

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139706.5 - phase: 0
(424 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
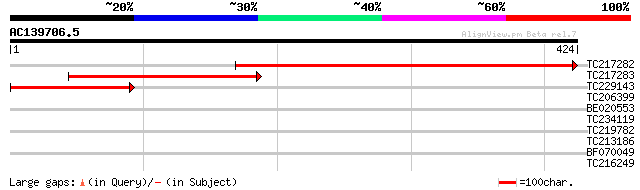
Score E
Sequences producing significant alignments: (bits) Value
TC217282 similar to UP|Q9ST36 (Q9ST36) JPR ORF1 protein, partial... 474 e-134
TC217283 homologue to UP|Q9ST36 (Q9ST36) JPR ORF1 protein, parti... 272 2e-73
TC229143 homologue to UP|Q9ST36 (Q9ST36) JPR ORF1 protein, parti... 140 8e-34
TC206399 similar to GB|BAB09797.1|9759182|AB013388 ATP-dependent... 29 4.6
BE020553 similar to GP|15528811|dbj P0431G06.18 {Oryza sativa (j... 29 4.6
TC234119 similar to UP|Q9LTR5 (Q9LTR5) Similarity to bactericida... 28 6.0
TC219782 similar to UP|Q9SC91 (Q9SC91) Glutamine synthetase I, p... 28 6.0
TC213186 similar to UP|Q70G58 (Q70G58) NADPH thioredoxin reducta... 28 6.0
BF070049 28 6.0
TC216249 weakly similar to UP|SAP1_YEAST (P39955) SAP1 protein, ... 28 7.8
>TC217282 similar to UP|Q9ST36 (Q9ST36) JPR ORF1 protein, partial (59%)
Length = 1014
Score = 474 bits (1221), Expect = e-134
Identities = 229/255 (89%), Positives = 246/255 (95%)
Frame = +3
Query: 170 LIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKMTP 229
+ D ++ +DA EINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKAT+PVWAKMTP
Sbjct: 81 MADLSDKLVVDAFEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATIPVWAKMTP 260
Query: 230 NITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVHPI 289
NITDISQPAR+ALSSGCEGV+AINTIMSVMGINLNTLRPEPCVEGYSTPGGYSA+AVHPI
Sbjct: 261 NITDISQPARIALSSGCEGVSAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSARAVHPI 440
Query: 290 ALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYGLV 349
ALGKVMSIAKMMKSEFDSENY+LSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYGLV
Sbjct: 441 ALGKVMSIAKMMKSEFDSENYTLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYGLV 620
Query: 350 KKLSLELKDFMKKHNFTSIEDFRGASLQYFTTHTDLVHRQQEAIKQRKAIKKGLQSDKDW 409
KKL LEL+DFMKKHNFTSIEDFRG SL+YFTTHTDLV RQ+EA+++RKAIKKGLQSDKDW
Sbjct: 621 KKLCLELQDFMKKHNFTSIEDFRGVSLEYFTTHTDLVRRQKEAVQKRKAIKKGLQSDKDW 800
Query: 410 TGDGFVKESESMVSN 424
TGDGFV+E+ESMVSN
Sbjct: 801 TGDGFVQETESMVSN 845
>TC217283 homologue to UP|Q9ST36 (Q9ST36) JPR ORF1 protein, partial (34%)
Length = 600
Score = 272 bits (696), Expect = 2e-73
Identities = 133/144 (92%), Positives = 138/144 (95%)
Frame = +2
Query: 45 SEPDLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTP 104
+EPDLSV VNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWG VIAKTVSLDAAKVINVTP
Sbjct: 167 TEPDLSVTVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGAVIAKTVSLDAAKVINVTP 346
Query: 105 RYARLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNK 164
RYARLRA A+ SAKGEIIGW+NIELISDRPLE MLKEFKQLKEEYPDRILIASIMEEYNK
Sbjct: 347 RYARLRAGADRSAKGEIIGWENIELISDRPLEIMLKEFKQLKEEYPDRILIASIMEEYNK 526
Query: 165 AAWEELIDRVEQTGIDAIEINFSC 188
AAWEELIDRV+QTG+DA EINFSC
Sbjct: 527 AAWEELIDRVQQTGVDAFEINFSC 598
>TC229143 homologue to UP|Q9ST36 (Q9ST36) JPR ORF1 protein, partial (14%)
Length = 587
Score = 140 bits (354), Expect = 8e-34
Identities = 71/93 (76%), Positives = 76/93 (81%)
Frame = +2
Query: 1 MANLSMTQLKTRNSASRFSFNFSKKVYPRPSRVDFKVFASEGQVSEPDLSVKVNGLHMPN 60
MA+LSM Q++T N AS F N + KV PSRV FKVFASE Q +EPDLSV VNGL MPN
Sbjct: 212 MASLSMIQIRTGNCASGFGLNCAGKVKVGPSRVGFKVFASETQATEPDLSVTVNGLRMPN 391
Query: 61 PFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVS 93
PFVIGSGPPGTNYTVMKRAFDEGWG VIAKTVS
Sbjct: 392 PFVIGSGPPGTNYTVMKRAFDEGWGAVIAKTVS 490
>TC206399 similar to GB|BAB09797.1|9759182|AB013388 ATP-dependent Clp
protease regulatory subunit CLPX {Arabidopsis thaliana;}
, partial (41%)
Length = 1207
Score = 28.9 bits (63), Expect = 4.6
Identities = 14/49 (28%), Positives = 27/49 (54%), Gaps = 5/49 (10%)
Frame = +2
Query: 315 IGGVETGGDAAEFILL-----GANTVQVCTGVMMHGYGLVKKLSLELKD 358
I ++TG D + +++ G+ T C G ++HG G +K+ ++KD
Sbjct: 677 IPDIKTGSDRVDAVVIDEESVGSLTAPGCGGKILHGDGALKQYLAKMKD 823
>BE020553 similar to GP|15528811|dbj P0431G06.18 {Oryza sativa (japonica
cultivar-group)}, partial (13%)
Length = 335
Score = 28.9 bits (63), Expect = 4.6
Identities = 24/94 (25%), Positives = 39/94 (40%)
Frame = +1
Query: 58 MPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYARLRANANGSA 117
+P V+G G +Y++ G + T+SLD + IN + A N S
Sbjct: 34 VPIAQVVGKGKVTEDYSLSSTDLFHSPAGTVQLTLSLDPSLAINSSVNLIPESAK-NSSI 210
Query: 118 KGEIIGWQNIELISDRPLETMLKEFKQLKEEYPD 151
E+I L+ + E ML + + E+PD
Sbjct: 211 SSEVI------LLDRKISEVMLDPVEYARIEFPD 294
>TC234119 similar to UP|Q9LTR5 (Q9LTR5) Similarity to bactericidal
permeability-increasing protein, partial (4%)
Length = 676
Score = 28.5 bits (62), Expect = 6.0
Identities = 19/69 (27%), Positives = 34/69 (48%), Gaps = 8/69 (11%)
Frame = +1
Query: 202 GQDCALLEEVCGWINAKATVPVWAKMTPNITDISQPARVALSSGCEGVA--------AIN 253
G + + E+ ++ ATV K + ++ + AR L+S CEG+A AI
Sbjct: 151 GAEFKRVAEIVLVLSTMATVRAGRKPSDAEVELMREARAKLASLCEGLAPKDIVTREAIG 330
Query: 254 TIMSVMGIN 262
T++ +G+N
Sbjct: 331 TVIEDLGLN 357
>TC219782 similar to UP|Q9SC91 (Q9SC91) Glutamine synthetase I, partial (82%)
Length = 1368
Score = 28.5 bits (62), Expect = 6.0
Identities = 14/37 (37%), Positives = 21/37 (55%), Gaps = 4/37 (10%)
Frame = +2
Query: 356 LKDFMKKHNFTSIEDFRGASLQYFTTHTD----LVHR 388
LK+FM T+I+ R A + ++T H D L+HR
Sbjct: 1034 LKEFMSDKLLTTIKAIRKAEIDHYTKHKDAYKQLIHR 1144
>TC213186 similar to UP|Q70G58 (Q70G58) NADPH thioredoxin reductase, partial
(36%)
Length = 745
Score = 28.5 bits (62), Expect = 6.0
Identities = 17/43 (39%), Positives = 22/43 (50%)
Frame = +1
Query: 28 PRPSRVDFKVFASEGQVSEPDLSVKVNGLHMPNPFVIGSGPPG 70
PRPS + +V +S S SV G + N +IGSGP G
Sbjct: 157 PRPSPLPLRVASSSSSSS----SVAAPGKGVENVVIIGSGPAG 273
>BF070049
Length = 520
Score = 28.5 bits (62), Expect = 6.0
Identities = 12/28 (42%), Positives = 19/28 (67%)
Frame = +2
Query: 314 AIGGVETGGDAAEFILLGANTVQVCTGV 341
A GG++T DAA + LG + + +C+GV
Sbjct: 272 AAGGLDTPADAALMMHLGCDGLYICSGV 355
>TC216249 weakly similar to UP|SAP1_YEAST (P39955) SAP1 protein, partial
(14%)
Length = 2222
Score = 28.1 bits (61), Expect = 7.8
Identities = 18/73 (24%), Positives = 37/73 (50%)
Frame = +2
Query: 140 KEFKQLKEEYPDRILIASIMEEYNKAAWEELIDRVEQTGIDAIEINFSCPHGMPERKMGA 199
K+ + L + +P++I I +E A+W++ +DR ++ ++I + H R +
Sbjct: 497 KQNRTLTKLFPNKITIHMPQDEALLASWKQQLDR----DVETLKIKGNLHH---LRTVLG 655
Query: 200 AVGQDCALLEEVC 212
G +C LE +C
Sbjct: 656 RCGMECEGLETLC 694
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.133 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,880,242
Number of Sequences: 63676
Number of extensions: 193084
Number of successful extensions: 854
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 850
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 854
length of query: 424
length of database: 12,639,632
effective HSP length: 100
effective length of query: 324
effective length of database: 6,272,032
effective search space: 2032138368
effective search space used: 2032138368
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC139706.5


