

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140550.8 + phase: 0 /pseudo
(496 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
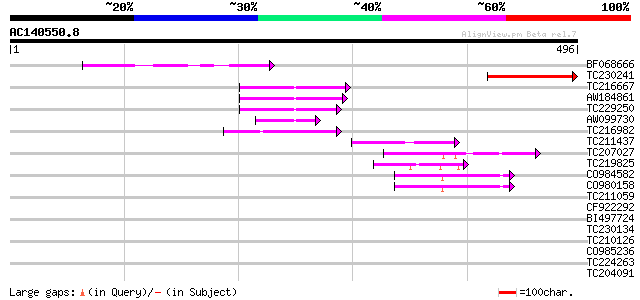
Score E
Sequences producing significant alignments: (bits) Value
BF068666 143 2e-34
TC230241 similar to UP|Q6WBD3 (Q6WBD3) Glycophorin A-like protei... 121 6e-28
TC216667 weakly similar to GB|BAC06968.1|22093674|AP003815 kinas... 70 2e-12
AW184861 62 5e-10
TC229250 similar to UP|Q9AR00 (Q9AR00) PSTVd RNA-biding protein,... 58 9e-09
AW099730 weakly similar to GP|29569106|gb| global transcription ... 50 2e-06
TC216982 similar to UP|Q9AR19 (Q9AR19) Histone acetyltransferase... 46 3e-05
TC211437 similar to UP|Q6VBJ4 (Q6VBJ4) Epa5p, partial (7%) 45 7e-05
TC207027 weakly similar to UP|Q9LW95 (Q9LW95) KED, partial (17%) 42 5e-04
TC219825 weakly similar to UP|MID2_YEAST (P36027) Mating process... 42 6e-04
CO984582 42 8e-04
CO980158 42 8e-04
TC211059 similar to UP|Q8LRK9 (Q8LRK9) HAF1, partial (7%) 41 0.001
CF922292 41 0.001
BI497724 similar to GP|16323117|gb| AT3g01770/F28J7_10 {Arabidop... 40 0.002
TC230134 weakly similar to UP|NUCL_HUMAN (P19338) Nucleolin (Pro... 40 0.003
TC210126 similar to UP|NUCL_HUMAN (P19338) Nucleolin (Protein C2... 39 0.004
CO985236 39 0.004
TC224263 weakly similar to UP|Q6GQH4 (Q6GQH4) Egr1 protein, part... 39 0.005
TC204091 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1... 39 0.005
>BF068666
Length = 405
Score = 143 bits (361), Expect = 2e-34
Identities = 88/170 (51%), Positives = 103/170 (59%), Gaps = 2/170 (1%)
Frame = +2
Query: 64 QHS-NLANINPDGSIRKP-VGRVKVKLKTSKMLDSQPNSSDAPSQSDTDKSSQQQHGLER 121
QH+ N+ANINPDGS+ K +GRVKVKLKT K+LDSQP SSDAP+ + +D
Sbjct: 5 QHNYNVANINPDGSVDKAAIGRVKVKLKTPKLLDSQPTSSDAPTPTQSD----------- 151
Query: 122 HGVNADRIEDSVTSFPDLKYGASYKKAGSIKIKSSKVLGLNADQTSKPLPISSEIIHPKE 181
+R+EDS PD+K K+A SIKIKSS S S+ H
Sbjct: 152 ----TERMEDSANLLPDMKLSTLSKRAASIKIKSS----------SNHQTFSASTQHN-- 283
Query: 182 RKMPPLNPRYNKHELDTSLMIIRKVMKMDAAEPFNVPVNPEALGIPDYFD 231
NK ELD SL++IRKVMKMDAAEPFNVPVNPEALGIPDYFD
Sbjct: 284 ----------NKQELDASLVVIRKVMKMDAAEPFNVPVNPEALGIPDYFD 403
>TC230241 similar to UP|Q6WBD3 (Q6WBD3) Glycophorin A-like protein
(Fragment), partial (25%)
Length = 534
Score = 121 bits (304), Expect = 6e-28
Identities = 60/78 (76%), Positives = 63/78 (79%)
Frame = +2
Query: 419 LAEESEVGDTAALHDEYKNTQQEGQAAVVQQQKKHKEPQDKHQKAKLLESLCSENPMLSS 478
LAE+S VGD ALHDEY+ QQEGQAA V QQKK K QDK QK KLLESL SENP LSS
Sbjct: 2 LAEKSNVGDATALHDEYRTKQQEGQAAAVPQQKKPKGSQDKQQKGKLLESLYSENPKLSS 181
Query: 479 LCGTLFPKNSQSVWSGPH 496
LCG LFP+NSQSVWSGPH
Sbjct: 182 LCGILFPQNSQSVWSGPH 235
>TC216667 weakly similar to GB|BAC06968.1|22093674|AP003815 kinase-like
protein {Oryza sativa (japonica cultivar-group);} ,
partial (21%)
Length = 892
Score = 70.1 bits (170), Expect = 2e-12
Identities = 40/97 (41%), Positives = 54/97 (55%)
Frame = +3
Query: 202 IIRKVMKMDAAEPFNVPVNPEALGIPDYFDIIDTPMDFGTICSNLEKNDKYMNSEDVYND 261
I++ +M + F+ PV+P AL IPDYF II PMD GTI S LEKN Y +E+ D
Sbjct: 507 ILKSLMSHTYSWVFSKPVDPIALSIPDYFTIISHPMDLGTIKSKLEKN-IYSGTEEFAAD 683
Query: 262 VRYIWENCYKYNNKGDYIVDLMKRVKKNFMKYWAAAG 298
VR + N KYN + + + K + K F + W G
Sbjct: 684 VRLTFSNAMKYNPPSNDVHLMAKELSKIFDRKWKDLG 794
>AW184861
Length = 448
Score = 62.4 bits (150), Expect = 5e-10
Identities = 34/94 (36%), Positives = 51/94 (54%)
Frame = +2
Query: 202 IIRKVMKMDAAEPFNVPVNPEALGIPDYFDIIDTPMDFGTICSNLEKNDKYMNSEDVYND 261
++ K+M+ FN PV+ E LG+ DYF II PMD GT+ + L KN Y + ++ D
Sbjct: 8 LLEKLMRHKHGWVFNSPVDVETLGLHDYFTIITHPMDLGTVKTRLNKN-WYKSPKEFAED 184
Query: 262 VRYIWENCYKYNNKGDYIVDLMKRVKKNFMKYWA 295
VR + N YN +G + + + + K F WA
Sbjct: 185 VRLTFRNAMTYNPQGQDVHIMAELLSKIFEDRWA 286
>TC229250 similar to UP|Q9AR00 (Q9AR00) PSTVd RNA-biding protein, Virp1
(PSTVd RNA-binding protein Virp1a) (PSTVd RNA-binding
protein Virp1b) (PSTVd RNA-binding protein Virp1c)
(PSTVd RNA-binding protein Virp1d), partial (34%)
Length = 1450
Score = 58.2 bits (139), Expect = 9e-09
Identities = 32/89 (35%), Positives = 47/89 (51%)
Frame = +2
Query: 202 IIRKVMKMDAAEPFNVPVNPEALGIPDYFDIIDTPMDFGTICSNLEKNDKYMNSEDVYND 261
+++K++K F PV+ L + DY DII PMD GT+ SNL KN Y D +D
Sbjct: 677 VLQKLIKHKHGWVFKAPVDVVGLKLHDYCDIIKQPMDLGTVKSNLSKN-VYATPADFASD 853
Query: 262 VRYIWENCYKYNNKGDYIVDLMKRVKKNF 290
VR + N YN KG + + +++ F
Sbjct: 854 VRLTFNNALAYNPKGHDVYTMAEQLLARF 940
>AW099730 weakly similar to GP|29569106|gb| global transcription factor group
E {Zea mays}, partial (8%)
Length = 177
Score = 50.4 bits (119), Expect = 2e-06
Identities = 25/57 (43%), Positives = 32/57 (55%)
Frame = +2
Query: 216 NVPVNPEALGIPDYFDIIDTPMDFGTICSNLEKNDKYMNSEDVYNDVRYIWENCYKY 272
N PV+ L + DY+D+I PMD GT+ SNL N KY D +DVR + N Y
Sbjct: 8 NAPVDVVGLQLTDYYDVIKQPMDLGTVKSNLSMN-KYTTPSDFASDVRLTFNNALAY 175
>TC216982 similar to UP|Q9AR19 (Q9AR19) Histone acetyltransferase GCN5
(At3g54610) (Expressed protein), partial (53%)
Length = 1203
Score = 46.2 bits (108), Expect = 3e-05
Identities = 28/103 (27%), Positives = 45/103 (43%)
Frame = +1
Query: 188 NPRYNKHELDTSLMIIRKVMKMDAAEPFNVPVNPEALGIPDYFDIIDTPMDFGTICSNLE 247
N KH +++ + A PF PV +A +PDY+DII PMD T+ ++
Sbjct: 556 NATNQKHLNGFMRSLLKSMFDHADAWPFKEPV--DARDVPDYYDIIKDPMDLKTMSKRVD 729
Query: 248 KNDKYMNSEDVYNDVRYIWENCYKYNNKGDYIVDLMKRVKKNF 290
Y+ E D R ++ N YN+ R++ +F
Sbjct: 730 SEQYYVTFEMFVADARRMFANARTYNSPETIYYKCSTRLEAHF 858
>TC211437 similar to UP|Q6VBJ4 (Q6VBJ4) Epa5p, partial (7%)
Length = 857
Score = 45.1 bits (105), Expect = 7e-05
Identities = 30/94 (31%), Positives = 44/94 (45%)
Frame = +1
Query: 300 YSEQPKGSTDFKQEESMLVESPGGEDSLSNTDELLGTDGDVDDDKGEVTKMETSEKNCSP 359
Y + T+ +++ E ED + +E G +GD D + E N
Sbjct: 343 YLQNKNSETEDEEDGDEDDEDGPDEDEDGDDEEFSGEEGDEGGDPED-------EANGEG 501
Query: 360 SDGRHEVNEVDDDGMEEENHGPEDVEEDEEEDGE 393
SD E ++ DD+G EE++ ED EEDEEED E
Sbjct: 502 SDDGDEDDDGDDNGDEEDDDDGEDEEEDEEEDEE 603
Score = 40.0 bits (92), Expect = 0.002
Identities = 21/71 (29%), Positives = 37/71 (51%), Gaps = 6/71 (8%)
Frame = +1
Query: 337 DGDVDDDKGEVTKMETSEKNCSPSDG------RHEVNEVDDDGMEEENHGPEDVEEDEEE 390
DGD DD+ G + ++ S +G E N D +E++ G ++ +E++++
Sbjct: 382 DGDEDDEDGPDEDEDGDDEEFSGEEGDEGGDPEDEANGEGSDDGDEDDDGDDNGDEEDDD 561
Query: 391 DGEGENEEEIE 401
DGE E E+E E
Sbjct: 562 DGEDEEEDEEE 594
>TC207027 weakly similar to UP|Q9LW95 (Q9LW95) KED, partial (17%)
Length = 1005
Score = 42.4 bits (98), Expect = 5e-04
Identities = 35/160 (21%), Positives = 70/160 (42%), Gaps = 23/160 (14%)
Frame = +3
Query: 328 SNTDELLGTDGDVDDDKGEVTKMETSEKNCSPSDGRHEVNEVDDDGMEEEN--------- 378
++ +++ G + D +DD+ + K + +K+ ++ + +DDG ++E
Sbjct: 81 TDVEKVKGKENDGEDDEEKKEKKKKEKKDKDGKTDAEKIKKKEDDGKDDEEKKEKKKKYD 260
Query: 379 -------HGPEDVEEDE-------EEDGEGENEEEIEMNRVKRRMDETLRHGGTLAEESE 424
G ED +DE +E G+G+ EE+ K + D+ ++ E
Sbjct: 261 KIDAXKVKGKEDDGKDEGNKEKKDKEKGDGDGEEK------KEKKDKEKEKKKEKKDKDE 422
Query: 425 VGDTAALHDEYKNTQQEGQAAVVQQQKKHKEPQDKHQKAK 464
DT L ++ KN + E +++K KE + H+K K
Sbjct: 423 ETDT--LKEKGKNDEGEDDEGNKKKKKDKKEKEKDHKKEK 536
Score = 36.2 bits (82), Expect = 0.035
Identities = 33/155 (21%), Positives = 68/155 (43%), Gaps = 25/155 (16%)
Frame = +3
Query: 338 GDVDDDKGEVTKMETSEKNCSPSDGRHEVNEV---DDDGMEEENHGPEDVEEDEEEDG-- 392
GD ++K E K E +K + +V +V ++DG ++E + +E +++DG
Sbjct: 3 GDDGEEKKEKDKKEKKKKENKDKGDKTDVEKVKGKENDGEDDEEKKEKKKKEKKDKDGKT 182
Query: 393 ----------EGENEEE----------IEMNRVKRRMDETLRHGGTLAEESEVGDTAALH 432
+G+++EE I+ +VK + D+ G ++ E GD
Sbjct: 183 DAEKIKKKEDDGKDDEEKKEKKKKYDKIDAXKVKGKEDDGKDEGNKEKKDKEKGD----- 347
Query: 433 DEYKNTQQEGQAAVVQQQKKHKEPQDKHQKAKLLE 467
+E + ++++K KE +DK ++ L+
Sbjct: 348 ----GDGEEKKEKKDKEKEKKKEKKDKDEETDTLK 440
>TC219825 weakly similar to UP|MID2_YEAST (P36027) Mating process protein
MID2 (Serine-rich protein SMS1) (Protein kinase A
interference protein), partial (19%)
Length = 862
Score = 42.0 bits (97), Expect = 6e-04
Identities = 34/94 (36%), Positives = 46/94 (48%), Gaps = 11/94 (11%)
Frame = -2
Query: 319 ESPGGEDSLSNTDELLGTDGDVDDDKGEVTK--METSEKNCSPSDGRHEVNEVDDDGME- 375
ES EDS + ++G DG+ +D+GE + +E E+ P +G E E DG E
Sbjct: 321 ESEDDEDSEEEEEGVVGVDGEDHEDQGEEDEGVVEDHEEG-DP*EGEEEALEDAGDGGEG 145
Query: 376 -EENHGPEDVEEDEEED-------GEGENEEEIE 401
E G VE + ED GEGE EEE+E
Sbjct: 144 GGEGDGLGGVEGGDGEDGGGGGEEGEGEEEEEVE 43
>CO984582
Length = 849
Score = 41.6 bits (96), Expect = 8e-04
Identities = 30/108 (27%), Positives = 46/108 (41%), Gaps = 3/108 (2%)
Frame = -1
Query: 337 DGDVDDDKGEVTKMETSEKNCSPSDGRHEVNEVDDDGMEEE--NHGPEDVEEDEEEDGEG 394
D ++DD ++ E+ +H E DD E N P++V E++EED
Sbjct: 579 DINIDDSSCSYPLVQVDEEEDYYDGTQHSDYESDDSNAENNPLNDYPDEVSEEDEEDENS 400
Query: 395 ENEEEIE-MNRVKRRMDETLRHGGTLAEESEVGDTAALHDEYKNTQQE 441
ENE E E N DE + HG + + + D DEY+ +
Sbjct: 399 ENESEREDGNTSSESSDENIEHGFSEDDADPLYDED--FDEYEGAHSD 262
>CO980158
Length = 803
Score = 41.6 bits (96), Expect = 8e-04
Identities = 30/108 (27%), Positives = 46/108 (41%), Gaps = 3/108 (2%)
Frame = -3
Query: 337 DGDVDDDKGEVTKMETSEKNCSPSDGRHEVNEVDDDGMEEE--NHGPEDVEEDEEEDGEG 394
D ++DD ++ E+ +H E DD E N P++V E++EED
Sbjct: 474 DINIDDSSCSYPLVQVDEEEDYYDGTQHSDYESDDSNAENNPLNDYPDEVSEEDEEDENS 295
Query: 395 ENEEEIE-MNRVKRRMDETLRHGGTLAEESEVGDTAALHDEYKNTQQE 441
ENE E E N DE + HG + + + D DEY+ +
Sbjct: 294 ENESEREDGNTSSESSDENIEHGFSEDDADPLYDED--FDEYEGAHSD 157
>TC211059 similar to UP|Q8LRK9 (Q8LRK9) HAF1, partial (7%)
Length = 800
Score = 41.2 bits (95), Expect = 0.001
Identities = 26/82 (31%), Positives = 39/82 (46%)
Frame = +2
Query: 193 KHELDTSLMIIRKVMKMDAAEPFNVPVNPEALGIPDYFDIIDTPMDFGTICSNLEKNDKY 252
K D S + ++ V K +A PDY DII+ PMD I + +N +Y
Sbjct: 263 KDRYDLSYLFLKPVSKKEA---------------PDYLDIIERPMDLSRIRERV-RNMEY 394
Query: 253 MNSEDVYNDVRYIWENCYKYNN 274
+ ED +D+ I N +KYN+
Sbjct: 395 KSREDFRHDMWQITFNAHKYND 460
>CF922292
Length = 722
Score = 40.8 bits (94), Expect = 0.001
Identities = 33/161 (20%), Positives = 71/161 (43%)
Frame = -2
Query: 301 SEQPKGSTDFKQEESMLVESPGGEDSLSNTDELLGTDGDVDDDKGEVTKMETSEKNCSPS 360
S P + D QE + V+ + S +N+ + GD+ +D + T +KN S
Sbjct: 502 SVTPVQNVDVSQENNSEVKEQATDPSNNNSQQFEDNRGDLSEDATKGDGSVTPDKN---S 332
Query: 361 DGRHEVNEVDDDGMEEENHGPEDVEEDEEEDGEGENEEEIEMNRVKRRMDETLRHGGTLA 420
D + + E D+ +E+ ED + + ++ E + + ++ K DE+ + +
Sbjct: 331 DVKEKQEEKSDEKSQEK--PSEDTKTENQDTSVSEKRSDSDESQQKSDSDESQQKSD--S 164
Query: 421 EESEVGDTAALHDEYKNTQQEGQAAVVQQQKKHKEPQDKHQ 461
+ESE +A ++ ++ + + + + +K E D Q
Sbjct: 163 DESEKKSDSAESEKKSDSDESEKKSDSDETEKSSESNDNKQ 41
>BI497724 similar to GP|16323117|gb| AT3g01770/F28J7_10 {Arabidopsis
thaliana}, partial (10%)
Length = 408
Score = 40.0 bits (92), Expect = 0.002
Identities = 19/45 (42%), Positives = 27/45 (59%)
Frame = +1
Query: 202 IIRKVMKMDAAEPFNVPVNPEALGIPDYFDIIDTPMDFGTICSNL 246
++++VM + F+ PV+ IPDYF II PMD GT+ S L
Sbjct: 244 LLKRVMSHQFGKVFDKPVDIVKWNIPDYFTIIKHPMDLGTVKSKL 378
>TC230134 weakly similar to UP|NUCL_HUMAN (P19338) Nucleolin (Protein C23),
partial (8%)
Length = 581
Score = 39.7 bits (91), Expect = 0.003
Identities = 25/92 (27%), Positives = 43/92 (46%), Gaps = 7/92 (7%)
Frame = +2
Query: 324 EDSLSNTDELLGTDGDVDDDKGEVTKMETSEKNCSPSD-------GRHEVNEVDDDGMEE 376
ED + D G DGD DD+ E + + D G + ++ DDD ++
Sbjct: 281 EDDDDDDDVNDGEDGDDDDEDDEDFSGDDGGEEADSDDDPEANGGGESDDDDEDDDDDDD 460
Query: 377 ENHGPEDVEEDEEEDGEGENEEEIEMNRVKRR 408
+N + E+D++E+ E E+EE + KR+
Sbjct: 461 DNDEDDGDEDDDDEEDEDEDEETPQPPSKKRK 556
Score = 36.6 bits (83), Expect = 0.027
Identities = 23/95 (24%), Positives = 47/95 (49%), Gaps = 2/95 (2%)
Frame = +2
Query: 301 SEQPKGSTDFKQEESM--LVESPGGEDSLSNTDELLGTDGDVDDDKGEVTKMETSEKNCS 358
SE+ K ++D + ++ + + G+D + ++ G DG + D + + ++
Sbjct: 251 SEENKDASDTEDDDDDDDVNDGEDGDDDDEDDEDFSGDDGGEEADSDDDPEANGGGES-- 424
Query: 359 PSDGRHEVNEVDDDGMEEENHGPEDVEEDEEEDGE 393
D + ++ DDD E++ +D EEDE+ED E
Sbjct: 425 -DDDDEDDDDDDDDNDEDDGDEDDDDEEDEDEDEE 526
Score = 33.5 bits (75), Expect = 0.22
Identities = 27/113 (23%), Positives = 48/113 (41%)
Frame = +2
Query: 352 TSEKNCSPSDGRHEVNEVDDDGMEEENHGPEDVEEDEEEDGEGENEEEIEMNRVKRRMDE 411
TSE+N SD ++ DDD + + G +D E+DE+ G+ EE D+
Sbjct: 248 TSEENKDASDTE---DDDDDDDVNDGEDGDDDDEDDEDFSGDDGGEE-------ADSDDD 397
Query: 412 TLRHGGTLAEESEVGDTAALHDEYKNTQQEGQAAVVQQQKKHKEPQDKHQKAK 464
+GG +++ + D D ++ E + + + PQ +K K
Sbjct: 398 PEANGGGESDDDDEDDDDDDDDNDEDDGDEDDDDEEDEDEDEETPQPPSKKRK 556
>TC210126 similar to UP|NUCL_HUMAN (P19338) Nucleolin (Protein C23), partial
(7%)
Length = 735
Score = 39.3 bits (90), Expect = 0.004
Identities = 27/86 (31%), Positives = 39/86 (44%), Gaps = 1/86 (1%)
Frame = +3
Query: 324 EDSLSNTDELLGTDGDVDDDKGEVTKMETSEKNCSPSDGRHEVNEVDDDGMEEENHGPED 383
+D + D+ G D D D + E + E E+ D DDD + +N +D
Sbjct: 345 DDKDGDEDDEDGHDQDDDGEDEEFSGEEGDEEGDPEDDPEANGGGSDDDDDDNDNEEDDD 524
Query: 384 VEEDEEED-GEGENEEEIEMNRVKRR 408
EDEEE+ E E+EEE K+R
Sbjct: 525 DGEDEEEEEDEEEDEEETSQPPTKKR 602
Score = 36.6 bits (83), Expect = 0.027
Identities = 24/108 (22%), Positives = 50/108 (46%)
Frame = +3
Query: 355 KNCSPSDGRHEVNEVDDDGMEEENHGPEDVEEDEEEDGEGENEEEIEMNRVKRRMDETLR 414
KN D + +E D+DG ++++ G ++ EE D EG+ E++ E N
Sbjct: 324 KNSETEDDDKDGDEDDEDGHDQDDDGEDEEFSGEEGDEEGDPEDDPEAN----------- 470
Query: 415 HGGTLAEESEVGDTAALHDEYKNTQQEGQAAVVQQQKKHKEPQDKHQK 462
GG ++ + D D+ ++ ++E + +++ +P K +K
Sbjct: 471 -GGGSDDDDDDNDNEEDDDDGEDEEEEEDEE--EDEEETSQPPTKKRK 605
Score = 33.1 bits (74), Expect = 0.29
Identities = 21/76 (27%), Positives = 34/76 (44%)
Frame = +3
Query: 337 DGDVDDDKGEVTKMETSEKNCSPSDGRHEVNEVDDDGMEEENHGPEDVEEDEEEDGEGEN 396
DGD DD+ G + ++ S +G E + DD +D + D EED + +
Sbjct: 354 DGDEDDEDGHDQDDDGEDEEFSGEEGDEEGDPEDDPEANGGGSDDDDDDNDNEEDDD-DG 530
Query: 397 EEEIEMNRVKRRMDET 412
E+E E + +ET
Sbjct: 531 EDEEEEEDEEEDEEET 578
>CO985236
Length = 466
Score = 39.3 bits (90), Expect = 0.004
Identities = 40/151 (26%), Positives = 69/151 (45%), Gaps = 15/151 (9%)
Frame = -1
Query: 324 EDSLSNTDELLGTDG--DVDDDKGEVTKMETSEKNCSPSDGRHEVNEVDDDG------ME 375
E+ + E DG +V++D+ E E E DG V EV++DG
Sbjct: 439 EEGVEEATEDKKEDGVEEVNEDRKEDGVGEVKEDK--EVDGVSCVKEVEEDGEVDGVKKV 266
Query: 376 EENHGPE--DVEEDEEEDGEGENEEEIEMNRVKRRMDETLRHGGTLAEESEVGDTAALH- 432
E+ G E ++EED+++D EN++ E +K M+ +G EESE A+
Sbjct: 265 EDKKGDELKEIEEDKKDDHISENDKMDEDTEIKETMEGEPENG---KEESEKPVVDAMEV 95
Query: 433 ----DEYKNTQQEGQAAVVQQQKKHKEPQDK 459
E + +++ + VV ++ + E +DK
Sbjct: 94 EGGIKEKEESKENEKVEVVMEEDEKDEDEDK 2
>TC224263 weakly similar to UP|Q6GQH4 (Q6GQH4) Egr1 protein, partial (8%)
Length = 790
Score = 38.9 bits (89), Expect = 0.005
Identities = 23/80 (28%), Positives = 38/80 (46%), Gaps = 2/80 (2%)
Frame = -2
Query: 350 METSEKNCSPSDGRHEVNEVDDDGMEEENHGPEDVEEDEEEDGEGENEEEIEMNRVKR-- 407
+E S + DG+ + D+D E++ ED ++D +ED E EE E + R
Sbjct: 678 LEVSVRLKEAHDGKEDEQGSDNDENEDDEDEDEDSDDDYDEDSGSEEYEETEEEFLNRYA 499
Query: 408 RMDETLRHGGTLAEESEVGD 427
+ E L +G + EE + D
Sbjct: 498 KAAEALENGSAIIEEGDDED 439
>TC204091 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(70%)
Length = 2075
Score = 38.9 bits (89), Expect = 0.005
Identities = 20/63 (31%), Positives = 35/63 (54%)
Frame = +2
Query: 349 KMETSEKNCSPSDGRHEVNEVDDDGMEEENHGPEDVEEDEEEDGEGENEEEIEMNRVKRR 408
K + + P + EV+++ +EEE E+ EE++EE+ E E EEE E + VK++
Sbjct: 86 KQQQPKPENPPQEYEEVEEEVEEEEVEEEVEEEEEEEEEDEEEEEEEEEEEEENDVVKQQ 265
Query: 409 MDE 411
+
Sbjct: 266 QQQ 274
Score = 38.5 bits (88), Expect = 0.007
Identities = 35/133 (26%), Positives = 56/133 (41%)
Frame = +2
Query: 348 TKMETSEKNCSPSDGRHEVNEVDDDGMEEENHGPEDVEEDEEEDGEGENEEEIEMNRVKR 407
T+ ++ P + E EV+++ EEE + EE+EEE+ E E EEE
Sbjct: 71 TEPPLKQQQPKPENPPQEYEEVEEEVEEEEVEEEVEEEEEEEEEDEEEEEEE-------- 226
Query: 408 RMDETLRHGGTLAEESEVGDTAALHDEYKNTQQEGQAAVVQQQKKHKEPQDKHQKAKLLE 467
EE E D VV+QQ++ ++ +D +KLLE
Sbjct: 227 ------------EEEEEEND------------------VVKQQQQQQQDEDDIPISKLLE 316
Query: 468 SLCSENPMLSSLC 480
L ++ +L+ LC
Sbjct: 317 PL-GKDQLLNLLC 352
Score = 35.4 bits (80), Expect = 0.059
Identities = 21/68 (30%), Positives = 35/68 (50%)
Frame = +2
Query: 344 KGEVTKMETSEKNCSPSDGRHEVNEVDDDGMEEENHGPEDVEEDEEEDGEGENEEEIEMN 403
K + K E + + E EV+++ EEE ED EE+EEE+ E E E ++
Sbjct: 86 KQQQPKPENPPQEYEEVEEEVEEEEVEEEVEEEEEEEEEDEEEEEEEE-EEEEENDVVKQ 262
Query: 404 RVKRRMDE 411
+ +++ DE
Sbjct: 263 QQQQQQDE 286
Score = 31.2 bits (69), Expect = 1.1
Identities = 20/48 (41%), Positives = 24/48 (49%), Gaps = 5/48 (10%)
Frame = +2
Query: 26 EQEEEEEEEEETSAFDVGSKEDNNSGMEVDTPSS-----TGTDQHSNL 68
++EEEEEEEEE DV ++ E D P S G DQ NL
Sbjct: 203 DEEEEEEEEEEEEENDVVKQQQQQQQDEDDIPISKLLEPLGKDQLLNL 346
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.305 0.127 0.356
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,866,996
Number of Sequences: 63676
Number of extensions: 247623
Number of successful extensions: 3072
Number of sequences better than 10.0: 220
Number of HSP's better than 10.0 without gapping: 1665
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2289
length of query: 496
length of database: 12,639,632
effective HSP length: 101
effective length of query: 395
effective length of database: 6,208,356
effective search space: 2452300620
effective search space used: 2452300620
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC140550.8


