

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140544.4 - phase: 0 /pseudo
(305 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
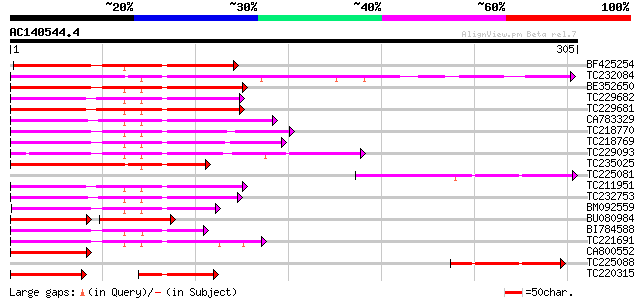
Score E
Sequences producing significant alignments: (bits) Value
BF425254 homologue to PIR|S26605|S26 myb-related protein 1 - gar... 134 7e-32
TC232084 UP|Q84PP2 (Q84PP2) Transcription factor MYB101 (Fragmen... 126 1e-29
BE352650 116 1e-26
TC229682 UP|Q9XIU9 (Q9XIU9) GmMYB29A2 protein, complete 110 6e-25
TC229681 UP|Q9S7E3 (Q9S7E3) GmMYB29A1 protein, complete 109 2e-24
CA783329 homologue to GP|6651292|gb| myb-related transcription f... 107 7e-24
TC218770 UP|Q9XIU5 (Q9XIU5) GmMYB29B2 protein, complete 104 4e-23
TC218769 UP|Q9XIU8 (Q9XIU8) GmMYB29A2 protein, complete 103 7e-23
TC229093 homologue to PIR|T51621|T51621 myb-like protein [import... 102 2e-22
TC235025 similar to UP|Q6R077 (Q6R077) MYB transcription factor,... 100 1e-21
TC225081 similar to UP|Q6R098 (Q6R098) MYB transcription factor,... 100 1e-21
TC211951 similar to GB|AAS10022.1|41619094|AY519552 MYB transcri... 96 2e-20
TC232753 GmMYB10 92 2e-19
BM092559 homologue to GP|13346186|gb GHMYB10 {Gossypium hirsutum... 87 1e-17
BU080984 62 1e-15
BI784588 homologue to GP|18419456|gb blind {Lycopersicon esculen... 78 4e-15
TC221691 GmMYB6 78 4e-15
CA800552 GP|5139814|dbj GmMYB29B2 {Glycine max}, partial (16%) 77 7e-15
TC225088 similar to UP|Q6WAT8 (Q6WAT8) Lipid tranfer protein II,... 77 1e-14
TC220315 similar to UP|Q8L5N8 (Q8L5N8) Myb-related transcription... 56 1e-14
>BF425254 homologue to PIR|S26605|S26 myb-related protein 1 - garden petunia,
partial (29%)
Length = 375
Score = 134 bits (336), Expect = 7e-32
Identities = 74/131 (56%), Positives = 87/131 (65%), Gaps = 10/131 (7%)
Frame = +3
Query: 3 RSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL-- 60
RSPCC+KVGLKKGPWT EEDQKLL+YIEEHGHGSWR+LP KA G+ R L
Sbjct: 3 RSPCCDKVGLKKGPWTPEEDQKLLAYIEEHGHGSWRALPAKA-----GLQRCGKSCRLRW 167
Query: 61 -------II*GRI-LRGESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLKKR 112
I G+ ++ E + + + L N + + +T LPKRTDNEIKNYWNTHLKK
Sbjct: 168 TNYLRPDIKRGKFXMQEEQTIIQLHALLGN--RWSSISTHLPKRTDNEIKNYWNTHLKKX 341
Query: 113 LTKMGIDPVTH 123
L KM IDPVTH
Sbjct: 342 LNKMSIDPVTH 374
>TC232084 UP|Q84PP2 (Q84PP2) Transcription factor MYB101 (Fragment), complete
Length = 1383
Score = 126 bits (317), Expect = 1e-29
Identities = 105/330 (31%), Positives = 161/330 (47%), Gaps = 26/330 (7%)
Frame = +2
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGR+PC ++ GLKKGPWT EED+ L+ YI++HGHGSWR+LP A + G +
Sbjct: 53 MGRTPCSDENGLKKGPWTPEEDRILVDYIQKHGHGSWRALPKHARLNRCGKSCRLRWTNY 232
Query: 61 II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLKKRLT 114
+ I RG E L+ + L N + A A LP RTDNEIKN+WNTHLKK+L
Sbjct: 233 LR-PDIKRGKFSEEEEQLIINLHAVLGN--KWSAIAGHLPGRTDNEIKNFWNTHLKKKLL 403
Query: 115 KMGIDPVTHKPKNETL-LPEN------HSKNVANLSHKAQWESA-RLEAEARLVRESKLR 166
+MG+DPVTH+P+++ L L N + V+NL++ +A RL+++A + + +L
Sbjct: 404 QMGLDPVTHRPRSDHLNLLSNLQQLLAATNIVSNLTNTWDTNNALRLQSDATQLAKLQLL 583
Query: 167 SQLQHQFG-----SNTSVFPSQSSSSSS------NQVLNIKEEG-EKEWKGYEDSTHLLE 214
+ H G SN + SSSS N+VL + + + + GY
Sbjct: 584 QNILHVLGDTTPTSNLDLLNPFGPSSSSLPDGFLNEVLGLNQSKLQNLYNGYSAQN---- 751
Query: 215 FKDLMENSSMAFSSTLQHEMTMINVAEEGFTNLLLDNSNSGDLSLSPESGGECNTCDGSG 274
+ + +F Q ++ + N +D + + PE G E
Sbjct: 752 -----QPNLQSFEPPHQQQV-------QPTGNQHMDGGSIDSSCMKPEKGDE-------- 871
Query: 275 SGGGSDFNEDNKNYWNNILNLVNSSPSDSS 304
D + +N++ NLV++SP S+
Sbjct: 872 ---QLDVPYSSSTPFNSLPNLVSASPECST 952
>BE352650
Length = 528
Score = 116 bits (290), Expect = 1e-26
Identities = 66/139 (47%), Positives = 85/139 (60%), Gaps = 11/139 (7%)
Frame = +1
Query: 1 MGRSPCCEKVG-LKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DG 59
MGRSPCCE+ +KKGPWT EED+KL+ YI +HGHGSWR+LP +A G+ R
Sbjct: 103 MGRSPCCEESSSVKKGPWTPEEDEKLIDYISKHGHGSWRTLPKRA-----GLNRCGKSCR 267
Query: 60 L----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHL 109
L + I RG E ++ + L N + A LP RTDNEIKNYW TH+
Sbjct: 268 LRWTNYLRPDIKRGKFSEDDERIIINFHSVLGN--KWSKIAAHLPGRTDNEIKNYWTTHI 441
Query: 110 KKRLTKMGIDPVTHKPKNE 128
+K+L KMGIDP T+KP+ +
Sbjct: 442 RKKLLKMGIDPETYKPRTD 498
>TC229682 UP|Q9XIU9 (Q9XIU9) GmMYB29A2 protein, complete
Length = 1249
Score = 110 bits (276), Expect = 6e-25
Identities = 65/136 (47%), Positives = 82/136 (59%), Gaps = 10/136 (7%)
Frame = +3
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
M R+PCCEK+GLKKGPWT EEDQ L+SYI++HGHG+WR+LP L G+ R L
Sbjct: 84 MVRAPCCEKMGLKKGPWTPEEDQILMSYIQKHGHGNWRALP-----KLAGLLRCGKSCRL 248
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG E ++ K ++ L N + A A +LP RTDNEIKN W+THLK
Sbjct: 249 RWINYLRPDIKRGNFSSEEEEIIIKMHELLGN--RWSAIAAKLPGRTDNEIKNVWHTHLK 422
Query: 111 KRLTKMGIDPVTHKPK 126
KRL I KP+
Sbjct: 423 KRLLNSDIQKRVSKPR 470
>TC229681 UP|Q9S7E3 (Q9S7E3) GmMYB29A1 protein, complete
Length = 795
Score = 109 bits (272), Expect = 2e-24
Identities = 64/136 (47%), Positives = 82/136 (60%), Gaps = 10/136 (7%)
Frame = +1
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
M R+PCCEK+GLKKGPWT EEDQ L+SYI++HGHG+WR+LP L G+ R L
Sbjct: 1 MVRAPCCEKMGLKKGPWTPEEDQILMSYIQKHGHGNWRALP-----KLAGLLRCGKSCRL 165
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG E ++ K ++ L N + A A +LP RTDNEIKN W+THLK
Sbjct: 166 RWINYLRPDIKRGNFSSEEEEIIIKMHELLGN--RWSAIAAKLPGRTDNEIKNVWHTHLK 339
Query: 111 KRLTKMGIDPVTHKPK 126
KRL + KP+
Sbjct: 340 KRLMNSDTNKRVSKPR 387
>CA783329 homologue to GP|6651292|gb| myb-related transcription factor
{Pimpinella brachycarpa}, partial (48%)
Length = 443
Score = 107 bits (267), Expect = 7e-24
Identities = 63/154 (40%), Positives = 88/154 (56%), Gaps = 10/154 (6%)
Frame = +3
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGR P C+K+G+KKGPWT+EED+KL+++I +G WR++P L G+ R L
Sbjct: 3 MGRQPRCDKLGVKKGPWTAEEDKKLINFILTNGQCCWRAVPK-----LAGLKRCGKSCRL 167
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ + RG E LV + L N + A RLP RTDNEIKN+WNTH+K
Sbjct: 168 RWTNYLRPDLKRGLLTEAEEQLVIDLHARLGN--RWSKIAARLPGRTDNEIKNHWNTHIK 341
Query: 111 KRLTKMGIDPVTHKPKNETLLPENHSKNVANLSH 144
K+L K+GIDPVTH+P + + + + H
Sbjct: 342 KKLLKIGIDPVTHEPLKKQAAASSQDSSSSPAEH 443
>TC218770 UP|Q9XIU5 (Q9XIU5) GmMYB29B2 protein, complete
Length = 1316
Score = 104 bits (260), Expect = 4e-23
Identities = 69/163 (42%), Positives = 91/163 (55%), Gaps = 10/163 (6%)
Frame = +1
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
M R+PCCEK+GLKKGPW +EEDQ L SYI++HGHG+WR+LP +A G+ R L
Sbjct: 112 MVRAPCCEKMGLKKGPWATEEDQILTSYIQKHGHGNWRALPKQA-----GLLRCGKSCRL 276
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG E + K ++ L N + A A +LP RTDNEIKN W+T+LK
Sbjct: 277 RWINYLRPDIKRGNFTIEEEETIIKLHEMLGN--RWSAIAAKLPGRTDNEIKNVWHTNLK 450
Query: 111 KRLTKMGIDPVTHKPKNETLLPENHSKNVANLSHKAQWESARL 153
KRL K KP ++ ++ +N S Q E A L
Sbjct: 451 KRLLKSD----QSKPSSKRATKPKIKRSDSNSSIITQSEPAHL 567
>TC218769 UP|Q9XIU8 (Q9XIU8) GmMYB29A2 protein, complete
Length = 828
Score = 103 bits (258), Expect = 7e-23
Identities = 68/159 (42%), Positives = 89/159 (55%), Gaps = 10/159 (6%)
Frame = +1
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
M R+PCCEK+GLKKGPW EEDQ L SYI++HGHG+WR+LP +A G+ R L
Sbjct: 1 MVRAPCCEKMGLKKGPWAPEEDQILTSYIDKHGHGNWRALPKQA-----GLLRCGKSCRL 165
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG E + K + L N + A A +LP RTDNEIKN W+T+LK
Sbjct: 166 RWINYLRPDIKRGNFTIEEEETIIKLHDMLGN--RWSAIAAKLPGRTDNEIKNVWHTNLK 339
Query: 111 KRLTKMGIDPVTHKPKNETLLPENHSKNVANLSHKAQWE 149
KRL K D KP ++ + ++ +N S Q E
Sbjct: 340 KRLLKS--DQSKSKPSSKRAIKPKIERSDSNSSIITQSE 450
>TC229093 homologue to PIR|T51621|T51621 myb-like protein [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(44%)
Length = 964
Score = 102 bits (255), Expect = 2e-22
Identities = 71/205 (34%), Positives = 112/205 (54%), Gaps = 14/205 (6%)
Frame = +3
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGR PCC+K G+KKGPWT EED L+SYI+EHG G+W+++P G++R + L
Sbjct: 153 MGRPPCCDK-GVKKGPWTPEEDIILVSYIQEHGPGNWKAVPANT-----GLSRCSKSCRL 314
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG E ++ L N + A A LP+RTDN+IKNYWNT+LK
Sbjct: 315 RWTNYLRPGIKRGNFTDQEEKMIIHLQALLGN--RWAAIAAYLPQRTDNDIKNYWNTYLK 488
Query: 111 KRLTKMGIDPVTHKPKNETLLPENHS----KNVANLSHKAQWESARLEAEARLVRESKLR 166
K+L K + +LL H+ +V+ + QWE RL+ + + +++
Sbjct: 489 KKLNK----KLEAGSDQGSLLGGGHNNIGFSSVSQPMSRGQWE-RRLQTDIHMAKKALSE 653
Query: 167 SQLQHQFGSNTSVFPSQSSSSSSNQ 191
+ ++ S++S ++S+S+ SN+
Sbjct: 654 ALSPNKASSSSSTLLAESNSNLSNE 728
>TC235025 similar to UP|Q6R077 (Q6R077) MYB transcription factor, partial
(35%)
Length = 434
Score = 100 bits (248), Expect = 1e-21
Identities = 57/115 (49%), Positives = 71/115 (61%), Gaps = 7/115 (6%)
Frame = +1
Query: 1 MGRSPCCEK-VGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DG 59
MGRSPCCE+ VG+KKGPWT EED+KL+ YI +HGHGSWR+LP +A + G +
Sbjct: 97 MGRSPCCEENVGVKKGPWTPEEDEKLIDYISKHGHGSWRTLPKRAGLNRCGKSCRLRWTN 276
Query: 60 LII*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTH 108
+ I RG E ++ + L N + AT LP RTDNEIKNYWNTH
Sbjct: 277 YLT-PDIKRGKFSEEDERIIINLHSVLGN--KWSKIATHLPGRTDNEIKNYWNTH 432
>TC225081 similar to UP|Q6R098 (Q6R098) MYB transcription factor, partial
(6%)
Length = 801
Score = 100 bits (248), Expect = 1e-21
Identities = 60/123 (48%), Positives = 70/123 (56%), Gaps = 4/123 (3%)
Frame = +3
Query: 187 SSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEMTMIN----VAEE 242
+S N +KEE E+EWK DS+ F + +M T E + + EE
Sbjct: 54 NSGNSSSLVKEEAEQEWKSTYDSSITTTFSSGLHEFTMNMEGTWASESLRTSGSHDIVEE 233
Query: 243 GFTNLLLDNSNSGDLSLSPESGGECNTCDGSGSGGGSDFNEDNKNYWNNILNLVNSSPSD 302
GFTNLLL +NS D SLS E GGE DG G G SDF EDN NYWN+ILNLVNSSPS
Sbjct: 234 GFTNLLL-KTNSDDPSLSSEDGGESKNGDGGG-GTNSDFYEDNNNYWNSILNLVNSSPSH 407
Query: 303 SSM 305
S M
Sbjct: 408 SPM 416
>TC211951 similar to GB|AAS10022.1|41619094|AY519552 MYB transcription factor
{Arabidopsis thaliana;} , partial (38%)
Length = 776
Score = 95.9 bits (237), Expect = 2e-20
Identities = 59/138 (42%), Positives = 79/138 (56%), Gaps = 10/138 (7%)
Frame = +2
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGR CC K L+KG W+ EED+KLL+YI +HGHG W S+P L G+ R L
Sbjct: 92 MGRHSCCYKQKLRKGLWSPEEDEKLLNYITKHGHGCWSSVP-----KLAGLQRCGKSCRL 256
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ + RG E+ + + + L N + A +LP RTDNEIKN WN+ LK
Sbjct: 257 RWINYLRPDLKRGAFSQQEENSIIELHAVLGN--RWSQIAAQLPGRTDNEIKNLWNSCLK 430
Query: 111 KRLTKMGIDPVTHKPKNE 128
K+L + GIDP TH+P +E
Sbjct: 431 KKLRQRGIDPNTHQPLSE 484
>TC232753 GmMYB10
Length = 423
Score = 92.4 bits (228), Expect = 2e-19
Identities = 58/135 (42%), Positives = 74/135 (53%), Gaps = 10/135 (7%)
Frame = +2
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGR CC K L+KG W+ EED+KL +YI G G W S+P L G+ R L
Sbjct: 32 MGRHSCCVKQKLRKGLWSPEEDEKLFNYITRFGVGCWSSVPK-----LAGLQRCGKSCRL 196
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ + RG E L+ ++ L N + A +LP RTDNEIKN+WN+ LK
Sbjct: 197 RWINYLRPDLKRGMFSQQEEDLIISLHEVLGN--RWAQIAAQLPGRTDNEIKNFWNSCLK 370
Query: 111 KRLTKMGIDPVTHKP 125
K+L K GIDP THKP
Sbjct: 371 KKLLKQGIDPSTHKP 415
>BM092559 homologue to GP|13346186|gb GHMYB10 {Gossypium hirsutum}, partial
(38%)
Length = 421
Score = 86.7 bits (213), Expect = 1e-17
Identities = 54/122 (44%), Positives = 68/122 (55%), Gaps = 10/122 (8%)
Frame = +2
Query: 2 GRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL- 60
GRSPCC K GL +G WT+ ED+ L YI HG G WR+LP +A G+ R L
Sbjct: 2 GRSPCCSKEGLNRGAWTAHEDKILREYIRVHGEGRWRNLPKRA-----GLKRCGKSCRLR 166
Query: 61 ---II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLKK 111
+ I RG E L+ + +K L N + A RLP RTDNEIKNYWNT+L K
Sbjct: 167 WLNYLRPDIKRGNISPDEEELIIRLHKLLGN--RWSLIAGRLPGRTDNEIKNYWNTNLGK 340
Query: 112 RL 113
++
Sbjct: 341 KV 346
>BU080984
Length = 420
Score = 62.0 bits (149), Expect(2) = 1e-15
Identities = 26/44 (59%), Positives = 32/44 (72%)
Frame = +2
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKA 44
MGR CC K L+KG W+ EED+KLL+YI +HGHG W S+P A
Sbjct: 137 MGRHSCCYKQKLRKGLWSPEEDEKLLNYITKHGHGCWSSVPKLA 268
Score = 38.1 bits (87), Expect(2) = 1e-15
Identities = 25/41 (60%), Positives = 25/41 (60%)
Frame = +3
Query: 49 RGVARAAD*DGLII*GRILRGESLVCKKNKPLFNFMHF*AT 89
RGVARAAD* G II G I R E KK PL NFM AT
Sbjct: 276 RGVARAAD*GG*IISGLI*REEHSHSKKRIPLLNFMQSLAT 398
>BI784588 homologue to GP|18419456|gb blind {Lycopersicon esculentum},
partial (36%)
Length = 407
Score = 78.2 bits (191), Expect = 4e-15
Identities = 48/118 (40%), Positives = 65/118 (54%), Gaps = 11/118 (9%)
Frame = +1
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHG-HGSWRSLPTKAVQDLRGVARAAD*DG 59
MGR+PCC+K +K+GPW+ EED KL YIE+HG G+W +LP KA G+ R
Sbjct: 73 MGRAPCCDKANVKRGPWSPEEDSKLKDYIEKHGTGGNWIALPQKA-----GLKRCGKSCR 237
Query: 60 L----II*GRILRGE------SLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNT 107
L + I GE ++C + + + A +LP RTDN+IKNYWNT
Sbjct: 238 LRWLNYLRPNIKHGEFSDEEDRIICSLYANIGS--RWSIIAAQLPGRTDNDIKNYWNT 405
>TC221691 GmMYB6
Length = 590
Score = 78.2 bits (191), Expect = 4e-15
Identities = 57/165 (34%), Positives = 81/165 (48%), Gaps = 27/165 (16%)
Frame = +3
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
M R+P C+K G +KG WT EED+KL++Y+ +G +WR LP A G+AR L
Sbjct: 78 MVRTPSCDKSGTRKGTWTPEEDRKLIAYVTRYGSWNWRQLPRFA-----GLARCGKSCRL 242
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ + RG + + + +K L N + A A LP RTDNEIKN+W+T LK
Sbjct: 243 RWMNYLRPNVKRGNFTQQEDECIIRMHKKLGN--KWSAIAAELPGRTDNEIKNHWHTTLK 416
Query: 111 K-----------RLTKMGIDPVTHK------PKNETLLPENHSKN 138
K T D V +K P N +L+ +N S +
Sbjct: 417 KWSQQNAITNEEARTSKSKDKVPNKGVTVTLPANSSLMSDNSSSS 551
>CA800552 GP|5139814|dbj GmMYB29B2 {Glycine max}, partial (16%)
Length = 434
Score = 77.4 bits (189), Expect = 7e-15
Identities = 31/44 (70%), Positives = 39/44 (88%)
Frame = +3
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKA 44
M R+PCCEK+GLKKGPW +EEDQ L SYI++HGHG+WR+LP +A
Sbjct: 111 MVRAPCCEKMGLKKGPWATEEDQILTSYIQKHGHGNWRALPKQA 242
>TC225088 similar to UP|Q6WAT8 (Q6WAT8) Lipid tranfer protein II, complete
Length = 1006
Score = 76.6 bits (187), Expect = 1e-14
Identities = 41/62 (66%), Positives = 45/62 (72%)
Frame = +1
Query: 238 NVAEEGFTNLLLDNSNSGDLSLSPESGGECNTCDGSGSGGGSDFNEDNKNYWNNILNLVN 297
++ EEGFTNLLL +NS D SLS E GGE DG G G SDF EDN NYWN+ILNLVN
Sbjct: 64 DIVEEGFTNLLL-KTNSDDPSLSSEDGGESKNGDGGG-GTNSDFYEDNNNYWNSILNLVN 237
Query: 298 SS 299
SS
Sbjct: 238 SS 243
>TC220315 similar to UP|Q8L5N8 (Q8L5N8) Myb-related transcription factor
VlMYBB1-1, partial (45%)
Length = 654
Score = 55.8 bits (133), Expect(2) = 1e-14
Identities = 21/41 (51%), Positives = 31/41 (75%)
Frame = +1
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLP 41
M R+P C+K G++KG WT EED+KL++Y+ +G +WR LP
Sbjct: 79 MVRTPSCDKSGMRKGTWTLEEDRKLIAYVTRYGSWNWRQLP 201
Score = 40.8 bits (94), Expect(2) = 1e-14
Identities = 21/43 (48%), Positives = 27/43 (61%)
Frame = +2
Query: 70 ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLKKR 112
E + + +K L N + A LP RTDNEIKN+W+T LKKR
Sbjct: 302 EEFIIRMHKKLGN--RWSTIAAELPGRTDNEIKNHWHTTLKKR 424
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.131 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,974,350
Number of Sequences: 63676
Number of extensions: 213845
Number of successful extensions: 1736
Number of sequences better than 10.0: 180
Number of HSP's better than 10.0 without gapping: 1634
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1716
length of query: 305
length of database: 12,639,632
effective HSP length: 97
effective length of query: 208
effective length of database: 6,463,060
effective search space: 1344316480
effective search space used: 1344316480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC140544.4


