

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
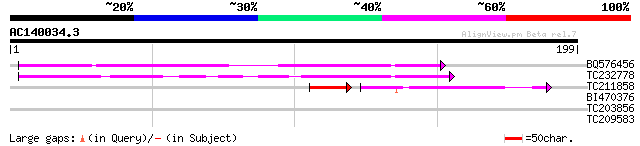
Query= AC140034.3 + phase: 0
(199 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BQ576456 69 1e-12
TC232778 similar to UP|Q9LNG5 (Q9LNG5) F21D18.16, partial (5%) 45 2e-05
TC211858 37 2e-04
BI470376 weakly similar to GP|14090338|dbj P0638D12.5 {Oryza sat... 28 3.6
TC203856 homologue to GB|AAG40366.1|11762176|AF325014 At2g01140 ... 27 4.8
TC209583 27 8.1
>BQ576456
Length = 424
Score = 69.3 bits (168), Expect = 1e-12
Identities = 49/150 (32%), Positives = 68/150 (44%)
Frame = +1
Query: 4 GEMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEYGGY 63
GE+T+TLDD ACL HLPI G + +E + E LLM L VS+ EA + + Y
Sbjct: 31 GELTITLDDMACLLHLPITGALHRFE-PLGVDEAVLLLMELLEVSREEARAKIVRAHRAY 207
Query: 64 ISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDRSNKR 123
+ LR+ Y S AR VR YL +LV C LF ++S
Sbjct: 208 VRLSWLREVYQSRC-----------------QARCWIVAVRAYLFHLVDCTLFANKSATH 336
Query: 124 IELIYLTTMADGYAGMRNYSWGGMTLAYLY 153
+ +++L D Y WG L ++Y
Sbjct: 337 VHVVHLKGF*D-LC*SGGYGWGVAALVHMY 423
>TC232778 similar to UP|Q9LNG5 (Q9LNG5) F21D18.16, partial (5%)
Length = 746
Score = 45.4 bits (106), Expect = 2e-05
Identities = 43/153 (28%), Positives = 66/153 (43%)
Frame = -1
Query: 4 GEMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEYGGY 63
GE T+TL D L L +G L + + A L LGV E E G
Sbjct: 422 GEATITLQDVLILLGLRTDGAPLIGSTNL---DWADLCEELLGVRPQEGEI-----EGSV 267
Query: 64 ISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDRSNKR 123
+ L Y N+ D +E+L R +R +++ +G ++F ++S+ R
Sbjct: 266 VKLSWL----AHYFSHINI-----DEGNVEQLQRF----IRAWILRFIGGVIFVNKSSSR 126
Query: 124 IELIYLTTMADGYAGMRNYSWGGMTLAYLYGEL 156
+ L YL + D + Y+WG LAYLY E+
Sbjct: 125 VSLRYLQFLRD-FE*CSTYAWGPAMLAYLYREM 30
>TC211858
Length = 524
Score = 36.6 bits (83), Expect(2) = 2e-04
Identities = 23/68 (33%), Positives = 35/68 (50%), Gaps = 1/68 (1%)
Frame = -1
Query: 124 IELIYLTTMAD-GYAGMRNYSWGGMTLAYLYGELADACRPGHRALGGSVTLLTVRKLKFY 182
+ ++YL D G +G Y+WG L ++Y +L +A R R + G +TLL
Sbjct: 470 VHVVYLDAFRDLGQSG--GYAWGVAALVHMYNQLDEASRTTTRQIVGYLTLL-------- 321
Query: 183 IYVNFIFL 190
VNF+FL
Sbjct: 320 -QVNFVFL 300
Score = 24.6 bits (52), Expect(2) = 2e-04
Identities = 9/15 (60%), Positives = 13/15 (86%)
Frame = -2
Query: 106 YLMYLVGCLLFDDRS 120
YL++LVGC LF ++S
Sbjct: 523 YLLHLVGCTLFANKS 479
>BI470376 weakly similar to GP|14090338|dbj P0638D12.5 {Oryza sativa
(japonica cultivar-group)}, partial (5%)
Length = 429
Score = 27.7 bits (60), Expect = 3.6
Identities = 11/23 (47%), Positives = 16/23 (68%)
Frame = -1
Query: 4 GEMTVTLDDFACLTHLPIEGRML 26
GE T+TL D + L +P++GR L
Sbjct: 273 GEATITLQDVSVLLGIPVDGRPL 205
>TC203856 homologue to GB|AAG40366.1|11762176|AF325014 At2g01140 {Arabidopsis
thaliana;} , partial (93%)
Length = 1641
Score = 27.3 bits (59), Expect = 4.8
Identities = 10/17 (58%), Positives = 15/17 (87%)
Frame = -1
Query: 177 RKLKFYIYVNFIFLLYF 193
R++K+Y VNF+FLL+F
Sbjct: 1542 RQIKYY*AVNFVFLLFF 1492
>TC209583
Length = 1349
Score = 26.6 bits (57), Expect = 8.1
Identities = 15/37 (40%), Positives = 20/37 (53%)
Frame = +2
Query: 70 RDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWY 106
RD YTS LG A LA ++DP E + + RW+
Sbjct: 851 RDDYTSVLGLAENLALSKDPCVKEFVKLLYMIMFRWF 961
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.142 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,272,732
Number of Sequences: 63676
Number of extensions: 126493
Number of successful extensions: 787
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 785
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 787
length of query: 199
length of database: 12,639,632
effective HSP length: 92
effective length of query: 107
effective length of database: 6,781,440
effective search space: 725614080
effective search space used: 725614080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC140034.3


