 492 KGIVGESRTNYILSNSLYLVVAGSDDIANTYFVAHARILQYDIPSYTDLMVNSASNFVKE 671
Query: 211 LYDLGAKKIS 220
LY+LGA++++
Sbjct: 672 LYNLGARRVA 701
>TC203468
Length = 887
Score = 182 bits (462), Expect = 2e-46
Identities = 97/210 (46%), Positives = 130/210 (61%), Gaps = 1/210 (0%)
Frame = +1
Query: 26 LSLVKSAKVPAIIVFGDSSVDAGNNNFIST-VARSNFQPYGRDFLGGKPTGRFSNGRIAT 84
+ L VPA+ VFGDS VD GNNN +T ARSNF PYGRDF GG PTGRFSNG++ +
Sbjct: 148 VKLPADVSVPAVFVFGDSVVDTGNNNNRTTSFARSNFPPYGRDFQGGIPTGRFSNGKVPS 327
Query: 85 DFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYK 144
D I E GIK +PAYL P+ S TGV FAS +GYD TS + S +PL Q++ K
Sbjct: 328 DLIVEELGIKELLPAYLKPNLQSSDLITGVCFASGGSGYDPLTSILESSMPLTGQVDLLK 507
Query: 145 EYQKKLGAYLGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIA 204
EY KL +GE +AK + +L+I+ G++D Y T R+ Y Y + L A
Sbjct: 508 EYIGKLKGLVGEDRAKFILANSLFIVVAGSSDISNTYRT---RSLLYDLPAYTDLLVNSA 678
Query: 205 QNFIHKLYDLGAKKISLGGLPPMGCLPLER 234
NF+ ++ +LGA+KI++ PP+GCLP ++
Sbjct: 679 SNFLTEINELGARKIAVFSAPPIGCLPFQK 768
>TC218565
Length = 626
Score = 168 bits (425), Expect = 4e-42
Identities = 75/151 (49%), Positives = 103/151 (67%)
Frame = +3
Query: 157 KKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIAQNFIHKLYDLGA 216
KK + K LY++SLGTNDFLENYY P R +T S+Y++FL IA+NF+ +LY LG
Sbjct: 171 KKPTRS*AKLLYLMSLGTNDFLENYYVFPTRRLHFTVSQYEDFLLRIAENFVRELYALGV 350
Query: 217 KKISLGGLPPMGCLPLERTTNFAGGNDCVSNYNNIALEFNGKLNKLTTKLKKDLPGIRLV 276
+K+S+ GL P+GCLPLER TN G + C YNN+A+ FN KL + TKL +DLP ++ +
Sbjct: 351 RKLSITGLIPVGCLPLERATNIFGDHGCNEEYNNVAMSFNKKLENVITKLNRDLPQLKAL 530
Query: 277 FSNPYDVLLGVVKKPGQYGFQVASMACCATG 307
+N Y + ++ KP YGF+V ACC+TG
Sbjct: 531 SANAYSIFSDIITKPSTYGFEVVEKACCSTG 623
Score = 104 bits (260), Expect = 5e-23
Identities = 51/85 (60%), Positives = 62/85 (72%)
Frame = +2
Query: 101 LDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKLGAYLGEKKAK 160
LDP+F I FATGV FASA TGYDNATS VL+VIPLWK+LEYYKEYQ KL A++G +KA
Sbjct: 2 LDPAFTIKDFATGVCFASAGTGYDNATSAVLNVIPLWKELEYYKEYQAKLRAHVGVEKAN 181
Query: 161 ETITKALYIISLGTNDFLENYYTIP 185
E I++A+ + LG F +P
Sbjct: 182 EIISEAVVLDELGNQRFPRKLLRVP 256
>TC228081 similar to UP|Q9FFC6 (Q9FFC6) GDSL-motif lipase/hydrolase-like
protein, partial (52%)
Length = 680
Score = 166 bits (420), Expect = 2e-41
Identities = 82/184 (44%), Positives = 114/184 (61%)
Frame = +3
Query: 34 VPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFGI 93
VPA+ +FGDS VD GNNN + T+ ++NF PYGRDF PTGRF NG++A+D+ +E G
Sbjct: 129 VPALFIFGDSVVDVGNNNHLYTIVKANFPPYGRDFKNHNPTGRFCNGKLASDYTAENLGF 308
Query: 94 KPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKLGAY 153
Y PAYL+ + G +FASAA+GY + T+ + IPL +QLE+YKE Q L
Sbjct: 309 TSYPPAYLNLKAKGNNLLNGANFASAASGYYDPTAKLYHAIPLSQQLEHYKECQNILVGT 488
Query: 154 LGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIAQNFIHKLYD 213
+G+ A I+ A+Y+IS G +DF++NYY P YT ++ + L FI LY
Sbjct: 489 VGQPNASSIISGAIYLISAGNSDFIQNYYINPLLYKVYTADQFSDILLQSYATFIQNLYA 668
Query: 214 LGAK 217
LGA+
Sbjct: 669 LGAR 680
>TC223028 weakly similar to UP|Q94CH7 (Q94CH7) Family II lipase EXL2, partial
(44%)
Length = 640
Score = 166 bits (419), Expect = 2e-41
Identities = 84/175 (48%), Positives = 108/175 (61%), Gaps = 1/175 (0%)
Frame = +3
Query: 18 ILLLQRLDLSLVKSAKVPAIIVFGDSSVDAGNNNF-ISTVARSNFQPYGRDFLGGKPTGR 76
I+ R + L + VPA++VFGDS VD GNNN + T AR N+ PYG+DF GGKPTGR
Sbjct: 105 IVCKTRAVVKLPPNVSVPAVLVFGDSIVDTGNNNNNLGTTARCNYPPYGKDFEGGKPTGR 284
Query: 77 FSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPL 136
FSNG++ +DFI+E GIK Y+PAYLDP + ATGV FAS GYD TS S I L
Sbjct: 285 FSNGKVPSDFIAEELGIKEYVPAYLDPHLQPGELATGVCFASGGAGYDPLTSQSASAISL 464
Query: 137 WKQLEYYKEYQKKLGAYLGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQY 191
QL+ +KEY KL +GE + + +LY++ G+ND Y+ R QY
Sbjct: 465 SGQLDLFKEYLGKLRGVVGEDRTNFILANSLYVVVFGSNDISNTYFLSRVRQLQY 629
>TC226564
Length = 1337
Score = 159 bits (402), Expect = 2e-39
Identities = 104/317 (32%), Positives = 163/317 (50%), Gaps = 6/317 (1%)
Frame = +3
Query: 32 AKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAF 91
++VP + +FGDS D+GNNN + T ++SNF+PYG DF G PTGR++NGR D I++
Sbjct: 129 SQVPCLFIFGDSMSDSGNNNELPTTSKSNFRPYGIDFPLG-PTGRYTNGRTEIDIITQFL 305
Query: 92 GIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATS-DVLSVIPLWKQLEYYKEYQKKL 150
G + +IP + + S S GV++AS +G N T + I L QL ++ ++
Sbjct: 306 GFEKFIPPFANTSG--SDILKGVNYASGGSGIRNETGWHYGAAIGLGLQLANHRVIVSEI 479
Query: 151 GAYLGEKK-AKETITKALYIISLGTNDFLENYYTIP--GRASQYTPSEYQNFLAGIAQNF 207
LG A++ + K LY +++G+ND++ NY+ P ++ YT E+ L
Sbjct: 480 ATKLGSPDLARQYLEKCLYYVNIGSNDYMGNYFLPPFYPTSTIYTIEEFTQVLIEELSLN 659
Query: 208 IHKLYDLGAKKISLGGLPPMGCLPLERTTNFAGGNDCVSNYNNIALEFNGKLNKLTTKLK 267
+ L+D+GA+K +L GL +GC P + + G+ C N A FN KL +
Sbjct: 660 LQALHDIGARKYALAGLGLIGCTPGMVSAHGTNGS-CAEEQNLAAFNFNNKLKARVDQFN 836
Query: 268 KDL--PGIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMD 325
D + +F N + + + K YGF V CC G+ G C +
Sbjct: 837 NDFYYANSKFIFINTQALAIELRDK---YGFPVPETPCCLPGL--TGECVPDQE--PCYN 995
Query: 326 ASRYVFWDSFHPTEKTN 342
+ YVF+D+FHPTE+ N
Sbjct: 996 RNDYVFFDAFHPTEQWN 1046
>TC207585 similar to UP|Q9AYM7 (Q9AYM7) CPRD47 protein, partial (73%)
Length = 840
Score = 152 bits (383), Expect = 3e-37
Identities = 94/282 (33%), Positives = 149/282 (52%), Gaps = 10/282 (3%)
Frame = +1
Query: 33 KVPAIIVFGDSSVDAGNNNFIS-TVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAF 91
K PA+ VFGDS VD GNNN++S ++ ++ YG DF KPTGRFSNG+ A D I+E
Sbjct: 4 KAPAVYVFGDSLVDVGNNNYLSLSIEKAILPHYGIDFPTKKPTGRFSNGKNAADLIAENL 183
Query: 92 GIKPYIPAYL--------DPSFNISQFATGVSFASAATGYDNAT-SDVLSVIPLWKQLEY 142
G+ P P YL + N+S F GV+FAS G NA+ IPL KQ++Y
Sbjct: 184 GL-PTSPPYLSLVSKVHNNNKKNVS-FLGGVNFASGGAGIFNASDKGFRQSIPLPKQVDY 357
Query: 143 YKEYQKKLGAYLGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAG 202
Y + ++L +G + ++K+++I+ +G ND Y+ + TP +Y + +A
Sbjct: 358 YSQVHEQLIQQIGASTLGKHLSKSIFIVVIGGNDIF-GYFDSKDLQKKNTPQQYVDSMAS 534
Query: 203 IAQNFIHKLYDLGAKKISLGGLPPMGCLPLERTTNFAGGNDCVSNYNNIALEFNGKLNKL 262
+ + +LY+ GAKK + G+ +GC P R N +CVS N++++++N L +
Sbjct: 535 TLKVQLQRLYNNGAKKFEIAGVGAIGCCPAYRVKN---KTECVSEANDLSVKYNEALQSM 705
Query: 263 TTKLKKDLPGIRLVFSNPYDVLLGVVKKPGQYGFQVASMACC 304
+ + + I + + Y L +V P Y + ACC
Sbjct: 706 LKEWRLENKHISYSYFDTYAGLQDLVHNPASYRYAYEKAACC 831
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.138 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,936,907
Number of Sequences: 63676
Number of extensions: 196351
Number of successful extensions: 1249
Number of sequences better than 10.0: 157
Number of HSP's better than 10.0 without gapping: 1093
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1099
length of query: 359
length of database: 12,639,632
effective HSP length: 98
effective length of query: 261
effective length of database: 6,399,384
effective search space: 1670239224
effective search space used: 1670239224
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC140670.14


