

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139356.2 + phase: 0
(180 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
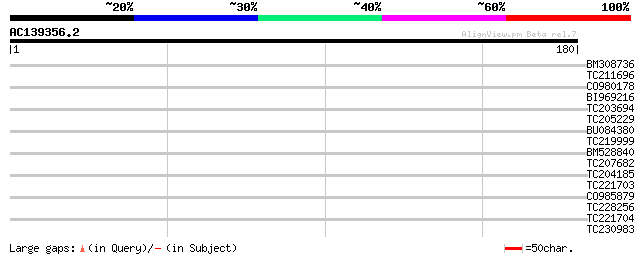
Score E
Sequences producing significant alignments: (bits) Value
BM308736 similar to GP|12006424|gb Ku70-like protein {Arabidopsi... 29 1.1
TC211696 homologue to UP|Q9ARC5 (Q9ARC5) Membrane transporter, c... 28 2.3
CO980178 28 2.3
BI969216 weakly similar to GP|28140231|gb protein disulphide iso... 28 2.3
TC203694 similar to UP|Q9XQB0 (Q9XQB0) Carbonic anhydrase, parti... 27 4.0
TC205229 similar to UP|Q8W0Z9 (Q8W0Z9) AT3g58560/F14P22_150, par... 27 4.0
BU084380 homologue to GP|15420162|gb| F-box containing protein O... 27 4.0
TC219999 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (5%) 27 4.0
BM528840 27 5.2
TC207682 similar to UP|Q6PCF8 (Q6PCF8) MGC69148 protein, partial... 27 6.8
TC204185 homologue to UP|TBB1_SOYBN (P12459) Tubulin beta-1 chai... 27 6.8
TC221703 27 6.8
CO985879 26 8.9
TC228256 weakly similar to UP|Q9ATY4 (Q9ATY4) MAP kinase phospha... 26 8.9
TC221704 homologue to UP|O24457 (O24457) Pyruvate dehydrogenase ... 26 8.9
TC230983 similar to GB|AAB61117.1|2194142|F20P5 ESTs gb|N38288,g... 26 8.9
>BM308736 similar to GP|12006424|gb Ku70-like protein {Arabidopsis thaliana},
partial (22%)
Length = 428
Score = 29.3 bits (64), Expect = 1.1
Identities = 14/42 (33%), Positives = 20/42 (47%)
Frame = -1
Query: 45 KIFSITGDVQCKSCQTKFQMPFDLRTKFDVISQYLVTNMNTM 86
KI I +QC SC F PFD + +I + T +N +
Sbjct: 377 KILCIFSSLQCHSCHI*FYCPFDTPKRIIIICEQKYTFVNCL 252
>TC211696 homologue to UP|Q9ARC5 (Q9ARC5) Membrane transporter, complete
Length = 1899
Score = 28.1 bits (61), Expect = 2.3
Identities = 14/33 (42%), Positives = 18/33 (54%)
Frame = -3
Query: 1 MTANHTHVPPRRRSAPRSGPIPIPFIWATNHRA 33
++ + THVPP PRS PI P+ HRA
Sbjct: 688 LSGSTTHVPP-----PRSSPIRKPYNLTVEHRA 605
>CO980178
Length = 702
Score = 28.1 bits (61), Expect = 2.3
Identities = 18/62 (29%), Positives = 27/62 (43%), Gaps = 4/62 (6%)
Frame = +2
Query: 119 KKNINWLFLLLGQMLGCCKLE----QLKYFCKYNNHHRTGAKNRVLYLTYFELCQQLDPS 174
K + W+F ++G M GC L+ + Y +H+ LT F LC + PS
Sbjct: 317 KGEVCWIF*VIGHMKGCSNLKDNNLNIIVILSYPSHN---------LLTPFSLCHLIIPS 469
Query: 175 GP 176
P
Sbjct: 470 LP 475
>BI969216 weakly similar to GP|28140231|gb protein disulphide isomerase
{Elaeis guineensis}, partial (26%)
Length = 634
Score = 28.1 bits (61), Expect = 2.3
Identities = 13/47 (27%), Positives = 23/47 (48%)
Frame = -1
Query: 88 DRAPKILMNPRLPRCAHCNQENSVKPVIAEKKKNINWLFLLLGQMLG 134
D + +L+ +P C HC + +A NI+ FL++ +M G
Sbjct: 589 DESKNVLLEIYVPWCXHCQSXEPISTKLANXXXNID--FLVIAKMDG 455
>TC203694 similar to UP|Q9XQB0 (Q9XQB0) Carbonic anhydrase, partial (58%)
Length = 731
Score = 27.3 bits (59), Expect = 4.0
Identities = 15/35 (42%), Positives = 18/35 (50%), Gaps = 1/35 (2%)
Frame = -1
Query: 15 APRSGPIPIPFIWATNHRAKIHNLNH-LLQNKIFS 48
AP G P PFI H + HNL H +QN I +
Sbjct: 611 APSHGKAPSPFIPPQAHVSNDHNLXHPKMQNSILN 507
>TC205229 similar to UP|Q8W0Z9 (Q8W0Z9) AT3g58560/F14P22_150, partial (57%)
Length = 1743
Score = 27.3 bits (59), Expect = 4.0
Identities = 18/55 (32%), Positives = 26/55 (46%), Gaps = 2/55 (3%)
Frame = -3
Query: 57 SCQTKFQMP--FDLRTKFDVISQYLVTNMNTMHDRAPKILMNPRLPRCAHCNQEN 109
SC + Q+ +D RT + + L + N H AP + P P C CNQE+
Sbjct: 1066 SCSSGTQLT*LYDQRTIQE--KEVLCLSSNFPHPIAPTMTPQPMNPLCKICNQES 908
>BU084380 homologue to GP|15420162|gb| F-box containing protein ORE9
{Arabidopsis thaliana}, partial (9%)
Length = 410
Score = 27.3 bits (59), Expect = 4.0
Identities = 14/44 (31%), Positives = 24/44 (53%), Gaps = 2/44 (4%)
Frame = -1
Query: 30 NHRAKIHNLNHLLQNKIFSITGDVQC--KSCQTKFQMPFDLRTK 71
N+R K+ N+N IF ++G V+C +CQ+ + F +K
Sbjct: 302 NNRQKLQNVNFFYIQAIFPLSGLVRCCYITCQSHIRSLFSASSK 171
>TC219999 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (5%)
Length = 641
Score = 27.3 bits (59), Expect = 4.0
Identities = 14/40 (35%), Positives = 20/40 (50%), Gaps = 8/40 (20%)
Frame = -1
Query: 9 PPRRRSAPRSGPIPIPF--------IWATNHRAKIHNLNH 40
PP +RS+P + P P+P + HR IH+L H
Sbjct: 314 PPSKRSSPSALPPPLPHRLRLLPPTLRRHRHRLLIHHLRH 195
>BM528840
Length = 406
Score = 26.9 bits (58), Expect = 5.2
Identities = 10/35 (28%), Positives = 17/35 (48%)
Frame = +2
Query: 7 HVPPRRRSAPRSGPIPIPFIWATNHRAKIHNLNHL 41
H P +R P + P+P P W + + + +HL
Sbjct: 80 HAPLNKRCKPSASPLPQPQQWNASDDGSVSSPSHL 184
>TC207682 similar to UP|Q6PCF8 (Q6PCF8) MGC69148 protein, partial (26%)
Length = 511
Score = 26.6 bits (57), Expect = 6.8
Identities = 11/31 (35%), Positives = 17/31 (54%), Gaps = 1/31 (3%)
Frame = -2
Query: 12 RRSAPRSGPIPIP-FIWATNHRAKIHNLNHL 41
+R P P+ + F W +HR K+ +L HL
Sbjct: 399 KRRGPTLSPVALLCFTWLPSHRGKVPDLQHL 307
>TC204185 homologue to UP|TBB1_SOYBN (P12459) Tubulin beta-1 chain,
complete
Length = 1732
Score = 26.6 bits (57), Expect = 6.8
Identities = 13/43 (30%), Positives = 20/43 (46%)
Frame = +1
Query: 15 APRSGPIPIPFIWATNHRAKIHNLNHLLQNKIFSITGDVQCKS 57
AP GP +PF T+ + + LNHL++ + C S
Sbjct: 46 APTKGPPLLPFSTCTSSNSILQLLNHLVELNYLQNERNPSCSS 174
>TC221703
Length = 781
Score = 26.6 bits (57), Expect = 6.8
Identities = 20/79 (25%), Positives = 35/79 (43%), Gaps = 8/79 (10%)
Frame = +2
Query: 49 ITGDVQCKSCQTKFQMPFDLRTKFDVISQYLVTNMNTMHDRAPKILMNPRLP-------- 100
++ D+QC+ CQ + D+ K +V ++ +V N K+ + RLP
Sbjct: 260 LSADMQCEKCQKRVT---DIIAKMNVETESVVIN-----GLEKKVTLTFRLPSIGKVTTR 415
Query: 101 RCAHCNQENSVKPVIAEKK 119
+ H N NS+ V K+
Sbjct: 416 KITHINNRNSLPKVAIIKR 472
>CO985879
Length = 695
Score = 26.2 bits (56), Expect = 8.9
Identities = 10/16 (62%), Positives = 10/16 (62%)
Frame = -2
Query: 5 HTHVPPRRRSAPRSGP 20
H H P RRR APR P
Sbjct: 400 HVHAPRRRRGAPRPRP 353
>TC228256 weakly similar to UP|Q9ATY4 (Q9ATY4) MAP kinase phosphatase
(Fragment), partial (10%)
Length = 1848
Score = 26.2 bits (56), Expect = 8.9
Identities = 9/17 (52%), Positives = 11/17 (63%)
Frame = +3
Query: 143 YFCKYNNHHRTGAKNRV 159
YFCK NNH + G N +
Sbjct: 861 YFCKSNNHAKDGGVNSI 911
>TC221704 homologue to UP|O24457 (O24457) Pyruvate dehydrogenase E1 alpha
subunit (At1g01090/T25K16_8) , partial (46%)
Length = 616
Score = 26.2 bits (56), Expect = 8.9
Identities = 12/25 (48%), Positives = 14/25 (56%)
Frame = -3
Query: 129 LGQMLGCCKLEQLKYFCKYNNHHRT 153
L Q+LG C EQ + FC YN T
Sbjct: 173 LTQLLGHCTSEQNQTFCLYNTSGHT 99
>TC230983 similar to GB|AAB61117.1|2194142|F20P5 ESTs
gb|N38288,gb|T43486,gb|AA395242 come from this gene.
{Arabidopsis thaliana;} , partial (34%)
Length = 938
Score = 26.2 bits (56), Expect = 8.9
Identities = 14/43 (32%), Positives = 21/43 (48%)
Frame = -1
Query: 104 HCNQENSVKPVIAEKKKNINWLFLLLGQMLGCCKLEQLKYFCK 146
H N+ P AEKKKN N++F CK ++ ++ K
Sbjct: 722 HQNRSKRWNPPAAEKKKNPNFIF---------CKSQEFRHLNK 621
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.138 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,344,007
Number of Sequences: 63676
Number of extensions: 191185
Number of successful extensions: 1574
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 1565
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1573
length of query: 180
length of database: 12,639,632
effective HSP length: 91
effective length of query: 89
effective length of database: 6,845,116
effective search space: 609215324
effective search space used: 609215324
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC139356.2


