

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140031.2 + phase: 0 /pseudo
(484 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
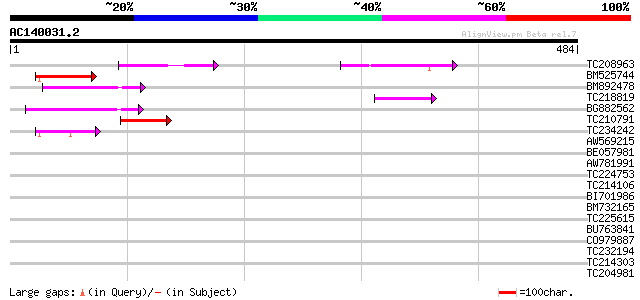
Score E
Sequences producing significant alignments: (bits) Value
TC208963 67 2e-11
BM525744 57 2e-08
BM892478 56 3e-08
TC218819 similar to UP|O03160 (O03160) NADH dehydrogenase subuni... 50 3e-06
BG882562 44 2e-04
TC210791 43 3e-04
TC234242 43 4e-04
AW569215 weakly similar to GP|4914326|gb| F14N23.12 {Arabidopsis... 41 0.001
BE057981 40 0.003
AW781991 37 0.015
TC224753 homologue to UP|Q09085 (Q09085) Hydroxyproline-rich gly... 34 0.17
TC214106 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 31 1.1
BI701986 31 1.1
BM732165 homologue to SP|Q9TV66|HCN4 Potassium/sodium hyperpolar... 31 1.1
TC225615 UP|Q84ZU7 (Q84ZU7) R 5 protein, complete 31 1.4
BU763841 similar to GP|28569968|dbj pherophorin {Volvox carteri ... 30 1.8
CO979887 30 2.4
TC232194 weakly similar to UP|Q9LSN6 (Q9LSN6) Arabidopsis thalia... 30 2.4
TC214303 similar to UP|Q9T0K5 (Q9T0K5) Extensin-like protein, pa... 30 2.4
TC204981 similar to UP|Q9LN36 (Q9LN36) F18O14.34, partial (71%) 30 3.1
>TC208963
Length = 1163
Score = 67.0 bits (162), Expect = 2e-11
Identities = 38/103 (36%), Positives = 58/103 (55%), Gaps = 3/103 (2%)
Frame = +1
Query: 283 FYSWLGGYMSRYALGNIVLEEERCRGTLCMDKEICGCV*RKSYGLPCACFVATKIRDDKP 342
FY L G++SR AL +I E +R + T +D ICGC+ R ++GLPCAC +A P
Sbjct: 673 FYVKLVGFVSRSALSHITEEYDRVK-TAGIDSSICGCIVRTTHGLPCACELARYSTMCHP 849
Query: 343 ILLDEIHLHWNRLCM---GQESNEDCFSVEEEWNGIQERLKKI 382
I L+ IH HW +L G N S++ E + + +R +++
Sbjct: 850 IPLEAIHAHWRKLKFSDHGTNDNGSELSLQPEVDALYKRFQEL 978
Score = 44.7 bits (104), Expect = 9e-05
Identities = 27/86 (31%), Positives = 43/86 (49%), Gaps = 1/86 (1%)
Frame = +1
Query: 94 NVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVKVRCILDCRYPMAKKDGKEVKH 153
++P+V+VT D LM+AV V P S + C FH+ +NVK +C K + H
Sbjct: 16 DMPQVIVTVGDIALMSAVQVVFPSSSNLLCRFHINQNVKAKC-------------KSIVH 156
Query: 154 -GDVVKKIMRVW*VIIESHPLKNYIQ 178
+ + +M W V++ S Y+Q
Sbjct: 157 LKEKQELMMDAWDVVVNSPNEGEYMQ 234
>BM525744
Length = 411
Score = 57.0 bits (136), Expect = 2e-08
Identities = 25/54 (46%), Positives = 39/54 (71%), Gaps = 2/54 (3%)
Frame = +3
Query: 23 HP--LSFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEK 74
HP + +F TVL +D+TY+ N Y++P+ EIVGVTST+LT+SV F ++ ++
Sbjct: 246 HPDAIKLLGSFHTVLFLDNTYKVNRYQLPLLEIVGVTSTELTFSVAFAYMESDE 407
>BM892478
Length = 429
Score = 56.2 bits (134), Expect = 3e-08
Identities = 26/88 (29%), Positives = 47/88 (52%)
Frame = +3
Query: 29 NNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCKL 88
++F VLV+D++Y+ +Y +P VGV +G + E EE+F W+ Q +
Sbjct: 150 SHFGDVLVLDTSYRKTVYLVPFATFVGVNHHKQPVLLGCALIADESEESFTWLFQTWLRA 329
Query: 89 LTSRMNVPKVVVTDKDTTLMNAVSNVLP 116
++ R+ P V+ D+D + A++ V P
Sbjct: 330 MSGRL--PLTVIADQDIAIQRAIAKVFP 407
>TC218819 similar to UP|O03160 (O03160) NADH dehydrogenase subunit 2, partial
(5%)
Length = 732
Score = 49.7 bits (117), Expect = 3e-06
Identities = 24/54 (44%), Positives = 31/54 (56%), Gaps = 1/54 (1%)
Frame = +1
Query: 312 MDKEICGCV*RKSYGLPCACFVATKIRDDKPILLDEIHLHWNRLCMG-QESNED 364
+D +CGC R ++GLPCAC +A R PI L IH+HW L QE N +
Sbjct: 34 IDGSLCGCTIRTTHGLPCACELAKYSRTWHPIPLQAIHVHWRTLNFSDQEMNNE 195
>BG882562
Length = 416
Score = 43.5 bits (101), Expect = 2e-04
Identities = 23/101 (22%), Positives = 49/101 (47%)
Frame = +3
Query: 14 LKTYFALIQHPLSFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHE 73
L+ F + + ++ F V+ DST +N Y++P+ VGV + +G + E
Sbjct: 120 LRNVFWIDSRSRAAYSYFGDVVAFDSTCLSNNYEIPLVAFVGVNHHGKSVLLGCGLLADE 299
Query: 74 KEENFVWVLQMMCKLLTSRMNVPKVVVTDKDTTLMNAVSNV 114
E ++W+ + +T R P+ ++T+ + +A++ V
Sbjct: 300 TFETYIWLFRAWLTCMTGR--PPQTIITNXCKAMQSAIAEV 416
>TC210791
Length = 456
Score = 43.1 bits (100), Expect = 3e-04
Identities = 19/44 (43%), Positives = 28/44 (63%)
Frame = +1
Query: 95 VPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVKVRCILD 138
+P V++T +D LM+AV V P S + C FH+ KNVK + +D
Sbjct: 28 MPPVIITVRDIALMDAVQVVFPSSSNLLCRFHISKNVKAKSQVD 159
>TC234242
Length = 478
Score = 42.7 bits (99), Expect = 4e-04
Identities = 22/62 (35%), Positives = 35/62 (55%), Gaps = 7/62 (11%)
Frame = +3
Query: 23 HP--LSFFNNFPTVLVMDSTYQTNMYKMPM-----FEIVGVTSTDLTYSVGFEFVTHEKE 75
HP + N V ++DSTY+TN Y++P+ + VG+T T +T+S F +V E+
Sbjct: 111 HPDSVKLVNACNLVFLIDSTYRTNRYRLPLLDFVVLDFVGMTPTGMTFSASFAYVEGERV 290
Query: 76 EN 77
N
Sbjct: 291 NN 296
>AW569215 weakly similar to GP|4914326|gb| F14N23.12 {Arabidopsis thaliana},
partial (19%)
Length = 425
Score = 41.2 bits (95), Expect = 0.001
Identities = 22/93 (23%), Positives = 45/93 (47%)
Frame = +1
Query: 35 LVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCKLLTSRMN 94
+V D+TY+ Y M + +GV + +T + E ++F W L+ + +
Sbjct: 10 VVFDTTYRVEAYDMLLGIWLGVDNNGMTCFFSCALLRDENIQSFSWALKAFLGFMKGK-- 183
Query: 95 VPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHV 127
P+ ++TD + L A++ LP++ C +H+
Sbjct: 184 APQTILTDHNMWLKEAIAVELPQTKHAFCIWHI 282
>BE057981
Length = 268
Score = 39.7 bits (91), Expect = 0.003
Identities = 15/44 (34%), Positives = 29/44 (65%)
Frame = +2
Query: 34 VLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEEN 77
V ++++ Y+TN Y++ + + GVT +T+SVGF ++ E+ N
Sbjct: 122 VFLINNAYKTNRYRLLLLDFAGVTPARMTFSVGFAYLEGERVNN 253
>AW781991
Length = 419
Score = 37.4 bits (85), Expect = 0.015
Identities = 16/53 (30%), Positives = 27/53 (50%)
Frame = +3
Query: 31 FPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQ 83
F + D+TY++N Y++P GV G F+ +E E +FVW+ +
Sbjct: 255 FGDTVTFDTTYRSNRYRLPFAFFTGVNHHGQPVLFGCAFLINESEASFVWLFK 413
>TC224753 homologue to UP|Q09085 (Q09085) Hydroxyproline-rich glycoprotein
(HRGP) (Fragment), partial (48%)
Length = 1088
Score = 33.9 bits (76), Expect = 0.17
Identities = 19/59 (32%), Positives = 30/59 (50%), Gaps = 1/59 (1%)
Frame = +3
Query: 414 HQLVESHLFGSVLIPKILIVYRHRHQQRHHFLKGNVLVLVKHHVPH-FHHLLGFQISLP 471
H ++ SHL +L +LI H H H ++ L+ H +PH HH++ I+LP
Sbjct: 180 HHIITSHLHHHLLHLHLLITTSHLHHHLPHLHLLTIINLLHHPLPHLLHHII---INLP 347
>TC214106 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (22%)
Length = 1296
Score = 31.2 bits (69), Expect = 1.1
Identities = 21/54 (38%), Positives = 26/54 (47%), Gaps = 3/54 (5%)
Frame = +2
Query: 414 HQLVESHLFGSVLIPKILIVYRHRHQQRHHFLKGNVLVL---VKHHVPHFHHLL 464
H L HLF IP +Y H HH L+ +L L + HH PH HHL+
Sbjct: 260 HHLHRRHLF----IP----IYHH-----HHLLQFTLLPLQSILHHHHPHLHHLV 382
Score = 30.0 bits (66), Expect = 2.4
Identities = 22/62 (35%), Positives = 25/62 (39%), Gaps = 7/62 (11%)
Frame = +2
Query: 414 HQLVESHLFGSVLIPKILIVYRHRHQQRHHFLKG-------NVLVLVKHHVPHFHHLLGF 466
H LV SHL L+ H H HH +K + L L HH H HHL F
Sbjct: 371 HHLV*SHLHRHHLV*N-----HHLHHHPHHHVKSIHHHHHHHTLHLTIHHHLHRHHLPRF 535
Query: 467 QI 468
I
Sbjct: 536 NI 541
>BI701986
Length = 422
Score = 31.2 bits (69), Expect = 1.1
Identities = 10/32 (31%), Positives = 21/32 (65%)
Frame = +2
Query: 424 SVLIPKILIVYRHRHQQRHHFLKGNVLVLVKH 455
S+L+P+++I++ H H HH + +L++H
Sbjct: 35 SLLLPRLMIIHHHHHHHHHHPRETLFSILLQH 130
>BM732165 homologue to SP|Q9TV66|HCN4 Potassium/sodium
hyperpolarization-activated cyclic nucleotide-gated
channel 4, partial (2%)
Length = 423
Score = 31.2 bits (69), Expect = 1.1
Identities = 9/33 (27%), Positives = 20/33 (60%)
Frame = +1
Query: 424 SVLIPKILIVYRHRHQQRHHFLKGNVLVLVKHH 456
S+L+P+++I++ H H HH + + ++ H
Sbjct: 1 SLLLPRLMIIHHHHHHHHHHHPRETLFSILLQH 99
>TC225615 UP|Q84ZU7 (Q84ZU7) R 5 protein, complete
Length = 2724
Score = 30.8 bits (68), Expect = 1.4
Identities = 24/99 (24%), Positives = 44/99 (44%), Gaps = 5/99 (5%)
Frame = +1
Query: 371 EWNGIQERLKKIPNRMNLEMKETKSTSQGSQEKSGYCEV--YRKGHQLVE-SHLFGSVLI 427
EW E K+IPN LE+ + + G ++K+ + ++ KG +L E H+ +
Sbjct: 1201 EWKSAVEHYKRIPNDEILEILKVSFDALGEEQKNVFLDIACCFKGCKLTEVEHMLRGL-- 1374
Query: 428 PKILIVYRHRHQQRHHF--LKGNVLVLVKHHVPHFHHLL 464
+ + +HH L L+ V+H + H L+
Sbjct: 1375 --------YNNCMKHHIDVLVDKSLIKVRHGTVNMHDLI 1467
>BU763841 similar to GP|28569968|dbj pherophorin {Volvox carteri f.
nagariensis}, partial (6%)
Length = 131
Score = 30.4 bits (67), Expect = 1.8
Identities = 15/38 (39%), Positives = 21/38 (54%), Gaps = 6/38 (15%)
Frame = -2
Query: 436 HRHQQRHHFLKG------NVLVLVKHHVPHFHHLLGFQ 467
H H H+ L G ++L+L HH+ H+ LLGFQ
Sbjct: 124 HHHHNLHYLLLGYHHPQHHLLLLRYHHLRHYPLLLGFQ 11
>CO979887
Length = 823
Score = 30.0 bits (66), Expect = 2.4
Identities = 26/78 (33%), Positives = 34/78 (43%), Gaps = 1/78 (1%)
Frame = +1
Query: 407 CEVYRKGHQLVESHLFGSVLIPKILIVYRHRHQQRHHFLKGNVLVLVKHHVPHFHHL-LG 465
C + GHQL H G+ KIL ++RHRH +H H P H L L
Sbjct: 136 CLTLQDGHQLF-LHEIGTEA--KILHLHRHRHHH*NHL----------QHCPTVHSLELN 276
Query: 466 FQISLPRFQSLFWIQLLF 483
+ L F +L + LLF
Sbjct: 277 VLLVLELFATLLNVALLF 330
>TC232194 weakly similar to UP|Q9LSN6 (Q9LSN6) Arabidopsis thaliana genomic
DNA, chromosome 3, TAC clone:K14A17, partial (40%)
Length = 550
Score = 30.0 bits (66), Expect = 2.4
Identities = 14/35 (40%), Positives = 19/35 (54%), Gaps = 3/35 (8%)
Frame = -3
Query: 436 HRHQQRHHFLKGNVLVLVKHH---VPHFHHLLGFQ 467
H H +RHH+ ++ + HH HFHHLL Q
Sbjct: 359 HHHLERHHYHFLSLFQRLSHHHLFQFHFHHLLQIQ 255
>TC214303 similar to UP|Q9T0K5 (Q9T0K5) Extensin-like protein, partial (18%)
Length = 1014
Score = 30.0 bits (66), Expect = 2.4
Identities = 20/55 (36%), Positives = 26/55 (46%)
Frame = +3
Query: 414 HQLVESHLFGSVLIPKILIVYRHRHQQRHHFLKGNVLVLVKHHVPHFHHLLGFQI 468
H LV SHL L + ++ H H HH L + ++HH H HHL F I
Sbjct: 201 HHLV*SHLLRHHLHHLVKSIHHHHH---HHTLH----LTIRHH-HHHHHLPQFNI 341
Score = 28.5 bits (62), Expect = 7.0
Identities = 16/47 (34%), Positives = 22/47 (46%), Gaps = 4/47 (8%)
Frame = +3
Query: 420 HLFGSVLIPKILIVYRHRHQQRHHFLKGNVLVLVKHH----VPHFHH 462
HL L+P I + HRH HH + ++L HH + H HH
Sbjct: 138 HLLQFTLLPHQFI-HHHRHPHLHHLV*SHLLRHHLHHLVKSIHHHHH 275
Score = 28.1 bits (61), Expect = 9.1
Identities = 15/41 (36%), Positives = 21/41 (50%), Gaps = 9/41 (21%)
Frame = +3
Query: 433 VYRHRHQQ------RHHFLKGNVLV--LVKHHV-PHFHHLL 464
++ H H Q RHH L+ +L + HH PH HHL+
Sbjct: 90 IHHHHHHQFIPIYHRHHLLQFTLLPHQFIHHHRHPHLHHLV 212
>TC204981 similar to UP|Q9LN36 (Q9LN36) F18O14.34, partial (71%)
Length = 774
Score = 29.6 bits (65), Expect = 3.1
Identities = 25/84 (29%), Positives = 38/84 (44%), Gaps = 19/84 (22%)
Frame = -2
Query: 14 LKTYFALIQHPLS-FFNNFPTVLVMDSTYQTNMYKM---------PMFEIVGVTSTDLTY 63
LK FA LS FPT++++ S +T Y+M PM E +G T+T L+
Sbjct: 488 LKFDFATCTRELS*MLVPFPTLMLLTSPLKTTPYQMDDPLPS*TSPMTEALGATNTSLST 309
Query: 64 S---------VGFEFVTHEKEENF 78
+ VG H KE+++
Sbjct: 308 TGALSNMFIRVGCVETKHNKEDDY 237
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.342 0.150 0.517
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,697,194
Number of Sequences: 63676
Number of extensions: 447129
Number of successful extensions: 4365
Number of sequences better than 10.0: 71
Number of HSP's better than 10.0 without gapping: 4134
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4308
length of query: 484
length of database: 12,639,632
effective HSP length: 101
effective length of query: 383
effective length of database: 6,208,356
effective search space: 2377800348
effective search space used: 2377800348
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 61 (28.1 bits)
Medicago: description of AC140031.2


