

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140549.8 - phase: 0
(122 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
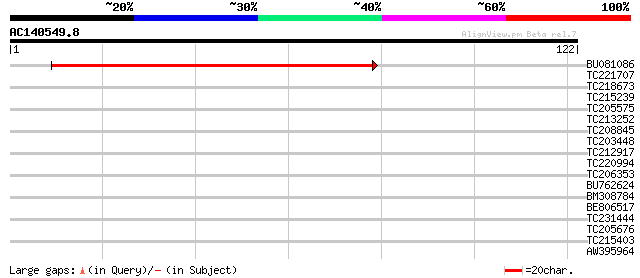
Score E
Sequences producing significant alignments: (bits) Value
BU081086 weakly similar to GP|27817864|db OJ1081_B12.13 {Oryza s... 84 9e-18
TC221707 similar to UP|Q9LR84 (Q9LR84) F21B7.1, partial (16%) 31 0.091
TC218673 similar to GB|AAL87403.1|19548077|AY081833 At2g20690/F5... 28 0.59
TC215239 UP|H3_PEA (P02300) Histone H3, partial (39%) 27 1.3
TC205575 weakly similar to UP|ZDH3_MOUSE (Q8R173) Zinc finger DH... 27 1.3
TC213252 26 2.9
TC208845 25 3.8
TC203448 similar to UP|Q6NKU3 (Q6NKU3) At1g55340, partial (48%) 25 5.0
TC212917 weakly similar to UP|Q9LX27 (Q9LX27) Calmodulin-like pr... 25 5.0
TC220994 similar to GB|AAO39958.1|28372952|BT003730 At1g08850 {A... 25 5.0
TC206353 25 5.0
BU762624 25 6.5
BM308784 25 6.5
BE806517 24 8.5
TC231444 24 8.5
TC205676 similar to GB|AAL87371.1|19548015|AY081718 AT4g33920/F1... 24 8.5
TC215403 similar to UP|Q40473 (Q40473) PS60 protein precursor, p... 24 8.5
AW395964 24 8.5
>BU081086 weakly similar to GP|27817864|db OJ1081_B12.13 {Oryza sativa
(japonica cultivar-group)}, partial (7%)
Length = 426
Score = 84.0 bits (206), Expect = 9e-18
Identities = 39/70 (55%), Positives = 52/70 (73%)
Frame = +2
Query: 10 LEGRVISGSNIDEKVFIPRLSLTPSDNRIPFKFKRRQFPISVSFAMIINKSQGQSLEHVG 69
+E +++SG V+IPRL+ +PS + PFK RR+FPI VS+AM INKSQGQ L VG
Sbjct: 215 IEAKIMSGKGQGNTVYIPRLATSPSQSPWPFKLIRRKFPIIVSYAMTINKSQGQLLASVG 394
Query: 70 VYLPSPIFSH 79
+YLP+P+FSH
Sbjct: 395 LYLPTPVFSH 424
>TC221707 similar to UP|Q9LR84 (Q9LR84) F21B7.1, partial (16%)
Length = 739
Score = 30.8 bits (68), Expect = 0.091
Identities = 23/82 (28%), Positives = 41/82 (49%), Gaps = 1/82 (1%)
Frame = +1
Query: 16 SGSNIDEKVFIPRLSLTPSDNRIPFKFKRRQFPISVSFAMIINKSQGQSLEHVGVYLPS- 74
S +N+ KVF+P ++ F S+S + + S+ + LEH+ VY PS
Sbjct: 43 SSANVTGKVFVPSGAIAAI------------FHNSLSHSQQLVNSKAKPLEHILVYTPSG 186
Query: 75 PIFSHGQLYVAISQVTSRGGLK 96
+F H +L +++ T+ GL+
Sbjct: 187 HVFQH-ELLASVALGTTDNGLR 249
>TC218673 similar to GB|AAL87403.1|19548077|AY081833 At2g20690/F5H14.34
{Arabidopsis thaliana;} , partial (72%)
Length = 1118
Score = 28.1 bits (61), Expect = 0.59
Identities = 15/34 (44%), Positives = 19/34 (55%)
Frame = +1
Query: 5 GGTYQLEGRVISGSNIDEKVFIPRLSLTPSDNRI 38
GGTY E V + NI + P L+LTP+ RI
Sbjct: 67 GGTYFTEKSVPTQHNIAKMTISPSLTLTPTTPRI 168
>TC215239 UP|H3_PEA (P02300) Histone H3, partial (39%)
Length = 340
Score = 26.9 bits (58), Expect = 1.3
Identities = 11/18 (61%), Positives = 14/18 (77%)
Frame = -3
Query: 47 FPISVSFAMIINKSQGQS 64
FP+ V FAM NKS+GQ+
Sbjct: 326 FPLIVCFAMTTNKSEGQT 273
>TC205575 weakly similar to UP|ZDH3_MOUSE (Q8R173) Zinc finger DHHC domain
containing protein 3 (Golgi-specific DHHC zinc finger
protein), partial (16%)
Length = 1563
Score = 26.9 bits (58), Expect = 1.3
Identities = 17/59 (28%), Positives = 25/59 (41%)
Frame = -1
Query: 24 VFIPRLSLTPSDNRIPFKFKRRQFPISVSFAMIINKSQGQSLEHVGVYLPSPIFSHGQL 82
V P +TP R P K QFP + AM+ + + G+ V + +S G L
Sbjct: 273 VTFPSFKVTPQAKRDPNVMKNHQFPRGNNMAMLRHDNLGEGKRSVFLRREVRAWSAGSL 97
>TC213252
Length = 689
Score = 25.8 bits (55), Expect = 2.9
Identities = 17/63 (26%), Positives = 32/63 (49%), Gaps = 3/63 (4%)
Frame = -3
Query: 41 KFKRRQFPISVSFAMIINKSQGQSLEHVGVYLPSPIFSHGQLYVAISQVTSRG---GLKI 97
K P++ +F +I + S+++ P+P S G++ +SQ+TS G K+
Sbjct: 378 KMGNNSMPLAPTFQYVIWTT*ISSIKNWESLQPTPSPSMGEIIQDVSQITSSSIS*GPKM 199
Query: 98 LIN 100
L+N
Sbjct: 198 LLN 190
>TC208845
Length = 766
Score = 25.4 bits (54), Expect = 3.8
Identities = 19/81 (23%), Positives = 35/81 (42%), Gaps = 12/81 (14%)
Frame = -3
Query: 19 NIDEKVFIPRLSLTPSDNRIPFKFKRRQFPISVSFAMII------------NKSQGQSLE 66
NI+ F L + + R+ +K + + +++ F ++ N S Q
Sbjct: 464 NINSSNFFCSLFIIKGNCRLSYKMIQNEARLNLPFLHLMHASSHLLK*RASNSSDAQMKL 285
Query: 67 HVGVYLPSPIFSHGQLYVAIS 87
+V +Y SP +GQ VAI+
Sbjct: 284 YVHLYFRSPHEDYGQYLVAIN 222
>TC203448 similar to UP|Q6NKU3 (Q6NKU3) At1g55340, partial (48%)
Length = 1257
Score = 25.0 bits (53), Expect = 5.0
Identities = 9/13 (69%), Positives = 11/13 (84%)
Frame = +1
Query: 96 KILINDDDDDDID 108
K ++NDDDDDD D
Sbjct: 982 KAILNDDDDDDED 1020
>TC212917 weakly similar to UP|Q9LX27 (Q9LX27) Calmodulin-like protein,
partial (49%)
Length = 1086
Score = 25.0 bits (53), Expect = 5.0
Identities = 11/27 (40%), Positives = 18/27 (65%)
Frame = -1
Query: 91 SRGGLKILINDDDDDDIDVASNVVYRE 117
S GG+K I+DDDDDD+ + ++ +
Sbjct: 969 SGGGVK-RIHDDDDDDVAIIFTYIHTQ 892
>TC220994 similar to GB|AAO39958.1|28372952|BT003730 At1g08850 {Arabidopsis
thaliana;} , partial (36%)
Length = 471
Score = 25.0 bits (53), Expect = 5.0
Identities = 12/34 (35%), Positives = 21/34 (61%)
Frame = +1
Query: 70 VYLPSPIFSHGQLYVAISQVTSRGGLKILINDDD 103
+Y+PS F HGQ ++A S G++ ++DD+
Sbjct: 247 IYVPSSFFHHGQAHMA---PRSFFGVEDFLDDDN 339
>TC206353
Length = 883
Score = 25.0 bits (53), Expect = 5.0
Identities = 7/24 (29%), Positives = 18/24 (74%)
Frame = -2
Query: 58 NKSQGQSLEHVGVYLPSPIFSHGQ 81
++ +G++++ +G +LP P+ +H Q
Sbjct: 534 HQGRGKTVDPLGFHLPPPLLAHSQ 463
>BU762624
Length = 408
Score = 24.6 bits (52), Expect = 6.5
Identities = 11/23 (47%), Positives = 15/23 (64%)
Frame = +3
Query: 19 NIDEKVFIPRLSLTPSDNRIPFK 41
NI V +++LTPS NRI F+
Sbjct: 270 NIKNSVMSVKVNLTPSQNRIFFR 338
>BM308784
Length = 408
Score = 24.6 bits (52), Expect = 6.5
Identities = 9/19 (47%), Positives = 14/19 (73%)
Frame = +1
Query: 27 PRLSLTPSDNRIPFKFKRR 45
P L+ +P+ R+PF+F RR
Sbjct: 211 PPLTASPTATRLPFRFLRR 267
>BE806517
Length = 427
Score = 24.3 bits (51), Expect = 8.5
Identities = 12/39 (30%), Positives = 21/39 (53%), Gaps = 4/39 (10%)
Frame = +2
Query: 21 DEKVFIPRLSLTPSDNRIPFKFK----RRQFPISVSFAM 55
D +FIP+++ NR+ +FK + FP+ S+ M
Sbjct: 26 DLLIFIPKMNKASVSNRLRIRFKGIILLKSFPVC*SWVM 142
>TC231444
Length = 966
Score = 24.3 bits (51), Expect = 8.5
Identities = 9/20 (45%), Positives = 14/20 (70%)
Frame = +3
Query: 88 QVTSRGGLKILINDDDDDDI 107
Q+T + ++ +DDDDDDI
Sbjct: 768 QMTDSASIFVI*SDDDDDDI 827
>TC205676 similar to GB|AAL87371.1|19548015|AY081718 AT4g33920/F17I5_110
{Arabidopsis thaliana;} , partial (81%)
Length = 1793
Score = 24.3 bits (51), Expect = 8.5
Identities = 8/12 (66%), Positives = 11/12 (91%)
Frame = -2
Query: 95 LKILINDDDDDD 106
++I +NDDDDDD
Sbjct: 103 IRIYLNDDDDDD 68
>TC215403 similar to UP|Q40473 (Q40473) PS60 protein precursor, partial (57%)
Length = 1090
Score = 24.3 bits (51), Expect = 8.5
Identities = 11/20 (55%), Positives = 14/20 (70%)
Frame = -3
Query: 56 IINKSQGQSLEHVGVYLPSP 75
I+ K Q SLE VG+ LP+P
Sbjct: 908 IVWKPQCNSLEQVGLQLPAP 849
>AW395964
Length = 308
Score = 24.3 bits (51), Expect = 8.5
Identities = 10/16 (62%), Positives = 12/16 (74%)
Frame = +3
Query: 93 GGLKILINDDDDDDID 108
G L LIN+DDD DI+
Sbjct: 195 GKLNWLINEDDDSDIE 242
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.140 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,209,241
Number of Sequences: 63676
Number of extensions: 49723
Number of successful extensions: 376
Number of sequences better than 10.0: 37
Number of HSP's better than 10.0 without gapping: 346
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 365
length of query: 122
length of database: 12,639,632
effective HSP length: 98
effective length of query: 24
effective length of database: 6,399,384
effective search space: 153585216
effective search space used: 153585216
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 51 (24.3 bits)
Medicago: description of AC140549.8


