

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139354.6 - phase: 0
(342 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
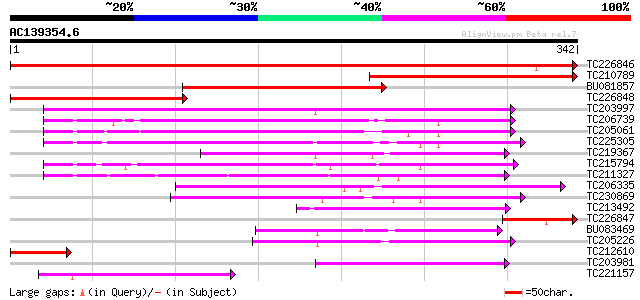
Score E
Sequences producing significant alignments: (bits) Value
TC226846 similar to UP|Q9SFE9 (Q9SFE9) T26F17.9 (GONST5 Golgi Nu... 645 0.0
TC210789 similar to UP|Q9SFE9 (Q9SFE9) T26F17.9 (GONST5 Golgi Nu... 244 5e-65
BU081857 225 3e-59
TC226848 homologue to UP|Q9SFE9 (Q9SFE9) T26F17.9 (GONST5 Golgi ... 206 2e-53
TC203997 similar to UP|Q9FFJ6 (Q9FFJ6) Phosphate/phosphoenolpyru... 179 1e-45
TC206739 similar to UP|O80688 (O80688) F8K4.1 protein, partial (... 112 3e-25
TC205061 homologue to UP|O64910 (O64910) Glucose-6-phosphate/pho... 108 3e-24
TC225305 similar to UP|CPTR_PEA (P21727) Triose phosphate/phosph... 100 7e-22
TC219367 weakly similar to UP|Q9FFJ6 (Q9FFJ6) Phosphate/phosphoe... 100 1e-21
TC215794 similar to UP|P93390 (P93390) Phosphate/phosphoenolpyru... 95 4e-20
TC211327 similar to UP|Q94JT2 (Q94JT2) AT5g17630/K10A8_110, part... 94 1e-19
TC206335 similar to GB|AAP42755.1|30984584|BT008742 At2g30460 {A... 93 2e-19
TC230869 similar to UP|P93390 (P93390) Phosphate/phosphoenolpyru... 89 3e-18
TC213492 similar to UP|Q9FYE5 (Q9FYE5) Phosphate/phosphoenolpyru... 82 5e-16
TC226847 similar to UP|Q9SFE9 (Q9SFE9) T26F17.9 (GONST5 Golgi Nu... 81 8e-16
BU083469 76 2e-14
TC205226 similar to GB|AAO11561.1|27363284|BT002645 At2g25520/F1... 71 6e-13
TC212610 similar to UP|Q9SFE9 (Q9SFE9) T26F17.9 (GONST5 Golgi Nu... 71 8e-13
TC203981 similar to UP|Q9FFJ6 (Q9FFJ6) Phosphate/phosphoenolpyru... 70 1e-12
TC221157 similar to UP|Q94EI9 (Q94EI9) AT3g14410/MLN21_19, parti... 62 4e-10
>TC226846 similar to UP|Q9SFE9 (Q9SFE9) T26F17.9 (GONST5 Golgi Nucleotide
sugar transporter), partial (97%)
Length = 1334
Score = 645 bits (1664), Expect = 0.0
Identities = 308/345 (89%), Positives = 332/345 (95%), Gaps = 3/345 (0%)
Frame = +2
Query: 1 MEEGLVCQWSVVRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAY 60
MEE V QWSV+RSLL+ILQWWAFNVTVII+NKWIFQKLDFKFPLSVSC+HFICS+IGAY
Sbjct: 146 MEESFVFQWSVIRSLLSILQWWAFNVTVIIVNKWIFQKLDFKFPLSVSCVHFICSSIGAY 325
Query: 61 VVIKVLKLKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATT 120
VVIK+LKLKPLI+VDP+DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATT
Sbjct: 326 VVIKLLKLKPLITVDPEDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATT 505
Query: 121 VVLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAE 180
VVLQWLVWRKYFDWRIWASLVPIVGGILLTS+TELSFNMFGFCAALFGCLATSTKTILAE
Sbjct: 506 VVLQWLVWRKYFDWRIWASLVPIVGGILLTSVTELSFNMFGFCAALFGCLATSTKTILAE 685
Query: 181 ALLHGYKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVL 240
+LLHGYKFDSINTVY+MAPFAT+I+ PA+LLEGNGILEW + HPYPW+A+IIIFSSGVL
Sbjct: 686 SLLHGYKFDSINTVYYMAPFATMILALPAMLLEGNGILEWLNTHPYPWSALIIIFSSGVL 865
Query: 241 AFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTF 300
AFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVL+SWLIFRNPISY+N+VGC +TLVGCTF
Sbjct: 866 AFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLVSWLIFRNPISYLNSVGCTVTLVGCTF 1045
Query: 301 YGYVRNMISQQPAVPG---TPRTPRTPRSKMELLPLVNDKLDDKV 342
YGYVR+ +SQQP VPG TPRTPRTPRSKMELLPLVNDKL+DKV
Sbjct: 1046YGYVRHKLSQQPQVPGTPRTPRTPRTPRSKMELLPLVNDKLEDKV 1180
>TC210789 similar to UP|Q9SFE9 (Q9SFE9) T26F17.9 (GONST5 Golgi Nucleotide
sugar transporter), partial (33%)
Length = 502
Score = 244 bits (622), Expect = 5e-65
Identities = 114/125 (91%), Positives = 122/125 (97%)
Frame = +3
Query: 218 LEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWL 277
LEW S HPYPW+A+IIIFSSGVLAFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVL+SWL
Sbjct: 9 LEWLSTHPYPWSALIIIFSSGVLAFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLVSWL 188
Query: 278 IFRNPISYMNAVGCAITLVGCTFYGYVRNMISQQPAVPGTPRTPRTPRSKMELLPLVNDK 337
IFRNPISY+N+VGCA+TLVGCTFYGYVR+M+SQQP VPGTPRTPRTPRSKMELLPLVNDK
Sbjct: 189 IFRNPISYLNSVGCAVTLVGCTFYGYVRHMLSQQPPVPGTPRTPRTPRSKMELLPLVNDK 368
Query: 338 LDDKV 342
LDDKV
Sbjct: 369 LDDKV 383
>BU081857
Length = 400
Score = 225 bits (573), Expect = 3e-59
Identities = 107/124 (86%), Positives = 115/124 (92%), Gaps = 1/124 (0%)
Frame = +2
Query: 105 PVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCA 164
PVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASL+PIVGGILLTS+TELSFN FGFCA
Sbjct: 2 PVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASLIPIVGGILLTSVTELSFNAFGFCA 181
Query: 165 ALFGCLATSTKTILAEALLHGYKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSI- 223
AL GCLATSTKTILAE+LLHGYKFDSINTVY+MAPFAT+I+ PALLLEGNG+ W +
Sbjct: 182 ALLGCLATSTKTILAESLLHGYKFDSINTVYYMAPFATMILAIPALLLEGNGVGXWA*VT 361
Query: 224 HPYP 227
HPYP
Sbjct: 362 HPYP 373
>TC226848 homologue to UP|Q9SFE9 (Q9SFE9) T26F17.9 (GONST5 Golgi Nucleotide
sugar transporter), partial (29%)
Length = 463
Score = 206 bits (523), Expect = 2e-53
Identities = 96/107 (89%), Positives = 104/107 (96%)
Frame = +3
Query: 1 MEEGLVCQWSVVRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAY 60
MEE V QWSV+RSLL+ILQWWAFNVTVII+NKWIFQKLDFKFPLSVSC+HFICS+IGAY
Sbjct: 141 MEESFVFQWSVIRSLLSILQWWAFNVTVIIVNKWIFQKLDFKFPLSVSCVHFICSSIGAY 320
Query: 61 VVIKVLKLKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVS 107
VVIK+LKLKPLI+VDP+DRWRRIFPMSFVFCINIVLGNVSLRYIPVS
Sbjct: 321 VVIKLLKLKPLITVDPEDRWRRIFPMSFVFCINIVLGNVSLRYIPVS 461
>TC203997 similar to UP|Q9FFJ6 (Q9FFJ6) Phosphate/phosphoenolpyruvate
translocator protein-like, partial (97%)
Length = 1271
Score = 179 bits (455), Expect = 1e-45
Identities = 94/287 (32%), Positives = 168/287 (57%), Gaps = 2/287 (0%)
Frame = +1
Query: 21 WWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQDRW 80
W++ N+ V+++NK++ FK+P+ ++ H ++ +YV I LK+ P+ ++ + ++
Sbjct: 121 WYSSNIGVLLLNKYLLSNYGFKYPIFLTMCHMTACSLFSYVAIAWLKMVPMQTIRSRLQF 300
Query: 81 RRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASL 140
+I +S +FC ++V GNVSLRY+PVSF Q + + TP T V +++ K W + +L
Sbjct: 301 LKIAALSLIFCFSVVFGNVSLRYLPVSFNQAVGATTPFFTAVFAYVMTFKREAWLTYLTL 480
Query: 141 VPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL--HGYKFDSINTVYHMA 198
VP+V G+++ S E SF++FGF + A + K++L LL G K +S+N + +MA
Sbjct: 481 VPVVTGVVIASGGEPSFHLFGFIVCIAATAARALKSVLQGILLSSEGEKLNSMNLLLYMA 660
Query: 199 PFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHSTTAV 258
P A + ++ L++E N + ++ + + + LA+ +N + F V T+A+
Sbjct: 661 PIAVVFLLPATLIMEENVVGITLALARDDVKIIWYLLFNSALAYFVNLTNFLVTKHTSAL 840
Query: 259 TFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVR 305
T V GN K AVAV++S LIFRNP+S +G ++T++G Y +
Sbjct: 841 TLQVLGNAKGAVAVVVSILIFRNPVSVTGMMGYSLTVLGVVLYSQAK 981
>TC206739 similar to UP|O80688 (O80688) F8K4.1 protein, partial (77%)
Length = 1668
Score = 112 bits (279), Expect = 3e-25
Identities = 78/296 (26%), Positives = 141/296 (47%), Gaps = 11/296 (3%)
Frame = +1
Query: 21 WWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYV--VIKVLKLKPLISVDPQD 78
WWA NV I NK + F +P S + ++ V +V ++ P +++D
Sbjct: 517 WWALNVVFNIYNKKVLNA--FPYPWLTSTLSLAAGSLMMLVSWATRVAEV-PKVNLD--- 678
Query: 79 RWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWA 138
W+ +FP++ I V VS+ + VSF IKS PA +V++ + + F ++
Sbjct: 679 FWKALFPVAVAHTIGHVAATVSMSKVAVSFTHIIKSGEPAFSVLVSRFLLGEAFPMPVYL 858
Query: 139 SLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHMA 198
SL+PI+GG L ++TEL+FNM GF A+ LA + I ++ + G +N ++
Sbjct: 859 SLLPIIGGCALAAVTELNFNMIGFMGAMISNLAFVFRNIFSKKGMKGMSVSGMNYYACLS 1038
Query: 199 PFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHSTTA- 257
+ LI+ A+ +EG + W + W + + + S+FY +++ +
Sbjct: 1039IMSLLILTPFAIAVEGPKV--WIA----GWQTAVSQIGPNFVWWVAAQSVFYHLYNQVSY 1200
Query: 258 --------VTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVR 305
+TF++ +K ++ S LIF P+ +NA+G AI ++G Y +
Sbjct: 1201MSLDQISPLTFSIGNTMKRISVIVSSILIFHTPVQPINALGAAIAILGTFLYSQAK 1368
>TC205061 homologue to UP|O64910 (O64910)
Glucose-6-phosphate/phosphate-translocator precursor,
partial (89%)
Length = 1898
Score = 108 bits (270), Expect = 3e-24
Identities = 77/298 (25%), Positives = 138/298 (45%), Gaps = 13/298 (4%)
Frame = +1
Query: 21 WWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQDRW 80
WWA NV I NK + + +P S + C ++ ++ + DP+ W
Sbjct: 682 WWALNVVFNIYNKKVLNA--YPYPWLTSTLSLACGSL-MMLISWATGIAEAPKTDPEF-W 849
Query: 81 RRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASL 140
+ +FP++ I V VS+ + VSF IKS PA +V++ + + F ++ SL
Sbjct: 850 KSLFPVAVAHTIGHVAATVSMSKVAVSFTHIIKSGEPAFSVLVSRFLLGESFPVPVYLSL 1029
Query: 141 VPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHMAPF 200
+PI+GG L ++TEL+FNM GF A+ LA + I ++ + G +N ++
Sbjct: 1030IPIIGGCALAAVTELNFNMIGFMGAMISNLAFVFRNIFSKKGMKGKSVSGMNYYACLSIL 1209
Query: 201 ATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGV----LAFCLNFSIFYVIHSTT 256
+ I+ A+ +EG P WAA S + + + S+FY +++
Sbjct: 1210SLAILTPFAIAVEG----------PQMWAAGWQTAMSQIGPQFIWWLAAQSVFYHLYNQV 1359
Query: 257 A---------VTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVR 305
+ +TF++ +K ++ S +IF P+ +NA+G AI ++G Y +
Sbjct: 1360SYMSLDQISPLTFSIGNTMKRISVIVSSIIIFHTPVQPINALGAAIAILGTFLYSQAK 1533
>TC225305 similar to UP|CPTR_PEA (P21727) Triose phosphate/phosphate
translocator, chloroplast precursor (CTPT) (p36) (E30),
complete
Length = 1707
Score = 100 bits (250), Expect = 7e-22
Identities = 77/303 (25%), Positives = 139/303 (45%), Gaps = 12/303 (3%)
Frame = +3
Query: 21 WWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQDRW 80
W+ NV I+NK I+ F +P VS IH AY ++ P + +
Sbjct: 411 WYFLNVIFNILNKKIYNY--FPYPYFVSVIHLFVGV--AYCLVSWAVGLPKRAPIDSNLL 578
Query: 81 RRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASL 140
+ + P++ + V NVS + VSF TIK+ P + + +W SL
Sbjct: 579 KLLIPVAVCHALGHVTSNVSFAAVAVSFTHTIKALEPFFNAAASQFILGQSIPITLWLSL 758
Query: 141 VPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHMAPF 200
P+V G+ + S+TELSFN GF +A+ ++ + ++I ++ + DS N +++
Sbjct: 759 APVVIGVSMASLTELSFNWVGFISAMISNISFTYRSIYSKKAM--TDMDSTNIYAYISII 932
Query: 201 ATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNF---SIFYVIHSTTA 257
A ++ + PA++LEG +L+ H + A I G++ F + +FY +++ A
Sbjct: 933 ALIVCIPPAVILEGPTLLK----HGFNDA----IAKVGLVKFVSDLFWVGMFYHLYNQVA 1088
Query: 258 ---------VTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVRNMI 308
+T V LK + S ++F N IS +G AI + G Y +++ +
Sbjct: 1089TNTLERVAPLTHAVGNVLKRVFVIGFSIIVFGNKISTQTGIGTAIAIAGVALYSFIKARM 1268
Query: 309 SQQ 311
++
Sbjct: 1269EEE 1277
>TC219367 weakly similar to UP|Q9FFJ6 (Q9FFJ6) Phosphate/phosphoenolpyruvate
translocator protein-like, partial (65%)
Length = 939
Score = 100 bits (249), Expect = 1e-21
Identities = 60/191 (31%), Positives = 104/191 (54%), Gaps = 5/191 (2%)
Frame = +1
Query: 116 TPATTVVLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTK 175
TP T + +L+ K ++ +L+P+V GI++ S +E F++FGF + + K
Sbjct: 10 TPFFTAIFAFLITCKKETGEVYLALLPVVFGIVVASNSEPLFHLFGFLVCVGSTAGRALK 189
Query: 176 TILAEALL--HGYKFDSINTVYHMAPFATLIMVFPALLLEGNGI---LEWFSIHPYPWAA 230
+++ LL K S+N + +MAP A +I++ L +EGN + +E P+
Sbjct: 190 SVVQGILLTSEAEKLHSMNLLLYMAPLAAMILLPFTLYIEGNVLALTIEKAKGDPF---I 360
Query: 231 MIIIFSSGVLAFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVG 290
+ ++ + +A+ +N + F V T+A+T V GN K AVA ++S LIFRNP++ M G
Sbjct: 361 VFLLLGNATVAYLVNLTNFLVTKHTSALTLQVLGNAKAAVAAVVSVLIFRNPVTVMGMAG 540
Query: 291 CAITLVGCTFY 301
IT++G Y
Sbjct: 541 FGITIMGVVLY 573
>TC215794 similar to UP|P93390 (P93390) Phosphate/phosphoenolpyruvate
translocator, partial (76%)
Length = 1693
Score = 95.1 bits (235), Expect = 4e-20
Identities = 79/297 (26%), Positives = 146/297 (48%), Gaps = 10/297 (3%)
Frame = +3
Query: 21 WWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKL----KPLISVDP 76
W+ FN+ I NK + + F +P++V+ + F A+G +V + L +P +S
Sbjct: 489 WYLFNIYFNIYNKQVLKA--FHYPVTVTVVQF---AVGTVLVAFMWGLNLYKRPKLS--- 644
Query: 77 QDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRI 136
I P++ V + + N+SL + VSF TIK+ P +V+L + ++ +
Sbjct: 645 GAMLGAILPLAAVHTLGNLFTNMSLGKVAVSFTHTIKAMEPFFSVILSAMFLGEFPTPWV 824
Query: 137 WASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSIN--TV 194
SLVPIVGG+ L S+TE SFN GF +A+ + ++ +L++ + K DS++ T+
Sbjct: 825 VGSLVPIVGGVALASVTEASFNWAGFWSAMASNVTNQSRNVLSKKAM-VKKEDSMDNITL 1001
Query: 195 YHMAPFATLIMVFP-ALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNF---SIFY 250
+ + + ++ P A+ +EG + + + S + A C + +
Sbjct: 1002FSIITVMSFFLLAPVAIFMEGVKFTPAY-LQSAGVNVRQLYIRSLLAALCFHAYQQVSYM 1178
Query: 251 VIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVRNM 307
++ + VT +V +K V ++ S + F+ P+S +NA G AI L G Y V+ +
Sbjct: 1179ILQRVSPVTHSVGNCVKRVVVIVSSVIFFQTPVSPVNAFGTAIALAGVFLYSRVKRI 1349
>TC211327 similar to UP|Q94JT2 (Q94JT2) AT5g17630/K10A8_110, partial (72%)
Length = 1224
Score = 93.6 bits (231), Expect = 1e-19
Identities = 79/287 (27%), Positives = 136/287 (46%), Gaps = 6/287 (2%)
Frame = +1
Query: 21 WWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQDRW 80
W+ N+ I NK + F FP ++ +I +V+ LKL+P +
Sbjct: 115 WYFQNIVFNIYNKKVLNI--FPFPWLLASFQLFVGSIWM-LVLWSLKLQPCPKISKPFII 285
Query: 81 RRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASL 140
+ P F I + VS + VSF IKS P + + ++ KY ++W S+
Sbjct: 286 ALLGPALF-HTIGHISACVSFSKVAVSFTHVIKSAEPVFSGMFSSVLGDKY-PIQVWLSI 459
Query: 141 VPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYK-FDSINTVYHMAP 199
+PIV G L ++TE+SFN+ G AL + + I ++ L +K D +N +Y
Sbjct: 460 LPIVLGCSLAAVTEVSFNVQGLWCALISNVGFVLRNIYSKRSLQNFKEVDGLN-LYGWIT 636
Query: 200 FATLIMVFP-ALLLEGNGILEWF--SIHPYPWAAMII--IFSSGVLAFCLNFSIFYVIHS 254
+L+ +FP A+ +EG+ + + +I A+ + SGV N S + +
Sbjct: 637 ILSLLYLFPVAIFVEGSQWIPGYYKAIEAIGKASTFYTWVLVSGVFYHLYNQSSYQALDE 816
Query: 255 TTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFY 301
+ +TF+V +K V ++ S L+FRNP+ +N +G AI ++G Y
Sbjct: 817 ISPLTFSVGNTMKRVVVIVSSVLVFRNPVRPLNGLGSAIAILGTFLY 957
>TC206335 similar to GB|AAP42755.1|30984584|BT008742 At2g30460 {Arabidopsis
thaliana;} , partial (60%)
Length = 1150
Score = 92.8 bits (229), Expect = 2e-19
Identities = 64/240 (26%), Positives = 114/240 (46%), Gaps = 5/240 (2%)
Frame = +1
Query: 101 LRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFNMF 160
L + V F Q K TV+L+ + + K F RI +L ++ G+ + ++T+L N
Sbjct: 1 LGFNSVGFYQMTKLAIIPCTVLLEIIFFGKRFSKRIQFALSILLLGVGIATVTDLQLNAL 180
Query: 161 GFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHMAPF--ATLIMVFPAL--LLEGNG 216
G + + T I+ + YK S +Y P+ ATL++ P L LL
Sbjct: 181 GSFLSFLAVITTCVAQIMTNTIQKKYKVSSTQLLYQSCPYQAATLLISGPYLDKLLTNQN 360
Query: 217 ILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISW 276
+ + Y + I S +++ +NFS F VI T+ VT+ V G+LK + + +
Sbjct: 361 VFGF----NYTTQVTVFIILSCLISISVNFSTFLVIGKTSPVTYQVLGHLKTCLVLAFGY 528
Query: 277 LIFRNPISYMNAVGCAITLVGCTFYGYVRNMISQQPAVPGTPRTPRTPRSKM-ELLPLVN 335
++ R+P S+ N +G I ++G Y Y + +QQ V ++ + +++ E PL+N
Sbjct: 529 ILLRDPFSWRNILGILIAMIGMILYSYYCTLENQQKTVEAASQSSQCFQAREDESDPLMN 708
>TC230869 similar to UP|P93390 (P93390) Phosphate/phosphoenolpyruvate
translocator, partial (53%)
Length = 1012
Score = 89.0 bits (219), Expect = 3e-18
Identities = 67/225 (29%), Positives = 115/225 (50%), Gaps = 11/225 (4%)
Frame = +2
Query: 98 NVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSF 157
N+SL + VSF TIK+ P TVV L+ + + + +SLVP+VGG+ L S+TE+SF
Sbjct: 5 NISLGKVAVSFTHTIKAMEPFFTVVXXALLLGEMPTFWVVSSLVPVVGGVALASMTEVSF 184
Query: 158 NMFGFCAALFGCLATSTKTILAEALLHGYK--FDSINTVYHMAPFATLIMVFPALLLEGN 215
N GF A+ + ++ +L++ L+ + D+IN + + L++V A+L+EG
Sbjct: 185 NWIGFTTAMASNVTNQSRNVLSKKLMTNEEETLDNINLYSVITIISFLLLVPCAILVEG- 361
Query: 216 GILEWFSIHPYPWAA------MIIIFSSGVLAFCLNF---SIFYVIHSTTAVTFNVAGNL 266
FS +AA + S + AFC + + ++ + VT +V +
Sbjct: 362 ---VKFSPSYLQFAASQGLNVRELCVRSILAAFCYHAYQQVSYMILQMVSPVTHSVGNCV 532
Query: 267 KVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVRNMISQQ 311
K V ++ S + F+ P+S +N +G + LVG Y + + S Q
Sbjct: 533 KRVVVIVSSVIFFQIPVSPVNTLGTGLALVGVFLYSRAKRIKSVQ 667
>TC213492 similar to UP|Q9FYE5 (Q9FYE5) Phosphate/phosphoenolpyruvate
translocator-like protein, partial (42%)
Length = 430
Score = 81.6 bits (200), Expect = 5e-16
Identities = 45/129 (34%), Positives = 78/129 (59%)
Frame = +1
Query: 174 TKTILAEALLHGYKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMII 233
T++IL + G K +S+N + +M+P A L+++ AL++E N + ++ + ++
Sbjct: 1 TRSILLSS--EGEKLNSMNLLLYMSPIAVLVLLPAALIMEPNVVDVTLTLAKDHKSMWLL 174
Query: 234 IFSSGVLAFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAI 293
+F + V+A+ N + F V T+A+T V GN K AVAV+IS L+FRNP++ + G I
Sbjct: 175 LFLNSVIAYAANLTNFLVTKHTSALTLQVLGNAKGAVAVVISILLFRNPVTVLGMGGYTI 354
Query: 294 TLVGCTFYG 302
T++G YG
Sbjct: 355 TVMGVAAYG 381
>TC226847 similar to UP|Q9SFE9 (Q9SFE9) T26F17.9 (GONST5 Golgi Nucleotide
sugar transporter), partial (12%)
Length = 283
Score = 80.9 bits (198), Expect = 8e-16
Identities = 38/48 (79%), Positives = 42/48 (87%), Gaps = 3/48 (6%)
Frame = +3
Query: 298 CTFYGYVRNMISQQPAVPGTPRTPR---TPRSKMELLPLVNDKLDDKV 342
C FYGYVR+ +SQQP +PGTPRTPR TPRSKMELLPLVNDKL+DKV
Sbjct: 6 CRFYGYVRHKLSQQPQIPGTPRTPRTPRTPRSKMELLPLVNDKLEDKV 149
>BU083469
Length = 448
Score = 76.3 bits (186), Expect = 2e-14
Identities = 54/152 (35%), Positives = 82/152 (53%), Gaps = 3/152 (1%)
Frame = +1
Query: 149 LTSITELSFNMFGFCAALFGCLATSTKTILAEALLH--GYKFDSINTVYHMAPFATLIMV 206
++S E+ FN+ G + G A + + +L + LL G + I ++Y++AP + + +
Sbjct: 1 ISSYGEIHFNIVGTVYQVTGIFAEALRLVLTQVLLQKKGLTLNPITSLYYIAPCSFVFLF 180
Query: 207 FPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHSTTAVTFNVAGNL 266
P LLE ++E I W I FS+ + A LNFSIF VI T AVT VAG L
Sbjct: 181 VPWYLLE-KPVMEVSQIQFNFW----IFFSNAICALALNFSIFLVIGRTGAVTIRVAGVL 345
Query: 267 KVAVAVLISWLIF-RNPISYMNAVGCAITLVG 297
K + + +S +IF + I+ +N VG AI L G
Sbjct: 346 KDWILIALSTVIFPESTITGLNIVGYAIALCG 441
>TC205226 similar to GB|AAO11561.1|27363284|BT002645 At2g25520/F13B15.18
{Arabidopsis thaliana;} , partial (52%)
Length = 970
Score = 71.2 bits (173), Expect = 6e-13
Identities = 37/161 (22%), Positives = 82/161 (49%), Gaps = 2/161 (1%)
Frame = +2
Query: 147 ILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLH--GYKFDSINTVYHMAPFATLI 204
+ + + E F+ +G L +T+ +L + LL+ G + I ++Y++AP +
Sbjct: 2 VAVAAYGEAKFDAWGVTXQLMAVAFEATRLVLIQILLNSKGISLNPITSLYYIAPCCLVF 181
Query: 205 MVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHSTTAVTFNVAG 264
+ P +++E + + S H I ++ AF LN ++F ++ T+A+T NVAG
Sbjct: 182 LSVPWIIMEYPSLRDNSSFH----LDFAIFGTNSACAFALNLAVFLLVGKTSALTMNVAG 349
Query: 265 NLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVR 305
+K + + SW + ++ ++ +N +G + +G +Y + +
Sbjct: 350 VVKDWLLIAFSWSVIKDTVTPINLIGYGLAFLGVAYYNHCK 472
>TC212610 similar to UP|Q9SFE9 (Q9SFE9) T26F17.9 (GONST5 Golgi Nucleotide
sugar transporter), partial (9%)
Length = 548
Score = 70.9 bits (172), Expect = 8e-13
Identities = 31/37 (83%), Positives = 34/37 (91%)
Frame = +3
Query: 1 MEEGLVCQWSVVRSLLAILQWWAFNVTVIIMNKWIFQ 37
MEE V QWSV+RSLL+ILQWWAFNVTVII+NKWIFQ
Sbjct: 135 MEESFVFQWSVIRSLLSILQWWAFNVTVIIVNKWIFQ 245
>TC203981 similar to UP|Q9FFJ6 (Q9FFJ6) Phosphate/phosphoenolpyruvate
translocator protein-like, partial (42%)
Length = 634
Score = 70.1 bits (170), Expect = 1e-12
Identities = 39/117 (33%), Positives = 66/117 (56%)
Frame = +3
Query: 185 GYKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCL 244
G K +S+N + +MAP A + ++ L++E N + ++ + + + LA+ +
Sbjct: 15 GEKLNSMNLLLYMAPMAVVFLLPATLIMEENVVGITLALARDDSKIIWYLLFNSALAYFV 194
Query: 245 NFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFY 301
N + F V T+A+T V GN K AVAV++S LIFRNP+S +G ++T+ G Y
Sbjct: 195 NLTNFLVTKHTSALTLQVLGNAKGAVAVVVSILIFRNPVSVTGMMGYSLTVFGVILY 365
>TC221157 similar to UP|Q94EI9 (Q94EI9) AT3g14410/MLN21_19, partial (60%)
Length = 661
Score = 62.0 bits (149), Expect = 4e-10
Identities = 33/122 (27%), Positives = 65/122 (53%), Gaps = 3/122 (2%)
Frame = +3
Query: 18 ILQWWAFNVTVIIMNKWIF--QKLDFKFPLSVSCIHFICSAIGAYVVIKVLK-LKPLISV 74
IL + A + I NKW+ ++++F +PL ++ +H + S++ +V+ K+LK +K +
Sbjct: 66 ILLYIALSSGQIFFNKWVLSSKEINFPYPLGLTLLHMVFSSVLCFVLTKILKVMKVEEGM 245
Query: 75 DPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDW 134
P+ + P+ +F + + LGN + YI V+F Q +K+ P VL ++ W
Sbjct: 246 TPEIYATSVVPIGAMFAMTLWLGNTAYLYISVAFAQMLKAIMPVAVFVLGVAAGLEFTKW 425
Query: 135 RI 136
+
Sbjct: 426 EV 431
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.331 0.142 0.464
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,534,734
Number of Sequences: 63676
Number of extensions: 325538
Number of successful extensions: 2534
Number of sequences better than 10.0: 81
Number of HSP's better than 10.0 without gapping: 2484
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2511
length of query: 342
length of database: 12,639,632
effective HSP length: 98
effective length of query: 244
effective length of database: 6,399,384
effective search space: 1561449696
effective search space used: 1561449696
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC139354.6


