

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139354.8 + phase: 0 /pseudo
(481 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
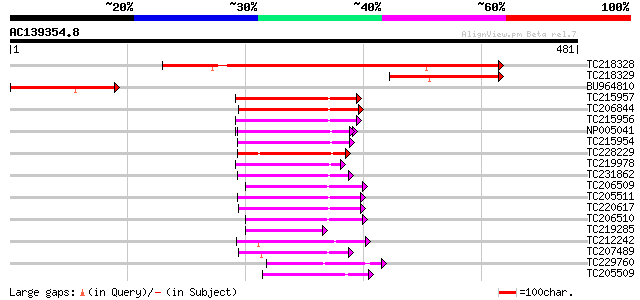
Score E
Sequences producing significant alignments: (bits) Value
TC218328 similar to UP|Q84VX9 (Q84VX9) At4g08320, partial (32%) 364 e-101
TC218329 130 1e-30
BU964810 similar to GP|28416691|gb| At4g08320 {Arabidopsis thali... 120 2e-27
TC215957 similar to UP|Q9SI76 (Q9SI76) F23N19.10, partial (32%) 85 8e-17
TC206844 similar to UP|Q8H1H4 (Q8H1H4) Type 5 serine/threonine p... 82 5e-16
TC215956 similar to UP|Q9SI76 (Q9SI76) F23N19.10, partial (25%) 79 3e-15
NP005041 sti (stress inducible protein) 68 8e-12
TC215954 similar to UP|Q9LNB6 (Q9LNB6) F5O11.2 (At1g12270/F5O11_... 65 9e-11
TC228229 60 2e-09
TC219978 weakly similar to UP|Q94GR7 (Q94GR7) Chloroplast protei... 60 3e-09
TC231862 similar to GB|AAL60196.1|18139887|AF441079 O-linked N-a... 57 1e-08
TC206509 similar to UP|Q84UV7 (Q84UV7) SGT1, partial (53%) 54 2e-07
TC205511 similar to UP|Q9MAH1 (Q9MAH1) F12M16.20 (At1g53300), pa... 54 2e-07
TC220617 weakly similar to UP|Q9SIN1 (Q9SIN1) Expressed protein ... 50 3e-06
TC206510 similar to UP|Q84UV7 (Q84UV7) SGT1, partial (51%) 49 4e-06
TC219285 weakly similar to UP|Q84UV7 (Q84UV7) SGT1, partial (37%) 49 6e-06
TC212242 similar to UP|Q75LL1 (Q75LL1) Expressed protein, partia... 47 2e-05
TC207489 weakly similar to UP|Q94AA8 (Q94AA8) AT5g63200/MDC12_17... 46 3e-05
TC229760 similar to UP|Q9C566 (Q9C566) Cyclophilin-40 (Expressed... 46 4e-05
TC205509 similar to UP|Q9MAH1 (Q9MAH1) F12M16.20 (At1g53300), pa... 46 4e-05
>TC218328 similar to UP|Q84VX9 (Q84VX9) At4g08320, partial (32%)
Length = 1086
Score = 364 bits (934), Expect = e-101
Identities = 200/304 (65%), Positives = 227/304 (73%), Gaps = 14/304 (4%)
Frame = +3
Query: 130 QFFAVLEKKHYFRTNIDGGDDIVQLEKASRLFDDGFTLYRR-------WKDLGVSSLI*R 182
+FFA LEK YF +N DG DD VQLEKAS LF++ R K+L S
Sbjct: 12 RFFAALEKNCYFWSNTDGSDDPVQLEKASCLFNEACMELERSGCHQFSLKNLAES----- 176
Query: 183 IWLNH*KH*AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRS 242
L + AMQSK+Y DAIELYNCAIA++EKSAVYYCNRAAAYTQIN+YTEAIQD LRS
Sbjct: 177 --LKTLGNKAMQSKKYSDAIELYNCAIAVHEKSAVYYCNRAAAYTQINKYTEAIQDCLRS 350
Query: 243 IEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEER 302
IEIDPNYSKAYSRLGL YYAQGNYRDAI KGF+KALQLDPNNESVKENIRVAE KL+EE+
Sbjct: 351 IEIDPNYSKAYSRLGLVYYAQGNYRDAIHKGFRKALQLDPNNESVKENIRVAERKLLEEQ 530
Query: 303 HRADHNQNSR-SQEFQNHYAR-GSRSHAAPASFGSMPFNPSNLASMFMAAAN----GGQG 356
HRA NQNSR SQEF N A+ GSRSH+ P F SM FNP ++ASMFM N QG
Sbjct: 531 HRAYQNQNSRSSQEFPNQSAQGGSRSHSVPPPFSSMSFNPRDIASMFMNITNPTNAQPQG 710
Query: 357 SHS-QEGQEDANSSGANEPEIRFGGNVNVNQDQIPQELRGAFQSVMHMFSGNAPPGQPHD 415
SHS QE QED+N SG +EPEIR GGN++VN + +P+++ GA QS+M M SG APPGQP D
Sbjct: 711 SHSHQERQEDSNGSGTSEPEIRIGGNISVNMEDMPEDITGALQSMMEMLSGAAPPGQPQD 890
Query: 416 QMNG 419
Q NG
Sbjct: 891 QTNG 902
>TC218329
Length = 672
Score = 130 bits (328), Expect = 1e-30
Identities = 63/101 (62%), Positives = 75/101 (73%), Gaps = 4/101 (3%)
Frame = +3
Query: 323 GSRSHAAPASFGSMPFNPSNLASMFMAAANGG----QGSHSQEGQEDANSSGANEPEIRF 378
GSRSH P F SMPFNP ++ASMFM N Q SHSQE QED+N SGA+EPEIR
Sbjct: 21 GSRSHGEPPPFSSMPFNPRDIASMFMNFTNATNAQQQVSHSQERQEDSNGSGASEPEIRI 200
Query: 379 GGNVNVNQDQIPQELRGAFQSVMHMFSGNAPPGQPHDQMNG 419
GGN++VN + +P+++ GAFQS+M M SG APPGQP DQMNG
Sbjct: 201 GGNISVNMEDMPEDITGAFQSMMEMLSGAAPPGQPQDQMNG 323
>BU964810 similar to GP|28416691|gb| At4g08320 {Arabidopsis thaliana},
partial (10%)
Length = 364
Score = 120 bits (300), Expect = 2e-27
Identities = 64/95 (67%), Positives = 70/95 (73%), Gaps = 2/95 (2%)
Frame = +3
Query: 1 MSHNRITTDSPLSHRIVRSFLHFLNQVEPSPGVDAEGIEVARECLVEAFKINNS--ASVT 58
M+HN I TDSPLS RIVR+FLHFLN VEP PGVDAEGIEVARECL EAFK+N S A
Sbjct: 75 MAHNYIATDSPLSRRIVRAFLHFLNSVEPGPGVDAEGIEVARECLAEAFKLNQSPVAGDD 254
Query: 59 GEPDSLIDIFKSFDANKQCERSRLDSMKASSSVSA 93
+ DSLIDIFKS +A KQCE S+ D SV A
Sbjct: 255 VKSDSLIDIFKSLEAKKQCEPSKSDVGPQPDSVDA 359
>TC215957 similar to UP|Q9SI76 (Q9SI76) F23N19.10, partial (32%)
Length = 632
Score = 84.7 bits (208), Expect = 8e-17
Identities = 43/107 (40%), Positives = 66/107 (61%)
Frame = +3
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A S Y AI ++ AIA+ + V Y NR+AAY I YT+A+ D+ +++E+ P++SK
Sbjct: 63 AFSSGDYPAAIHHFSDAIALAPTNHVLYTNRSAAYASIQNYTDALADAKKTVELKPDWSK 242
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKL 298
YSRLG A+ Y DA+ ++K L++DPNNE +K + A+ L
Sbjct: 243 GYSRLGAAHLGLSQYGDAV-SAYEKGLKIDPNNEPLKSGLADAQKAL 380
>TC206844 similar to UP|Q8H1H4 (Q8H1H4) Type 5 serine/threonine phosphatase
55 kDa isoform (Type 5 protein serine/threonine
phosphatase 55 kDa isoform), complete
Length = 1859
Score = 82.0 bits (201), Expect = 5e-16
Identities = 39/106 (36%), Positives = 68/106 (63%)
Frame = +3
Query: 195 SKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYS 254
+++Y AI+LY AI + ++AVY+ NRA A+ ++ Y AIQD+ ++IEIDP YSK Y
Sbjct: 255 ARKYSQAIDLYTQAIELNSQNAVYFSNRAFAHLRLEEYGSAIQDATKAIEIDPKYSKGYY 434
Query: 255 RLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
R G A+ G +++A+ K F++ ++ PN+ + ++ E +M+
Sbjct: 435 RRGAAHLGLGKFKEAL-KDFQQVKKMCPNDPDATKKLKECEKAVMK 569
>TC215956 similar to UP|Q9SI76 (Q9SI76) F23N19.10, partial (25%)
Length = 753
Score = 79.3 bits (194), Expect = 3e-15
Identities = 40/107 (37%), Positives = 64/107 (59%)
Frame = +2
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A S Y AI ++ IA+ + V Y NR+AAY + Y +A+ D+ +++E+ P++SK
Sbjct: 161 AFSSGDYPAAIHHFSDTIALAPSNHVLYSNRSAAYASLKNYADALADAKKTVELKPDWSK 340
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKL 298
YSRLG A+ Y DAI +K+ L++DP+NE +K + A+ L
Sbjct: 341 GYSRLGAAHLGLSQYDDAI-LAYKRGLEIDPHNEPLKSGLADAQKAL 478
>NP005041 sti (stress inducible protein)
Length = 1710
Score = 68.2 bits (165), Expect = 8e-12
Identities = 34/99 (34%), Positives = 59/99 (59%)
Frame = +1
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
+ ++Y +A + Y AI K A Y NRAA YT++ E ++D+ + IE+DP +SK Y
Sbjct: 1177 KQQKYPEATKHYTEAIKRNPKDAKAYSNRAACYTKLGAMPEGLKDAEKCIELDPTFSKGY 1356
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIR 292
+R G ++ Y A++ +++ L+ DPNN+ + + IR
Sbjct: 1357 TRKGAVQFSMKEYDKALET-YREGLKHDPNNQELLDGIR 1470
Score = 43.9 bits (102), Expect = 2e-04
Identities = 30/104 (28%), Positives = 53/104 (50%)
Frame = +1
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A + + A+ ++ AIA+ + V Y NR+AA T +++++ P++ K
Sbjct: 34 AFSAGDFAAAVRHFSDAIALSPSNHVLYSNRSAA-TLPPELRGGPSRRQKTVDLKPDWPK 210
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAE 295
AYSRLG A+ +RDA K A +P+N ++K + A+
Sbjct: 211 AYSRLGAAHLGLRRHRDA-SPPTKPASNSNPDNAALKSGLADAQ 339
>TC215954 similar to UP|Q9LNB6 (Q9LNB6) F5O11.2 (At1g12270/F5O11_1), partial
(76%)
Length = 1839
Score = 64.7 bits (156), Expect = 9e-11
Identities = 33/99 (33%), Positives = 56/99 (56%)
Frame = +3
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
+ ++Y DA++ Y +I K Y NRAA YT++ E ++D+ + IE+DP + K Y
Sbjct: 1047 KQQKYPDAVKHYTESIRRNPKDPRAYSNRAACYTKLGAMPEGLKDAEKCIELDPTFVKGY 1226
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIR 292
+R G Y Y A++ +++ L+ D NN+ + E IR
Sbjct: 1227 TRKGAVQYFMKEYDKALET-YREGLKYDSNNQELLEGIR 1340
Score = 36.2 bits (82), Expect = 0.033
Identities = 17/58 (29%), Positives = 32/58 (54%)
Frame = +2
Query: 241 RSIEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKL 298
R + + P ++ R+G A + Y DAI +K+ +++DP+NE +K + A+ L
Sbjct: 5 RQLNLSPTGARVTDRIGAALFGLSQYDDAI-LAYKRGVEIDPHNERLKFGMADAQKAL 175
>TC228229
Length = 1725
Score = 60.1 bits (144), Expect = 2e-09
Identities = 33/96 (34%), Positives = 58/96 (60%)
Frame = +3
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
+ K++ +A + Y+ +IA+ +AV Y NRA A ++ R+ EA D ++ +D Y KAY
Sbjct: 498 KQKKFKEARDCYSRSIAL-SPTAVAYANRAMANIKLRRFQEAEDDCTEALNLDDRYIKAY 674
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKE 289
SR A G ++++D + AL+L+PNN+ +K+
Sbjct: 675 SRRATARKELGKIKESMDDA-EFALRLEPNNQEIKK 779
>TC219978 weakly similar to UP|Q94GR7 (Q94GR7) Chloroplast
protein-translocon-like protein, partial (33%)
Length = 1237
Score = 59.7 bits (143), Expect = 3e-09
Identities = 33/94 (35%), Positives = 55/94 (58%)
Frame = +2
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A + +Q+ A+ Y+ AI + + YYCNRAAA+ ++ + +A +D ++I +D K
Sbjct: 581 AFKERQWSKALSYYSEAIKLNGTNTTYYCNRAAAHLKLGCFQQAAEDCGKAILLDKKNVK 760
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNE 285
AY R G A + Y +A++ FK AL L+P N+
Sbjct: 761 AYLRRGTARESLLCYEEALE-DFKHALVLEPQNK 859
>TC231862 similar to GB|AAL60196.1|18139887|AF441079 O-linked N-acetyl
glucosamine transferase {Arabidopsis thaliana;} ,
partial (25%)
Length = 750
Score = 57.4 bits (137), Expect = 1e-08
Identities = 34/98 (34%), Positives = 52/98 (52%)
Frame = +1
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
Q Y DAI YN + I +A NR Y +I R ++AIQD +R+I + P ++A+
Sbjct: 322 QQGNYVDAISCYNEVLRIDPLAADGLVNRGNTYKEIGRVSDAIQDYIRAIAVRPTMAEAH 501
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENI 291
+ L AY G+ A+ K +K+AL L P+ N+
Sbjct: 502 ANLASAYKDSGHVEAAV-KSYKQALILRPDFPEATGNL 612
Score = 33.1 bits (74), Expect = 0.28
Identities = 18/57 (31%), Positives = 28/57 (48%)
Frame = +1
Query: 226 YTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDP 282
Y + N A Q ++ + S Y+ L + Y QGNY DAI + + L++DP
Sbjct: 214 YMEWNMVAAAAQYYKATLNVTTGLSAPYNNLAIIYKQQGNYVDAI-SCYNEVLRIDP 381
Score = 28.9 bits (63), Expect = 5.3
Identities = 18/57 (31%), Positives = 27/57 (46%)
Frame = +3
Query: 201 AIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLG 257
AI Y A+A + Y N A + R EAIQ + + + PN+ +A + LG
Sbjct: 36 AILHYKQAVACDPRFLEAYNNLGNALKDVGRVEEAIQCYNQCLTLQPNHPQALTNLG 206
>TC206509 similar to UP|Q84UV7 (Q84UV7) SGT1, partial (53%)
Length = 853
Score = 53.9 bits (128), Expect = 2e-07
Identities = 31/103 (30%), Positives = 53/103 (51%)
Frame = +1
Query: 201 AIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAY 260
A++L + AI + A +Y +RA A ++N +TEA+ D+ ++IE++P+ KAY R G A
Sbjct: 211 AVDLLSQAIHLEPNKAEFYADRAQANIKLNNFTEAVADANKAIELNPSLPKAYLRKGTAC 390
Query: 261 YAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERH 303
Y A + L P+N I+ + + EE +
Sbjct: 391 MKLEEYETA-KAALEVGASLSPDNSRFATLIKECDKLIAEESY 516
>TC205511 similar to UP|Q9MAH1 (Q9MAH1) F12M16.20 (At1g53300), partial (50%)
Length = 1458
Score = 53.9 bits (128), Expect = 2e-07
Identities = 30/109 (27%), Positives = 61/109 (55%)
Frame = +2
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
+S++Y +A Y + + ++V YCNRAA + ++ ++ +I+DS +++ I PNY+KA
Sbjct: 401 KSERYTEACLAYGEGLRLDPSNSVLYCNRAACWFKLGQWERSIEDSNQALHIQPNYTKAL 580
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEER 302
R + + +A+ K ++ + PN+ V E++ A+ L + R
Sbjct: 581 LRRAASNSKLERWEEAV-KDYEILRKELPNDNEVAESLFHAQVALKKSR 724
>TC220617 weakly similar to UP|Q9SIN1 (Q9SIN1) Expressed protein
(At2g42580/F14N22.15), partial (32%)
Length = 744
Score = 49.7 bits (117), Expect = 3e-06
Identities = 27/108 (25%), Positives = 55/108 (50%)
Frame = +2
Query: 195 SKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYS 254
S + +A Y + + V YCNRA ++++ + +++QD +++ I PNY+KA
Sbjct: 173 SGMFSEACSAYGEGLKYDNSNHVLYCNRAICWSKLGLWEQSVQDCSQALNIQPNYTKALF 352
Query: 255 RLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEER 302
R + + + + K ++ + PN+ V E++R A+ L + R
Sbjct: 353 RRAASNTKLERWSEVV-KDYQALKRELPNDNEVAESLRQAQLALEKSR 493
>TC206510 similar to UP|Q84UV7 (Q84UV7) SGT1, partial (51%)
Length = 885
Score = 49.3 bits (116), Expect = 4e-06
Identities = 30/103 (29%), Positives = 51/103 (49%)
Frame = +2
Query: 201 AIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAY 260
A++L + AI + A Y +RA A ++N +TEA+ D+ ++IE++ + KAY R G A
Sbjct: 266 AVDLLSQAIHLEPNKAELYADRAQANIKLNNFTEAVADANKAIELNSSLPKAYLRKGTAC 445
Query: 261 YAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERH 303
Y A + L P+N I+ + + EE +
Sbjct: 446 MKLEEYETA-KAALEVGASLSPDNSRFATLIKECDKLIAEESY 571
>TC219285 weakly similar to UP|Q84UV7 (Q84UV7) SGT1, partial (37%)
Length = 657
Score = 48.5 bits (114), Expect = 6e-06
Identities = 24/69 (34%), Positives = 41/69 (58%)
Frame = +1
Query: 201 AIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAY 260
A++LY+ AI + A + +RA A+ ++N +TEA+ D+ +SI+++P+ KAY R A
Sbjct: 145 AVDLYSEAIRLDPNDANLFADRAQAHIKLNAFTEAVSDANKSIQLNPSLPKAYLRKATAC 324
Query: 261 YAQGNYRDA 269
Y A
Sbjct: 325 IKLQEYHTA 351
>TC212242 similar to UP|Q75LL1 (Q75LL1) Expressed protein, partial (28%)
Length = 646
Score = 46.6 bits (109), Expect = 2e-05
Identities = 37/119 (31%), Positives = 60/119 (50%), Gaps = 5/119 (4%)
Frame = +3
Query: 193 MQSKQYFDAIELYNCAI---AIYE-KSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPN 248
M K Y DAI+ Y AI A+ + ++++ + NRA + A+ DS ++++ P+
Sbjct: 153 MGKKHYSDAIDCYTRAIDQKALSDSETSILFANRAHVNLLLGNLRRALTDSNEALKLCPS 332
Query: 249 YSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME-ERHRAD 306
KA R A + +A + K LQ DPNNE +K+ R K+ E E+H A+
Sbjct: 333 NIKAIYRAAKASLSLNMLAEAREYCLK-GLQFDPNNEDLKKLDRQIGLKISEKEKHEAE 506
>TC207489 weakly similar to UP|Q94AA8 (Q94AA8) AT5g63200/MDC12_17, partial
(40%)
Length = 800
Score = 46.2 bits (108), Expect = 3e-05
Identities = 30/99 (30%), Positives = 52/99 (52%), Gaps = 2/99 (2%)
Frame = +1
Query: 195 SKQYFDAIELYNCAIAIY--EKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
S++ A E+Y A+++ +++ V N Y +Y A +S+E+ P Y+ A
Sbjct: 283 SEEPSQAEEVYKQALSLATTKQAHVILSNLGILYRHQKQYQRAKAMFTKSLELQPGYAPA 462
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENI 291
++ LGL + A+G +A F KALQ DP ++ K N+
Sbjct: 463 FNNLGLVFVAEGLLEEA-KYCFDKALQSDPLLDAAKSNL 576
>TC229760 similar to UP|Q9C566 (Q9C566) Cyclophilin-40 (Expressed protein),
partial (70%)
Length = 1164
Score = 45.8 bits (107), Expect = 4e-05
Identities = 31/101 (30%), Positives = 53/101 (51%)
Frame = +2
Query: 219 YCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKAL 278
+ N +A+ ++ A+ D+ ++ N +KA R G AY A + DA + FKKAL
Sbjct: 506 FTNSSASKLKLGDIKGALLDTEFAMREGDNNAKALFRQGQAYMALHDI-DAAVESFKKAL 682
Query: 279 QLDPNNESVKENIRVAEHKLMEERHRADHNQNSRSQEFQNH 319
L+PN+ +K+ + A K+ + R D + + S+ FQ H
Sbjct: 683 TLEPNDAGIKKELAAARKKIADRR---DLEKKAYSKMFQ*H 796
>TC205509 similar to UP|Q9MAH1 (Q9MAH1) F12M16.20 (At1g53300), partial (30%)
Length = 1218
Score = 45.8 bits (107), Expect = 4e-05
Identities = 27/94 (28%), Positives = 50/94 (52%)
Frame = +1
Query: 215 SAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGF 274
++V YCNRAA + ++ ++ +I+D +++ I P+Y+KA R + + +A+
Sbjct: 19 NSVLYCNRAACWFKLGQWERSIEDCNQALHIQPDYTKAILRRAASNSKLERWEEAVTDYE 198
Query: 275 KKALQLDPNNESVKENIRVAEHKLMEERHRADHN 308
+L +NE V EN+ A+ L + R HN
Sbjct: 199 LLRRELPDDNE-VAENLFHAQVALKKSRGEEVHN 297
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.132 0.387
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,673,311
Number of Sequences: 63676
Number of extensions: 230005
Number of successful extensions: 1207
Number of sequences better than 10.0: 78
Number of HSP's better than 10.0 without gapping: 1158
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1195
length of query: 481
length of database: 12,639,632
effective HSP length: 101
effective length of query: 380
effective length of database: 6,208,356
effective search space: 2359175280
effective search space used: 2359175280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC139354.8


