

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139743.2 - phase: 0 /pseudo
(461 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
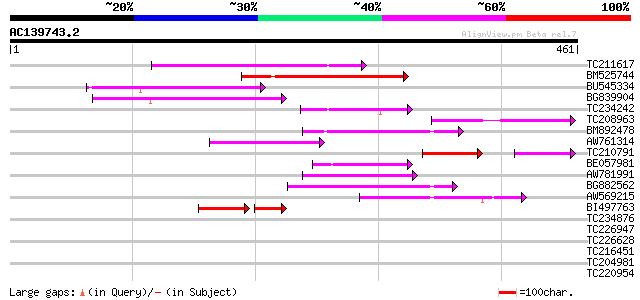
Score E
Sequences producing significant alignments: (bits) Value
TC211617 133 1e-31
BM525744 125 4e-29
BU545334 84 2e-16
BG839904 74 1e-13
TC234242 74 2e-13
TC208963 70 2e-12
BM892478 68 1e-11
AW761314 65 5e-11
TC210791 47 6e-11
BE057981 61 1e-09
AW781991 53 2e-07
BG882562 52 4e-07
AW569215 weakly similar to GP|4914326|gb| F14N23.12 {Arabidopsis... 50 3e-06
BI497763 39 4e-06
TC234876 similar to UP|Q7RNR5 (Q7RNR5) 60S ribosomal protein L27... 39 0.006
TC226947 similar to UP|IF39_TOBAC (P56821) Eukaryotic translatio... 31 1.0
TC226628 similar to UP|HAT5_ARATH (Q02283) Homeobox-leucine zipp... 31 1.0
TC216451 similar to UP|Q93XR7 (Q93XR7) Fructose-6-phosphate 2-ki... 31 1.3
TC204981 similar to UP|Q9LN36 (Q9LN36) F18O14.34, partial (71%) 30 1.7
TC220954 weakly similar to UP|Q84UC4 (Q84UC4) HASTY, partial (19%) 30 2.3
>TC211617
Length = 534
Score = 133 bits (335), Expect = 1e-31
Identities = 65/175 (37%), Positives = 100/175 (57%)
Frame = -1
Query: 116 CERSGSHIVPEKKPKHANTGSRKCGCLFMISGYQSKQTKEWGLNILNGVHNHPMEPALEG 175
CERSG + +K+ +T +RKCGC F I G + W + ++ G+HNH + L
Sbjct: 531 CERSGKYKCRKKEFVRRDTCTRKCGCPFKIRGKPMHGGEGWTVKLICGIHNHELAKTLVE 352
Query: 176 HILAGRLKEDDKKIVRDLTKSKMLPRNILIHLKNQRPHCMTNVKQVYNERQQIWKANRGD 235
H AGRL +D+K I+ D+TKS + PRNI + LK + T +KQ+YN R + RGD
Sbjct: 351 HPYAGRLTDDEKNIIADMTKSNVKPRNIQLTLKEHNTNSCTTIKQIYNARSAYHSSIRGD 172
Query: 236 KKSLQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDST 290
+Q L+ L+ Y ++ R + + + + D+FW HP ++KL N V ++DST
Sbjct: 171 DTEMQHLMRLLKRDQYIHWHRLK-DQDVVRDLFWCHPDAVKLCNTCHLVFLIDST 10
>BM525744
Length = 411
Score = 125 bits (314), Expect = 4e-29
Identities = 60/136 (44%), Positives = 89/136 (65%)
Frame = +3
Query: 189 IVRDLTKSKMLPRNILIHLKNQRPHCMTNVKQVYNERQQIWKANRGDKKSLQFLISKLEE 248
+V D+TK + PRNIL+ LK+ T ++ +YN RQ + +G + +Q L+ LE
Sbjct: 6 LVIDMTKKMVEPRNILLTLKDHNND--TTIRHIYNARQAYRSSQKGPRTEMQHLLKLLEH 179
Query: 249 HNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRMPMFEVVGVTS 308
Y +SR +S+ I DIFWAHP +IKL +F TVL +D+TYK N Y++P+ E+VGVTS
Sbjct: 180 DQYVCWSRKVDDSDAIRDIFWAHPDAIKLLGSFHTVLFLDNTYKVNRYQLPLLEIVGVTS 359
Query: 309 TDLTYSVGFGFMTHEK 324
T+LT+SV F +M ++
Sbjct: 360 TELTFSVAFAYMESDE 407
>BU545334
Length = 556
Score = 83.6 bits (205), Expect = 2e-16
Identities = 56/152 (36%), Positives = 80/152 (51%), Gaps = 6/152 (3%)
Frame = -1
Query: 63 QPHEVDTAKFFSDDVKSKNRDELLEWVRRQANKAGFTIVTQRS-----SLINPMFRLV-C 116
+PH VD + F+ RD +L+ R A++ GF V RS S F L+ C
Sbjct: 466 EPH-VDCSYAFNTXQVFGTRDYVLQ*XRSVAHENGFAAVIMRSDTHTGSRGRSSFVLIGC 290
Query: 117 ERSGSHIVPEKKPKHANTGSRKCGCLFMISGYQSKQTKEWGLNILNGVHNHPMEPALEGH 176
ER+G + K GSRK GC F + G + W + ++ G+HNH + +L H
Sbjct: 289 ERNGQYKCRNKXXVIRAIGSRKYGCPFKLRGKPVIGGQGWMMKLICGIHNHELAKSLVEH 110
Query: 177 ILAGRLKEDDKKIVRDLTKSKMLPRNILIHLK 208
AGRL +D+K I+ D+TKS + PRNIL+ LK
Sbjct: 109 PYAGRLTKDEKTIIVDITKSMVKPRNILLILK 14
>BG839904
Length = 712
Score = 74.3 bits (181), Expect = 1e-13
Identities = 50/172 (29%), Positives = 77/172 (44%), Gaps = 14/172 (8%)
Frame = -3
Query: 68 DTAKFFSDDVKSKNRDELLEWVRRQANKAGFTIVTQRSSLINPMFR------------LV 115
D + F+ +R+ +L W R A + GF ++ RS R +
Sbjct: 518 DCSDAFNTTEVFPSREAMLNWARDVAKENGFVLIILRSETSTTCTRSTRCNQRKTFVIMG 339
Query: 116 CERSGSHIVPEKKPKHAN-TGSRKCGCLFMISGYQSKQTKEWGLNILNGVHNHPMEPALE 174
C+RSG + P K +G+RKC C F + G K+ + W + ++ G HNH + L
Sbjct: 338 CDRSGKYRGPYKNELSRKVSGTRKCECPFKLKGKALKKAEGWIVKVMCGCHNHDLXETLV 159
Query: 175 GHILAGRLKEDDKKIVRDLTKS-KMLPRNILIHLKNQRPHCMTNVKQVYNER 225
GH AGRL ++K R K R+ + H K +T +KQ+YN R
Sbjct: 158 GHPYAGRLSAEEKSFGRCFDKEHDEAXRHSIDH*KIITWVTVTTIKQIYNAR 3
>TC234242
Length = 478
Score = 73.6 bits (179), Expect = 2e-13
Identities = 35/96 (36%), Positives = 57/96 (58%), Gaps = 5/96 (5%)
Frame = +3
Query: 237 KSLQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMY 296
+ +Q L+ LE Y ++ R + + + + DIFW HP S+KL N V ++DSTY+TN Y
Sbjct: 12 EEMQHLMKFLERDQYIHWHRIK-DEDVVRDIFWCHPDSVKLVNACNLVFLIDSTYRTNRY 188
Query: 297 RMPM-----FEVVGVTSTDLTYSVGFGFMTHEKEEN 327
R+P+ + VG+T T +T+S F ++ E+ N
Sbjct: 189 RLPLLDFVVLDFVGMTPTGMTFSASFAYVEGERVNN 296
>TC208963
Length = 1163
Score = 70.5 bits (171), Expect = 2e-12
Identities = 45/118 (38%), Positives = 61/118 (51%), Gaps = 1/118 (0%)
Frame = +1
Query: 344 NMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVKQRCVLNCKYPLGFKKDGKEVS 403
+MP+VIVT D++LM AV VFP S L C FH+ NVK +C V
Sbjct: 16 DMPQVIVTVGDIALMSAVQVVFPSSSNLLCRFHINQNVKAKC-------------KSIVH 156
Query: 404 NRDVVKKIMNAWKAMVESPNQQLYANALVEFKDSCSDFSIFVDYAMTT-LDEVKDKIV 460
++ + +M+AW +V SPN+ Y L F++ C DF I DY T L K+K V
Sbjct: 157 LKEKQELMMDAWDVVVNSPNEGEYMQRLAFFENVCLDFPILGDYVKNTWLIPHKEKFV 330
>BM892478
Length = 429
Score = 67.8 bits (164), Expect = 1e-11
Identities = 35/131 (26%), Positives = 71/131 (53%)
Frame = +3
Query: 239 LQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRM 298
L++ S+ E +Y+ ++++ +IFWA S ++F VLV+D++Y+ +Y +
Sbjct: 33 LEYFQSRQAEDTGFFYAM-EVDNGNCMNIFWADGRSRYSCSHFGDVLVLDTSYRKTVYLV 209
Query: 299 PMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIVTDRDMSLM 358
P VGV +G + E EE+F W+ + +S + +P ++ D+D+++
Sbjct: 210 PFATFVGVNHHKQPVLLGCALIADESEESFTWLFQTWLRAMSGR--LPLTVIADQDIAIQ 383
Query: 359 KAVAHVFPESY 369
+A+A VFP ++
Sbjct: 384 RAIAKVFPVTH 416
>AW761314
Length = 331
Score = 65.5 bits (158), Expect = 5e-11
Identities = 33/94 (35%), Positives = 52/94 (55%)
Frame = +1
Query: 163 GVHNHPMEPALEGHILAGRLKEDDKKIVRDLTKSKMLPRNILIHLKNQRPHCMTNVKQVY 222
G NH + +L H A RL +D+ I+ ++TKS + P+NIL+ LK + T +KQ+Y
Sbjct: 49 GSDNHELVKSLVRHPYASRLTKDENIIITNMTKSMVKPKNILLTLKEHNVNSYTAIKQIY 228
Query: 223 NERQQIWKANRGDKKSLQFLISKLEEHNYTYYSR 256
N R + RG +Q L+ LE+ Y ++ R
Sbjct: 229 NARSAYCSSIRGSDTKMQQLMKLLEQDQYIHWHR 330
>TC210791
Length = 456
Score = 46.6 bits (109), Expect(2) = 6e-11
Identities = 24/49 (48%), Positives = 30/49 (60%)
Frame = +1
Query: 336 RKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVKQR 384
R L+ MP VI+T RD++LM AV VFP S L C FH+ NVK +
Sbjct: 1 RGLIFKDNEMPPVIITVRDIALMDAVQVVFPSSSNLLCRFHISKNVKAK 147
Score = 38.5 bits (88), Expect(2) = 6e-11
Identities = 20/51 (39%), Positives = 28/51 (54%), Gaps = 1/51 (1%)
Frame = +2
Query: 411 IMNAWKAMVESPNQQLYANALVEFKDSCSDFSIFVDYAMTT-LDEVKDKIV 460
+M+AW +++ SPN+ Y L + CSDF F DY T L K+K V
Sbjct: 188 VMDAWDSVMNSPNEGEYMQRLTLLEKVCSDFPTFGDYVKNTWLIPHKEKFV 340
>BE057981
Length = 268
Score = 60.8 bits (146), Expect = 1e-09
Identities = 28/81 (34%), Positives = 47/81 (57%)
Frame = +2
Query: 247 EEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRMPMFEVVGV 306
E Y ++ R + E + D+FW H ++KL N V ++++ YKTN YR+ + + GV
Sbjct: 14 ERDQYIHWHRLKDEF-VVRDLFWCHLDAVKLCNACHLVFLINNAYKTNRYRLLLLDFAGV 190
Query: 307 TSTDLTYSVGFGFMTHEKEEN 327
T +T+SVGF ++ E+ N
Sbjct: 191 TPARMTFSVGFAYLEGERVNN 253
>AW781991
Length = 419
Score = 53.1 bits (126), Expect = 2e-07
Identities = 27/93 (29%), Positives = 45/93 (48%)
Frame = +3
Query: 239 LQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRM 298
L +L E+ +Y+ E +I ++FWA P + + F + D+TY++N YR+
Sbjct: 129 LDYLRQMHAENPNFFYAVQGDEDQSITNVFWADPKARMNYTFFGDTVTFDTTYRSNRYRL 308
Query: 299 PMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWV 331
P GV G F+ +E E +FVW+
Sbjct: 309 PFAFFTGVNHHGQPVLFGCAFLINESEASFVWL 407
>BG882562
Length = 416
Score = 52.4 bits (124), Expect = 4e-07
Identities = 30/139 (21%), Positives = 65/139 (46%), Gaps = 1/139 (0%)
Frame = +3
Query: 227 QIWKANRGDKKSLQFLISKLEEHNYTYYSRTQL-ESNTIEDIFWAHPTSIKLFNNFPTVL 285
Q ++ +GD + + +++ N ++ L + + ++FW S ++ F V+
Sbjct: 6 QAFET*KGDPELISNYFCRIQLMNPNFFYVMDLNDDGQLRNVFWIDSRSRAAYSYFGDVV 185
Query: 286 VMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNM 345
DST +N Y +P+ VGV + +G G + E E ++W+ ++ +
Sbjct: 186 AFDSTCLSNNYEIPLVAFVGVNHHGKSVLLGCGLLADETFETYIWLFRAWLTCMTGR--P 359
Query: 346 PKVIVTDRDMSLMKAVAHV 364
P+ I+T+ ++ A+A V
Sbjct: 360 PQTIITNXCKAMQSAIAEV 416
>AW569215 weakly similar to GP|4914326|gb| F14N23.12 {Arabidopsis thaliana},
partial (19%)
Length = 425
Score = 49.7 bits (117), Expect = 3e-06
Identities = 33/138 (23%), Positives = 63/138 (44%), Gaps = 2/138 (1%)
Frame = +1
Query: 285 LVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMN 344
+V D+TY+ Y M + +GV + +T + E ++F W L + K
Sbjct: 10 VVFDTTYRVEAYDMLLGIWLGVDNNGMTCFFSCALLRDENIQSFSWALKAFLGFMKGK-- 183
Query: 345 MPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVKQ--RCVLNCKYPLGFKKDGKEV 402
P+ I+TD +M L +A+A P++ C +H+ + +L +Y +K + +
Sbjct: 184 APQTILTDHNMWLKEAIAVELPQTKHAFCIWHILSKFSDWFSLLLGSQYD-EWKAEFHRL 360
Query: 403 SNRDVVKKIMNAWKAMVE 420
N + V+ W+ MV+
Sbjct: 361 YNLEQVEDFEEGWRQMVD 414
>BI497763
Length = 421
Score = 38.9 bits (89), Expect(2) = 4e-06
Identities = 17/42 (40%), Positives = 26/42 (61%)
Frame = +2
Query: 154 KEWGLNILNGVHNHPMEPALEGHILAGRLKEDDKKIVRDLTK 195
K W + +L G HNH + +L H GRL +D+K I+ ++TK
Sbjct: 41 KGWMVKLLCGSHNHELTKSLVEHPYVGRLTKDEKTIIVNMTK 166
Score = 29.6 bits (65), Expect(2) = 4e-06
Identities = 12/26 (46%), Positives = 17/26 (65%)
Frame = +3
Query: 200 PRNILIHLKNQRPHCMTNVKQVYNER 225
PRNIL+ LK + T ++Q+YN R
Sbjct: 162 PRNILLMLKEYNANGCTTIRQIYNAR 239
>TC234876 similar to UP|Q7RNR5 (Q7RNR5) 60S ribosomal protein L27 homolog,
partial (15%)
Length = 920
Score = 38.5 bits (88), Expect = 0.006
Identities = 20/67 (29%), Positives = 32/67 (46%)
Frame = -3
Query: 136 SRKCGCLFMISGYQSKQTKEWGLNILNGVHNHPMEPALEGHILAGRLKEDDKKIVRDLTK 195
+RKCGCLF + + W ++ HNH + L H RL ++K I+ D+ K
Sbjct: 603 NRKCGCLFKLQVKPVLGGEGWMKKLIRESHNHALAKLLVRHPYVDRLTINEKIIIGDMIK 424
Query: 196 SKMLPRN 202
+ +N
Sbjct: 423 LIIKSKN 403
>TC226947 similar to UP|IF39_TOBAC (P56821) Eukaryotic translation initiation
factor 3 subunit 9 (eIF-3 eta) (eIF3 p110) (eIF3b),
partial (25%)
Length = 890
Score = 31.2 bits (69), Expect = 1.0
Identities = 23/73 (31%), Positives = 36/73 (48%)
Frame = -3
Query: 375 FHVQANVKQRCVLNCKYPLGFKKDGKEVSNRDVVKKIMNAWKAMVESPNQQLYANALVEF 434
FHV+ + K++ L Y +GF KD EVS + + +N M ES + QL + L
Sbjct: 741 FHVKFDNKKKTKLMKDYKIGFYKDEIEVSCPTTIPRQVN---KMFESKSSQLKNSLLTIL 571
Query: 435 KDSCSDFSIFVDY 447
K D ++D+
Sbjct: 570 K--VKDLFCYIDH 538
>TC226628 similar to UP|HAT5_ARATH (Q02283) Homeobox-leucine zipper protein
HAT5 (HD-ZIP protein 5) (HD-ZIP protein ATHB-1), partial
(20%)
Length = 767
Score = 31.2 bits (69), Expect = 1.0
Identities = 20/59 (33%), Positives = 28/59 (46%), Gaps = 8/59 (13%)
Frame = -2
Query: 165 HNHPMEPALEGHILAGRLKEDDKKIVRDLTK--SKMLPRNILIH------LKNQRPHCM 215
HN P +P G + GRL E ++ DL + S +LPR ++ KN RP M
Sbjct: 640 HNTPHKPVSSGEVKKGRLFEASSMLIIDLPRKASSVLPRKSMLFPREAEASKNIRPDSM 464
>TC216451 similar to UP|Q93XR7 (Q93XR7) Fructose-6-phosphate
2-kinase/fructose-2,6-bisphosphatase, partial (62%)
Length = 1595
Score = 30.8 bits (68), Expect = 1.3
Identities = 18/66 (27%), Positives = 30/66 (45%), Gaps = 2/66 (3%)
Frame = -3
Query: 329 VWVLTMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYALNCF--FHVQANVKQRCV 386
+W+ ++R LL+ K N+P + + +A+ F E Y + F FH + V
Sbjct: 609 IWLFCIIRTLLNFKANIP--------FNYISIIAYCFQEDYLASSFSQFHQHVSSFLATV 454
Query: 387 LNCKYP 392
KYP
Sbjct: 453 CGIKYP 436
>TC204981 similar to UP|Q9LN36 (Q9LN36) F18O14.34, partial (71%)
Length = 774
Score = 30.4 bits (67), Expect = 1.7
Identities = 20/66 (30%), Positives = 32/66 (48%), Gaps = 18/66 (27%)
Frame = -2
Query: 281 FPTVLVMDSTYKTNMYRM---------PMFEVVGVTSTDLTYS---------VGFGFMTH 322
FPT++++ S KT Y+M PM E +G T+T L+ + VG H
Sbjct: 434 FPTLMLLTSPLKTTPYQMDDPLPS*TSPMTEALGATNTSLSTTGALSNMFIRVGCVETKH 255
Query: 323 EKEENF 328
KE+++
Sbjct: 254 NKEDDY 237
>TC220954 weakly similar to UP|Q84UC4 (Q84UC4) HASTY, partial (19%)
Length = 724
Score = 30.0 bits (66), Expect = 2.3
Identities = 20/84 (23%), Positives = 36/84 (42%), Gaps = 8/84 (9%)
Frame = -1
Query: 5 IIVMVKNAKDVPGETKENFVENSGDVKPPNFEAKTNIDEASVHVEASKYVGGSGVL---- 60
+I+ VK + + + FV + +E SVHV+ASK++ G
Sbjct: 493 LIMDVKRSLETYSTSSA*FVTIARTTNAEQYEEILVTASPSVHVKASKHICSVGRTRHSC 314
Query: 61 ----PLQPHEVDTAKFFSDDVKSK 80
P H+ +KFF +D+ ++
Sbjct: 313 FRRKPTMEHDATVSKFFREDISNR 242
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,431,023
Number of Sequences: 63676
Number of extensions: 273208
Number of successful extensions: 1354
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 1345
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1350
length of query: 461
length of database: 12,639,632
effective HSP length: 101
effective length of query: 360
effective length of database: 6,208,356
effective search space: 2235008160
effective search space used: 2235008160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC139743.2


