

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140549.19 - phase: 0
(373 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
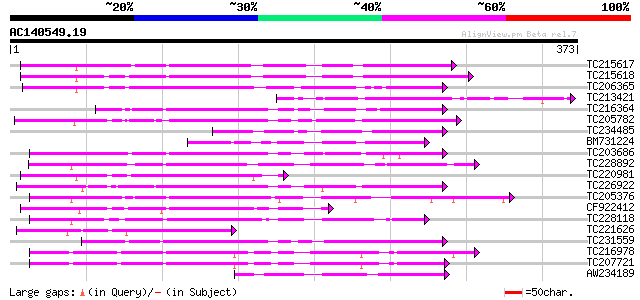
Score E
Sequences producing significant alignments: (bits) Value
TC215617 similar to UP|Q75PK5 (Q75PK5) Mitogen-activated kinase ... 196 1e-50
TC215618 similar to UP|Q75PK5 (Q75PK5) Mitogen-activated kinase ... 196 2e-50
TC206365 similar to UP|Q93ZH4 (Q93ZH4) AT5g66850/MUD21_11, parti... 176 2e-44
TC213421 similar to UP|M3K3_ARATH (O22042) Mitogen-activated pro... 164 5e-41
TC216364 similar to UP|Q9MAI7 (Q9MAI7) F12M16.4, partial (58%) 134 9e-32
TC205782 similar to UP|Q93WR8 (Q93WR8) MEK map kinase kinsae, pa... 130 7e-31
TC234485 similar to UP|Q6UY78 (Q6UY78) YDA, partial (16%) 129 2e-30
BM731224 weakly similar to PIR|T02550|T025 NPK1-related protein ... 119 2e-27
TC203686 similar to UP|Q93WR7 (Q93WR7) MAP kinase kinase, complete 116 2e-26
TC228892 UP|Q9XF25 (Q9XF25) SNF-1-like serine/threonine protein ... 110 1e-24
TC220981 similar to UP|Q93ZH4 (Q93ZH4) AT5g66850/MUD21_11, parti... 109 2e-24
TC226922 similar to UP|Q9LPQ3 (Q9LPQ3) F15H18.14, partial (54%) 108 4e-24
TC205376 similar to UP|Q9MB45 (Q9MB45) IRE homolog 1 (Fragment),... 107 7e-24
CF922412 106 1e-23
TC228118 similar to UP|KP19_ARATH (Q39030) Serine/threonine-prot... 103 1e-22
TC221626 weakly similar to UP|O23190 (O23190) MAP3K-like protein... 101 5e-22
TC231559 homologue to UP|Q9AYN9 (Q9AYN9) NQK1 MAPKK, partial (71%) 100 1e-21
TC216978 homologue to UP|Q8RXH5 (Q8RXH5) Osmotic stress-activate... 100 2e-21
TC207721 UP|Q43466 (Q43466) Protein kinase 3 , complete 100 2e-21
AW234189 99 4e-21
>TC215617 similar to UP|Q75PK5 (Q75PK5) Mitogen-activated kinase kinase
kinase alpha, partial (51%)
Length = 1451
Score = 196 bits (498), Expect = 1e-50
Identities = 110/291 (37%), Positives = 167/291 (56%), Gaps = 4/291 (1%)
Frame = +3
Query: 8 SEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAH----SEAGRDALENEVNILNTLNSS 63
S+W KGK++G G+FG V+L N +G L +K+ ++ ++ L+ ++ L+
Sbjct: 648 SKWKKGKLLGRGTFGHVYLGFNSDSGQLCAIKEVRVVCDDQSSKECLKQLNQEIHLLSQL 827
Query: 64 PSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLY 123
P IVQ G+D + L V++EY+SGGS+ + ++G + E V++ YTRQIV GL
Sbjct: 828 SHPNIVQYYGSDLGEET--LSVYLEYVSGGSIHKLLQEYG-AFKEPVIQNYTRQIVSGLS 998
Query: 124 HLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMA 183
+LH VH D+K N+L+ +G +KLADFG AK + SSS +++ G+P WMA
Sbjct: 999 YLHGRNTVHRDIKGANILVDPNGEIKLADFGMAKHI-------NSSSSMLSFKGSPYWMA 1157
Query: 184 PEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFK 243
PEV++ N S+ DIWSLGCT++EMAT +PPW + AA+FK
Sbjct: 1158PEVVMNTNGYSL----------PVDIWSLGCTILEMATSKPPW------NQYEGAAAIFK 1289
Query: 244 IACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLT 294
I +P+ P H S + +F++ CL RDP R TA +L+ HPF+ + T
Sbjct: 1290IGNSRDMPEIPDHLSSDAKNFIQLCLQRDPSARPTAQKLIEHPFIRDQSAT 1442
>TC215618 similar to UP|Q75PK5 (Q75PK5) Mitogen-activated kinase kinase
kinase alpha, partial (73%)
Length = 2291
Score = 196 bits (497), Expect = 2e-50
Identities = 116/305 (38%), Positives = 171/305 (56%), Gaps = 7/305 (2%)
Frame = +1
Query: 8 SEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAH-------SEAGRDALENEVNILNTL 60
S+W KGK++G G+FG V+L N G + +K+ S+ L E+N+LN L
Sbjct: 610 SKWRKGKLLGRGTFGHVYLGFNSENGQMCAIKEVKVVFDDHTSKECLKQLNQEINLLNQL 789
Query: 61 NSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVH 120
+ P IVQ G++ + L V++EY+SGGS+ + ++G E V++ YTRQIV
Sbjct: 790 SH---PNIVQYHGSEL--VEESLSVYLEYVSGGSIHKLLQEYG-PFKEPVIQNYTRQIVS 951
Query: 121 GLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPL 180
GL +LH VH D+K N+L+ +G +KLADFG AK + SS+ +++ G+P
Sbjct: 952 GLAYLHGRNTVHRDIKGANILVDPNGEIKLADFGMAKHI-------NSSASMLSFKGSPY 1110
Query: 181 WMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAA 240
WMAPEV++ N S+ DIWSLGCT+IEMAT +PPW + +AA
Sbjct: 1111WMAPEVVMNTNGYSL----------PVDIWSLGCTIIEMATSKPPW------NQYEGVAA 1242
Query: 241 MFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTHHKHYS 300
+FKI +P+ P H S + F++ CL RDP R TA +LL+HPF+ + T + S
Sbjct: 1243IFKIGNSKDMPEIPEHLSNDAKKFIKLCLQRDPLARPTAQKLLDHPFIRDQSATKAANVS 1422
Query: 301 ASSPA 305
+ A
Sbjct: 1423ITRDA 1437
>TC206365 similar to UP|Q93ZH4 (Q93ZH4) AT5g66850/MUD21_11, partial (37%)
Length = 1866
Score = 176 bits (445), Expect = 2e-44
Identities = 112/287 (39%), Positives = 161/287 (56%), Gaps = 7/287 (2%)
Frame = +1
Query: 9 EWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAH-------SEAGRDALENEVNILNTLN 61
+W KGK++G GS+GSV+ A N TG +K+ S LE E+ IL L+
Sbjct: 433 QWQKGKLIGRGSYGSVYHATNLETGASCALKEVDLFPDDPKSADCIKQLEQEIRILRQLH 612
Query: 62 SSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHG 121
P IVQ G++ ++L+++MEY+ GSL H+ G++ E VVR +TR I+ G
Sbjct: 613 H---PNIVQYYGSEI--VGDRLYIYMEYVHPGSLHKFMHEHCGAMTESVVRNFTRHILSG 777
Query: 122 LYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLW 181
L +LH +H D+K N+L+ +SG+VKLADFG +K + + S ++ G+P W
Sbjct: 778 LAYLHGTKTIHRDIKGANLLVDASGSVKLADFGVSKILTE-------KSYELSLKGSPYW 936
Query: 182 MAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAM 241
MAPE++ ++I ES A DIWSLGCT+IEM TG+PPW + P AM
Sbjct: 937 MAPELM----KAAIKKESSPDIAMAIDIWSLGCTIIEMLTGKPPWSE-----FEGPQ-AM 1086
Query: 242 FKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
FK+ P P S EG DFL++C R+P +R +A LL H F+
Sbjct: 1087FKVL--HKSPDLPESLSSEGQDFLQQCFRRNPAERPSAAVLLTHAFV 1221
>TC213421 similar to UP|M3K3_ARATH (O22042) Mitogen-activated protein kinase
kinase kinase 3 (Arabidospsis NPK1-related protein
kinase 3) , partial (6%)
Length = 724
Score = 164 bits (416), Expect = 5e-41
Identities = 105/202 (51%), Positives = 121/202 (58%), Gaps = 5/202 (2%)
Frame = +1
Query: 176 GGTPLWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISIS 235
GGTPLWMAPEVL +N S +DFAA DIWSLGCTVIEMATG PPW +S
Sbjct: 22 GGTPLWMAPEVL--RNES--------LDFAA-DIWSLGCTVIEMATGTPPWAHQ----VS 156
Query: 236 NPMAAMFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTH 295
NP A+ IA G GIP FP HFS+EG DFL RC R P KR T +LL HPF+VST
Sbjct: 157 NPTTAVLMIAHGHGIPHFPPHFSKEGLDFLTRCFERHPNKRPTVQDLLTHPFIVST---- 324
Query: 296 HKHYSASSPASVLEVHQFEDTYDDDDDDDDSDDNDELISPAGGNHFFITKELLC-----P 350
K Y ASSP SVLEV F+D+ DD+ + SD GNHF IT
Sbjct: 325 -KQY-ASSPTSVLEVQNFKDS--DDELETCSDQ---------GNHFSITNTTFAFHDDDL 465
Query: 351 QGTTMWQLEDSASGNHWITIRS 372
+G + + ED +SGN WIT+RS
Sbjct: 466 KGIPICKPEDISSGN-WITVRS 528
>TC216364 similar to UP|Q9MAI7 (Q9MAI7) F12M16.4, partial (58%)
Length = 2224
Score = 134 bits (336), Expect = 9e-32
Identities = 85/232 (36%), Positives = 130/232 (55%)
Frame = +1
Query: 57 LNTLNSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTR 116
++ L+ PYI + G+ Y +Q +L + MEYM+GGS+AD+ G L+E + R
Sbjct: 1 ISVLSQCRCPYITEYYGS-YLNQ-TKLWIIMEYMAGGSVADLIQS-GPPLDEMSIACILR 171
Query: 117 QIVHGLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANG 176
++H + +LH G +H D+K N+LL+ +G+VK+ADFG + ++ + ++ K+
Sbjct: 172 DLLHAVDYLHSEGKIHRDIKAANILLSENGDVKVADFGVSAQLTRTISRRKTFV------ 333
Query: 177 GTPLWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISN 236
GTP WMAPEV+ +N N++ ADIWSLG T IEMA G PP D +
Sbjct: 334 GTPFWMAPEVI--QNTDGYNEK--------ADIWSLGITAIEMAKGEPPLAD------LH 465
Query: 237 PMAAMFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
PM +F I + PQ HFS+ +F+ CL + P +R +A ELL F+
Sbjct: 466 PMRVLF-IIPRENPPQLDDHFSRPLKEFVSLCLKKVPAERPSAKELLKDRFI 618
>TC205782 similar to UP|Q93WR8 (Q93WR8) MEK map kinase kinsae, partial (91%)
Length = 1377
Score = 130 bits (328), Expect = 7e-31
Identities = 95/297 (31%), Positives = 154/297 (50%), Gaps = 3/297 (1%)
Frame = +2
Query: 4 IIKGSEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKA---HSEAGRDALENEVNILNTL 60
+I SE + +G GS G+V+ +++++G ++ +K H E+ R + E+ IL +
Sbjct: 263 VIPFSELERLNRIGSGSGGTVYKVVHRTSGRVYALKVIYGHHEESVRRQIHREIQILRDV 442
Query: 61 NSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVH 120
+ + +V+C YD Q++++ V +E+M GGSL + H E + +RQI+
Sbjct: 443 DDAN---VVKCHEM-YD-QNSEIQVLLEFMDGGSL-EGKH----ITQEQQLADLSRQILR 592
Query: 121 GLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPL 180
GL +LH+ IVH D+K N+L+ S VK+ADFG + + + M+ SS GT
Sbjct: 593 GLAYLHRRHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSV------GTIA 754
Query: 181 WMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAA 240
+M+PE + N+ IND D A DIWS G +++E GR P+ + A+
Sbjct: 755 YMSPE----RINTDINDGQ--YDAYAGDIWSFGVSILEFYMGRFPFA----VGRQGDWAS 904
Query: 241 MFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTHHK 297
+ C P+ P S DF+ RCL RDP +R +A LL HPF+ H++
Sbjct: 905 LMCAICMSQPPEAPPSASPHFKDFILRCLQRDPSRRWSASRLLEHPFIAPPLPNHNQ 1075
>TC234485 similar to UP|Q6UY78 (Q6UY78) YDA, partial (16%)
Length = 543
Score = 129 bits (324), Expect = 2e-30
Identities = 72/155 (46%), Positives = 93/155 (59%)
Frame = +1
Query: 134 DLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVLLMKNNS 193
D+K N+L+ +G VKLADFG AK + SC ++ GTP WMAPEV+ KN++
Sbjct: 61 DIKGANILVDPTGRVKLADFGMAKHIT-------GQSCPLSFKGTPYWMAPEVI--KNSN 213
Query: 194 SINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKIACGDGIPQF 253
N A DIWSLGCTV+EMAT +PPW + + AAMFKI +P
Sbjct: 214 GCN--------LAVDIWSLGCTVLEMATTKPPWFQYEGV------AAMFKIGNSKELPTI 351
Query: 254 PIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
P H S EG DF+R+CL R+P R +A ELL+HPF+
Sbjct: 352 PDHLSNEGKDFVRKCLQRNPHDRPSASELLDHPFV 456
>BM731224 weakly similar to PIR|T02550|T025 NPK1-related protein kinase
homolog T26B15.7 - Arabidopsis thaliana, partial (23%)
Length = 407
Score = 119 bits (298), Expect = 2e-27
Identities = 63/159 (39%), Positives = 92/159 (57%)
Frame = +3
Query: 118 IVHGLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGG 177
++ GL +LH G+VHCD+K N+L+ G K+ DFGCAK N SS+ I GG
Sbjct: 3 VLQGLQYLHNKGVVHCDIKGGNILIGEDG-AKIGDFGCAKFA------NDSSAVI---GG 152
Query: 178 TPLWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNP 237
TP++MAPEV + AD+W+LGCTV+EMATG PW + + +P
Sbjct: 153 TPMFMAPEVARGEEQGY-----------PADVWALGCTVLEMATGFAPWPN-----VEDP 284
Query: 238 MAAMFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKR 276
+ ++++A D +P+ P S+E DFL +C R+PK+R
Sbjct: 285 VTVLYRVAYSDDVPEIPCFLSEEAKDFLGKCFRRNPKER 401
>TC203686 similar to UP|Q93WR7 (Q93WR7) MAP kinase kinase, complete
Length = 1502
Score = 116 bits (290), Expect = 2e-26
Identities = 88/280 (31%), Positives = 135/280 (47%), Gaps = 5/280 (1%)
Frame = +2
Query: 14 KMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGRDALENEVNILNTLNSSPSPYIVQCLG 73
K+VG G+ G V L +K T F +K + L + PY+V C
Sbjct: 308 KVVGKGNGGVVQLVQHKWTSQFFALKVIQMNIEESMRKQIAQELKINQQAQCPYVVVCYQ 487
Query: 74 TDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYHLH-QHGIVH 132
+ Y++ + + +EYM GGSLAD+ K ++ ED + +Q++ GL +LH + I+H
Sbjct: 488 SFYEN--GVISIILEYMDGGSLADLLKKVK-TIPEDYLAAICKQVLKGLVYLHHEKHIIH 658
Query: 133 CDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVLLMKNN 192
DLK N+L+ G VK+ DFG + ++ + GT +M+PE
Sbjct: 659 RDLKPSNLLINHIGEVKITDFGVSAIMESTSGQANTFI------GTYNYMSPE------- 799
Query: 193 SSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKI--ACGDGI 250
IN R ++ + DIWSLG ++E A GR P+ D S ++F++ D
Sbjct: 800 -RINGSQRGYNYKS-DIWSLGLILLECALGRFPYAPPDQ---SETWESIFELIETIVDKP 964
Query: 251 PQFPI--HFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
P P FS E F+ CL +DPK R +A EL+ HPF+
Sbjct: 965 PPIPPSEQFSTEFCSFISACLQKDPKDRLSAQELMAHPFV 1084
>TC228892 UP|Q9XF25 (Q9XF25) SNF-1-like serine/threonine protein kinase,
complete
Length = 1931
Score = 110 bits (274), Expect = 1e-24
Identities = 88/302 (29%), Positives = 138/302 (45%), Gaps = 5/302 (1%)
Frame = +1
Query: 13 GKMVGCGSFGSVHLAMNKSTGGLFVVK-----KAHSEAGRDALENEVNILNTLNSSPSPY 67
GK +G GSFG V +A + TG +K K + + + E+ IL
Sbjct: 145 GKTLGIGSFGKVKIAEHVRTGHKVAIKILNRHKIKNMEMEEKVRREIKILRLFMHHHIIR 324
Query: 68 IVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYHLHQ 127
+ + + T D ++V MEY+ G L D + G L ED R + +QI+ G+ + H+
Sbjct: 325 LYEVVETPTD-----IYVVMEYVKSGELFDYIVE-KGRLQEDEARHFFQQIISGVEYCHR 486
Query: 128 HGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVL 187
+ +VH DLK +N+LL S N+K+ADFG + N+ + + + G+P + APEV+
Sbjct: 487 NMVVHRDLKPENLLLDSKFNIKIADFGLS-------NIMRDGHFLKTSCGSPNYAAPEVI 645
Query: 188 LMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKIACG 247
++ D+WS G + + G P+ D+++ + +FK G
Sbjct: 646 ----------SGKLYAGPEVDVWSCGVILYALLCGTLPFDDENIPN-------LFKKIKG 774
Query: 248 DGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTHHKHYSASSPASV 307
GI P H S D + R LV DP KR T E+ HP+ H Y A P
Sbjct: 775 -GIYTLPSHLSPGARDLIPRMLVVDPMKRMTIPEIRQHPWF----QVHLPRYLAVPPPDT 939
Query: 308 LE 309
L+
Sbjct: 940 LQ 945
>TC220981 similar to UP|Q93ZH4 (Q93ZH4) AT5g66850/MUD21_11, partial (27%)
Length = 646
Score = 109 bits (273), Expect = 2e-24
Identities = 66/186 (35%), Positives = 107/186 (57%), Gaps = 10/186 (5%)
Frame = +1
Query: 8 SEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAH-------SEAGRDALENEVNILNTL 60
S+W KGK++G G+FGSV++A N+ TG L +K+ S LE E+ +L+ L
Sbjct: 133 SQWKKGKLIGRGTFGSVYVATNRETGALCAMKEVELFPDDPKSAECIKQLEQEIKVLSNL 312
Query: 61 NSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVH 120
S IVQ G++ + +++L++++EY GS+ + G++ E V+R +TR I+
Sbjct: 313 KHSN---IVQYYGSEIE--EDRLYIYLEYDHPGSINKYVREHCGAITESVIRNFTRHILS 477
Query: 121 GLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRV---KKNMNMNKSSSCIIANGG 177
GL +LH +H D+K N+L+ S+G VKLADFG AK + + N+++ G
Sbjct: 478 GLAYLHSKKTIHRDIKGANLLVDSAGVVKLADFGMAKHLTAFEANLSLR----------G 627
Query: 178 TPLWMA 183
+P WMA
Sbjct: 628 SPYWMA 645
>TC226922 similar to UP|Q9LPQ3 (Q9LPQ3) F15H18.14, partial (54%)
Length = 1190
Score = 108 bits (270), Expect = 4e-24
Identities = 83/290 (28%), Positives = 144/290 (49%), Gaps = 6/290 (2%)
Frame = +1
Query: 5 IKGSEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAG----RDALENEVNILNTL 60
I ++ K ++G G+ G+V+ +K+T + +K HS+ R AL +E +IL +
Sbjct: 202 IAAADLEKLAILGHGNGGTVYKVRHKATSATYALKIIHSDTDATRRRRAL-SETSILRRV 378
Query: 61 NSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVH 120
P ++V+ + ++ + + MEYM GG+L + + G+ +E+ + R ++
Sbjct: 379 TDCP--HVVR-FHSSFEKPSGDVAILMEYMDGGTL-ETALAASGTFSEERLAKVARDVLE 546
Query: 121 GLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPL 180
GL +LH I H D+K N+L+ S G+VK+ADFG +K + +++ S GT
Sbjct: 547 GLAYLHARNIAHRDIKPANILVNSEGDVKIADFGVSKLMCRSLEACNSYV------GTCA 708
Query: 181 WMAPEVLLMKNNSSINDESRVVDF--AAADIWSLGCTVIEMATGRPPWVDDDLISISNPM 238
+M+P+ + E+ ++ AADIWSLG T+ E+ G P++
Sbjct: 709 YMSPD--------RFDPEAYGGNYNGFAADIWSLGLTLFELYVGHFPFLQ---AGQRPDW 855
Query: 239 AAMFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
A + C P P S E DF+ CL ++ +R T +LL HPF+
Sbjct: 856 ATLMCAICFGDPPSLPETASPEFRDFVECCLKKESGERWTTAQLLTHPFV 1005
>TC205376 similar to UP|Q9MB45 (Q9MB45) IRE homolog 1 (Fragment), partial
(45%)
Length = 2377
Score = 107 bits (268), Expect = 7e-24
Identities = 101/370 (27%), Positives = 160/370 (42%), Gaps = 51/370 (13%)
Frame = +2
Query: 14 KMVGCGSFGSVHLAMNKSTGGLFVVK--KAHSEAGRDALENEVNILNTLNSSPSPYIVQC 71
K + G+FG V LA ++TG LF +K K ++A+E+ + + L + +P++V+
Sbjct: 299 KPISRGAFGRVFLAKKRTTGDLFAIKVLKKADMIRKNAVESILAERDILITVRNPFVVRF 478
Query: 72 LGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYHLHQHGIV 131
+ + ++N L++ MEY++GG L + G L+E+V RVY ++V L +LH +V
Sbjct: 479 FYS-FTCREN-LYLVMEYLNGGDLYSLLRNLG-CLDEEVARVYIAEVVLALEYLHSLRVV 649
Query: 132 HCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANG--------------- 176
H DLK N+L+A G++KL DFG +K N + S + NG
Sbjct: 650 HRDLKPDNLLIAHDGHIKLTDFGLSKVGLINSTDDLSGPAV--NGTSLLEEDETDVFTSA 823
Query: 177 ------------GTPLWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRP 224
GTP ++APE+LL + AD WS+G + E+ G P
Sbjct: 824 DQRERREKRSAVGTPDYLAPEILLGTGHG-----------FTADWWSVGVILFELLVGIP 970
Query: 225 PW-------VDDDLISISNPMAAMFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKR- 276
P+ + D++++ P P P S + D + R L DP +R
Sbjct: 971 PFNAEHPQTIFDNILNRKIPW------------PAVPEEMSPQAQDLIDRLLTEDPNQRL 1114
Query: 277 --STALELLNHPFLVS---TTLTHHKHYSASSPASVLEVHQFEDTYDDDDDD-------- 323
A E+ H F TL K + S L+ F Y + D
Sbjct: 1115GSKGASEVKQHVFFKDINWDTLARQKAAFVPASESALDTSYFTSRYSWNTSDGFVYPASD 1294
Query: 324 -DDSDDNDEL 332
+DS D D L
Sbjct: 1295VEDSSDADSL 1324
>CF922412
Length = 701
Score = 106 bits (265), Expect = 1e-23
Identities = 71/212 (33%), Positives = 113/212 (52%), Gaps = 6/212 (2%)
Frame = -2
Query: 8 SEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSE----AGRDALENEVNILNTLNSS 63
+++M G +G G++G V+ ++ G +K+ E + + E+++L LN
Sbjct: 568 NKYMLGDEIGKGAYGRVYKGLDLENGDFVAIKQVSLENIAQEDLNIIMQEIDLLKNLNHK 389
Query: 64 PSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADV--SHKFGGSLNEDVVRVYTRQIVHG 121
IV+ LG+ + LH+ +EY+ GSLA++ +KFG E +V VY Q++ G
Sbjct: 388 N---IVKYLGSS--KTKSHLHIVLEYVENGSLANIIKPNKFG-PFPESLVAVYIAQVLEG 227
Query: 122 LYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLW 181
L +LH+ G++H D+K N+L G VKLADFG A ++ + ++N S GTP W
Sbjct: 226 LVYLHEQGVIHRDIKGANILTTKEGLVKLADFGVATKLTE-ADVNTHSVV-----GTPYW 65
Query: 182 MAPEVLLMKNNSSINDESRVVDFAAADIWSLG 213
MAPEV+ M AA+DIWS+G
Sbjct: 64 MAPEVIEMAGVC-----------AASDIWSVG 2
>TC228118 similar to UP|KP19_ARATH (Q39030) Serine/threonine-protein kinase
AtPK19 (Ribosomal-protein S6 kinase homolog) , partial
(65%)
Length = 1305
Score = 103 bits (258), Expect = 1e-22
Identities = 81/270 (30%), Positives = 134/270 (49%), Gaps = 7/270 (2%)
Frame = +1
Query: 14 KMVGCGSFGSVHLAMNKSTGGLFVVK-----KAHSEAGRDALENEVNILNTLNSSPSPYI 68
K+VG G+F V+ K T ++ +K K + + ++ E +I + P++
Sbjct: 178 KVVGQGAFAKVYQVRKKDTSEIYAMKVMRKDKIMEKNHAEYMKAERDIWTKIEH---PFV 348
Query: 69 VQCLGTDYDHQDN-QLHVFMEYMSGGSLA-DVSHKFGGSLNEDVVRVYTRQIVHGLYHLH 126
VQ Y Q +L++ +++++GG L + H+ G ED+ R+YT +IV + HLH
Sbjct: 349 VQLR---YSFQTKYRLYLVLDFVNGGHLFFQLYHQ--GLFREDLARIYTAEIVSAVSHLH 513
Query: 127 QHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEV 186
+GI+H DLK +N+LL + G+V L DFG AK+ +++ N S C GT +MAPE+
Sbjct: 514 ANGIMHRDLKPENILLDADGHVMLTDFGLAKQFEESTRSN--SMC-----GTLEYMAPEI 672
Query: 187 LLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKIAC 246
+L K + AAD WS+G + E TG+PP+ + I +
Sbjct: 673 ILGKGHDK-----------AADWWSVGVLLFETLTGKPPFCGGNRDKIQQKIVK------ 801
Query: 247 GDGIPQFPIHFSQEGFDFLRRCLVRDPKKR 276
D I + P S E L+ L ++ +R
Sbjct: 802 -DKI-KLPAFLSSEAHSLLKGLLQKEQARR 885
>TC221626 weakly similar to UP|O23190 (O23190) MAP3K-like protein kinase,
partial (10%)
Length = 542
Score = 101 bits (252), Expect = 5e-22
Identities = 58/151 (38%), Positives = 87/151 (57%), Gaps = 6/151 (3%)
Frame = +2
Query: 5 IKGSEWMKGKMVGCGSFGSVHLAMNKSTGGLF----VVKKAHSEAGRDALENEVNILNTL 60
I +W++G VG GSF +V LA+ + F VVK A ++ L NE ++L+ L
Sbjct: 98 IPNMDWVRGDAVGRGSFATVSLAIPTTNYNQFPSLTVVKSADAQTSC-WLRNEKHVLDRL 274
Query: 61 NSSPSPYIVQCLGTD--YDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQI 118
S P I++C G D +++ ++F+EY +GGSLAD G + E R YTR I
Sbjct: 275 GSCPR--IIRCFGDDCSFENGVEYYNLFLEYAAGGSLADELRNHDGRIPEPQAREYTRDI 448
Query: 119 VHGLYHLHQHGIVHCDLKCKNVLLASSGNVK 149
V GL H+H++G VHCD+K +N+L+ G +K
Sbjct: 449 VEGLSHVHKNGFVHCDIKLQNILVFEDGRIK 541
>TC231559 homologue to UP|Q9AYN9 (Q9AYN9) NQK1 MAPKK, partial (71%)
Length = 867
Score = 100 bits (248), Expect = 1e-21
Identities = 76/243 (31%), Positives = 122/243 (49%), Gaps = 2/243 (0%)
Frame = +2
Query: 48 DALENEVNILNTLNSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLN 107
D + V L +S P++V C + Y + + + +EYM GSLADV K ++
Sbjct: 26 DIRKQIVQELKINQASQCPHVVVCYHSFY--HNGVISLVLEYMDRGSLADVI-KQVKTIL 196
Query: 108 EDVVRVYTRQIVHGLYHLH-QHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMN 166
E + V +Q++ GL +LH + ++H D+K N+L+ G VK+ DFG + + +M
Sbjct: 197 EPYLAVVFKQVLQGLVYLHNERHVIHRDIKPSNLLVNHKGEVKITDFGVSAMLASSMGQR 376
Query: 167 KSSSCIIANGGTPLWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPW 226
+ GT +M+PE + + S D S +DIWSLG V+E A GR P+
Sbjct: 377 DTFV------GTYNYMSPE----RISGSTYDYS-------SDIWSLGMVVLECAIGRFPY 505
Query: 227 V-DDDLISISNPMAAMFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNH 285
+ +D S + + I P FS E F+ C+ +DP+ R T+L+LL+H
Sbjct: 506 IQSEDQQSWPSFYELLAAIVESPPPSAPPDQFSPEFCSFVSSCIQKDPRDRLTSLKLLDH 685
Query: 286 PFL 288
PF+
Sbjct: 686 PFI 694
>TC216978 homologue to UP|Q8RXH5 (Q8RXH5) Osmotic stress-activated protein
kinase, partial (96%)
Length = 1626
Score = 99.8 bits (247), Expect = 2e-21
Identities = 89/309 (28%), Positives = 138/309 (43%), Gaps = 13/309 (4%)
Frame = +1
Query: 14 KMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGRDALENEVNILNTLNSSPSPYIVQCLG 73
K +G G+FG L +K T L +K + E G EN + S P I++
Sbjct: 376 KDIGSGNFGVARLMRHKDTKELVAMK--YIERGHKIDENVAREIINHRSLRHPNIIRF-- 543
Query: 74 TDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYHLHQHGIVHC 133
+ L + MEY +GG L + G +ED R + +Q++ G+ + H I H
Sbjct: 544 KEVVLTPTHLGIVMEYAAGGELFERICS-AGRFSEDEARYFFQQLISGVSYCHSMQICHR 720
Query: 134 DLKCKNVLLASSG--NVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVLLMKN 191
DLK +N LL S +K+ DFG +K ++ ++ S + GTP ++APEVL
Sbjct: 721 DLKLENTLLDGSPAPRLKICDFGYSK---SSLLHSRPKSTV----GTPAYIAPEVL---- 867
Query: 192 NSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDD-----LISISNPMAAMFKIAC 246
R D AD+WS G T+ M G P+ D D +I+ MA +K
Sbjct: 868 ------SRREYDGKLADVWSCGVTLYVMLVGAYPFEDQDDPKNFRKTINRIMAVQYK--- 1020
Query: 247 GDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVS------TTLTHHKHYS 300
IP + +H SQ+ L R V +P +R T E+ +HP+ + T + +Y
Sbjct: 1021---IPDY-VHISQDCRHLLSRIFVANPARRITIKEIKSHPWFLKNLPRELTEVAQAAYYR 1188
Query: 301 ASSPASVLE 309
+P L+
Sbjct: 1189KENPTFSLQ 1215
>TC207721 UP|Q43466 (Q43466) Protein kinase 3 , complete
Length = 1416
Score = 99.8 bits (247), Expect = 2e-21
Identities = 85/283 (30%), Positives = 131/283 (46%), Gaps = 7/283 (2%)
Frame = +3
Query: 14 KMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGRDALENEVNILNTLNSSPSPYIVQCLG 73
K +G G+FG L NK T L +K + E G+ EN + S P I++
Sbjct: 147 KDLGAGNFGVARLMRNKETKELVAMK--YIERGQKIDENVAREIINHRSLRHPNIIRF-- 314
Query: 74 TDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYHLHQHGIVHC 133
+ L + MEY +GG L + G +ED R + +Q++ G+++ H I H
Sbjct: 315 KEVVLTPTHLAIVMEYAAGGELFERICN-AGRFSEDEARYFFQQLISGVHYCHAMQICHR 491
Query: 134 DLKCKNVLLASSG--NVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVLLMKN 191
DLK +N LL S +K+ DFG +K ++ ++ S + GTP ++APEVL
Sbjct: 492 DLKLENTLLDGSPAPRLKICDFGYSK---SSLLHSRPKSTV----GTPAYIAPEVL---- 638
Query: 192 NSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDD-----LISISNPMAAMFKIAC 246
R D AD+WS G T+ M G P+ D D +I MA +K
Sbjct: 639 ------SRREYDGKLADVWSCGVTLYVMLVGAYPFEDQDDPRNFRKTIQRIMAVQYK--- 791
Query: 247 GDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLV 289
IP + +H SQ+ L R V +P +R + E+ +HP+ +
Sbjct: 792 ---IPDY-VHISQDCRHLLSRIFVANPLRRISLKEIKSHPWFL 908
>AW234189
Length = 406
Score = 98.6 bits (244), Expect = 4e-21
Identities = 53/141 (37%), Positives = 80/141 (56%)
Frame = +1
Query: 149 KLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVLLMKNNSSINDESRVVDFAAAD 208
KLADFG A ++ S + ++ G+P+WMAPEV+ + +D
Sbjct: 16 KLADFGAAVEIES------SPAMLLFPRGSPMWMAPEVVRRERQGP-----------ESD 144
Query: 209 IWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKIACGDGIPQFPIHFSQEGFDFLRRC 268
+WSLGCTVIE+A G+P W D + ++S +I D +P+FPI S+ G DFL +C
Sbjct: 145 VWSLGCTVIEIAIGKPAWEDRGVDTLS-------RIGYSDELPEFPIQLSELGKDFLEKC 303
Query: 269 LVRDPKKRSTALELLNHPFLV 289
L R+P +R + +LL HP+L+
Sbjct: 304 LRREPSERWSCDQLLQHPYLL 366
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.135 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,058,816
Number of Sequences: 63676
Number of extensions: 286934
Number of successful extensions: 4248
Number of sequences better than 10.0: 723
Number of HSP's better than 10.0 without gapping: 2601
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3279
length of query: 373
length of database: 12,639,632
effective HSP length: 99
effective length of query: 274
effective length of database: 6,335,708
effective search space: 1735983992
effective search space used: 1735983992
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC140549.19


