

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139354.3 + phase: 0
(282 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
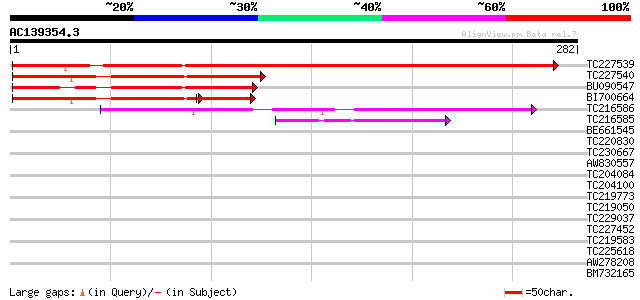
Score E
Sequences producing significant alignments: (bits) Value
TC227539 similar to UP|Q84WK2 (Q84WK2) At1g44770, partial (34%) 396 e-111
TC227540 similar to UP|Q84WK2 (Q84WK2) At1g44770, partial (18%) 160 5e-40
BU090547 similar to GP|27808560|gb| At1g44770 {Arabidopsis thali... 135 3e-32
BI700664 similar to GP|27808560|gb| At1g44770 {Arabidopsis thali... 110 4e-32
TC216586 similar to UP|Q6K819 (Q6K819) MADS box interactor-like,... 72 2e-13
TC216585 similar to UP|Q6K819 (Q6K819) MADS box interactor-like,... 42 2e-04
BE661545 37 0.013
TC220830 similar to UP|Q6PQQ5 (Q6PQQ5) AP2/EREBP transcription f... 29 0.017
TC230667 weakly similar to UP|AGA1_YEAST (P32323) A-agglutinin a... 35 0.038
AW830557 34 0.085
TC204084 UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-binding protein, co... 26 0.12
TC204100 homologue to UP|Q93WA5 (Q93WA5) Ribosomal protein L32, ... 26 0.16
TC219773 similar to UP|Q9VQA9 (Q9VQA9) CG15388-PA, partial (3%) 33 0.19
TC219050 similar to UP|AHM2_ARATH (O64474) Potential cadmium/zin... 33 0.19
TC229037 similar to GB|AAM16226.1|20334730|AY093965 At1g15730/F7... 32 0.25
TC227452 UP|Q9XGS2 (Q9XGS2) Ni-binding urease accessory protein ... 32 0.42
TC219583 weakly similar to UP|GRP2_NICSY (P27484) Glycine-rich p... 32 0.42
TC225618 similar to UP|Q9LSQ5 (Q9LSQ5) 1,4-benzoquinone reductas... 31 0.55
AW278208 31 0.55
BM732165 homologue to SP|Q9TV66|HCN4 Potassium/sodium hyperpolar... 30 0.94
>TC227539 similar to UP|Q84WK2 (Q84WK2) At1g44770, partial (34%)
Length = 1189
Score = 396 bits (1018), Expect = e-111
Identities = 201/273 (73%), Positives = 229/273 (83%), Gaps = 1/273 (0%)
Frame = +3
Query: 2 RTHNAVEMAKTVLEVADVAWTAVEY-HHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLEAL 60
+T +VE+AKTVLEVADVAWTAVE+ HH+ HR PV + CPSDQDLE+L
Sbjct: 138 QTMKSVEVAKTVLEVADVAWTAVEHTHHYRHRCSHPVAPA------AADVCPSDQDLESL 299
Query: 61 KLENRRLRNLLDQNLKLLNNLSESNSFLSNCPPDLHVRLAATVRSDEYLTRLKVLQEETA 120
+ ENRRLRNLLD NL LL NLS+S L NCPPDL+ RL AT+RSDEYLTRLK LQ+ETA
Sbjct: 300 RSENRRLRNLLDLNLNLLQNLSKSPC-LDNCPPDLNERLVATMRSDEYLTRLKFLQQETA 476
Query: 121 SGGNQFPFKEATEVDYQSADILINVDSKEPSWWVWVADETDPINFEECSGIDDESYLIIS 180
SGGNQFPFKEATE++ +SADIL+N+DS+EPSWWVWV DE DPIN EE SGIDDE+Y++IS
Sbjct: 477 SGGNQFPFKEATELEGRSADILVNIDSQEPSWWVWVTDEVDPINVEELSGIDDENYVVIS 656
Query: 181 EEHVVDGVANFMARCIMSNPKARNMSPEELQNNLSKAFAGTSKLEKVLDIWAAGKLFYAL 240
EEHVVDGVANFMARCI+SNPKA N SPEELQ LSKA GT+KLEK+LDIW AGKLFY L
Sbjct: 657 EEHVVDGVANFMARCILSNPKALNFSPEELQKALSKALRGTTKLEKILDIWEAGKLFYCL 836
Query: 241 STWGLALAGLYQTRSLLKVAAKGVHSGGKLALK 273
STWGLALAGLYQ+R++L+VAAKGVHSG KLALK
Sbjct: 837 STWGLALAGLYQSRAILRVAAKGVHSGSKLALK 935
>TC227540 similar to UP|Q84WK2 (Q84WK2) At1g44770, partial (18%)
Length = 477
Score = 160 bits (406), Expect = 5e-40
Identities = 87/128 (67%), Positives = 97/128 (74%), Gaps = 2/128 (1%)
Frame = +1
Query: 2 RTHNAVEMAKTVLEVADVAWTAVEYHHH--HHRHHDPVVTHDDKYDDKDNNCPSDQDLEA 59
+T +VE+AKTVLEVADVAWTAVE+ HH H H P D CPSDQDLE+
Sbjct: 118 QTMKSVEVAKTVLEVADVAWTAVEHTHHLRHRSHPVPAAPAADV-------CPSDQDLES 276
Query: 60 LKLENRRLRNLLDQNLKLLNNLSESNSFLSNCPPDLHVRLAATVRSDEYLTRLKVLQEET 119
L+ ENRRLRNLLDQNLKLL NLSE+ F NCPPDL+ RL AT+RSDEYLTRLK LQ+ET
Sbjct: 277 LRSENRRLRNLLDQNLKLLQNLSEATCF-DNCPPDLNDRLVATMRSDEYLTRLKYLQQET 453
Query: 120 ASGGNQFP 127
ASGGNQFP
Sbjct: 454 ASGGNQFP 477
>BU090547 similar to GP|27808560|gb| At1g44770 {Arabidopsis thaliana},
partial (16%)
Length = 420
Score = 135 bits (339), Expect = 3e-32
Identities = 78/122 (63%), Positives = 88/122 (71%)
Frame = +1
Query: 2 RTHNAVEMAKTVLEVADVAWTAVEYHHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLEALK 61
+T +VE+AKTVLEVADVAWTAVE H P+ D CPSDQDLE+L+
Sbjct: 100 QTMKSVEVAKTVLEVADVAWTAVE-------HTPPLPPAADV-------CPSDQDLESLR 237
Query: 62 LENRRLRNLLDQNLKLLNNLSESNSFLSNCPPDLHVRLAATVRSDEYLTRLKVLQEETAS 121
ENRRLRNLLD NL LL NLS+S L NCPPDL+ RL AT+RSDEYLTRLK LQ+ETAS
Sbjct: 238 SENRRLRNLLDLNLNLLQNLSKSPC-LDNCPPDLNERLVATMRSDEYLTRLKFLQQETAS 414
Query: 122 GG 123
GG
Sbjct: 415 GG 420
>BI700664 similar to GP|27808560|gb| At1g44770 {Arabidopsis thaliana},
partial (17%)
Length = 424
Score = 110 bits (274), Expect(2) = 4e-32
Identities = 60/97 (61%), Positives = 69/97 (70%), Gaps = 2/97 (2%)
Frame = +3
Query: 2 RTHNAVEMAKTVLEVADVAWTAVEYHHH--HHRHHDPVVTHDDKYDDKDNNCPSDQDLEA 59
+T +VE+AKTVLEVADVAWTAVE+ HH H H P D CPSDQDLE+
Sbjct: 57 QTMKSVEVAKTVLEVADVAWTAVEHTHHLRHRSHPVPAAPAADV-------CPSDQDLES 215
Query: 60 LKLENRRLRNLLDQNLKLLNNLSESNSFLSNCPPDLH 96
L+ ENRRLRNLLDQNLKLL NLSE+ F NCPPD++
Sbjct: 216 LRSENRRLRNLLDQNLKLLQNLSEATCF-DNCPPDVY 323
Score = 45.4 bits (106), Expect(2) = 4e-32
Identities = 22/29 (75%), Positives = 26/29 (88%)
Frame = +2
Query: 94 DLHVRLAATVRSDEYLTRLKVLQEETASG 122
+L+ RL AT+RSDEYLTRLK LQ+ETASG
Sbjct: 338 NLNDRLVATMRSDEYLTRLKYLQQETASG 424
>TC216586 similar to UP|Q6K819 (Q6K819) MADS box interactor-like, partial
(57%)
Length = 1145
Score = 72.4 bits (176), Expect = 2e-13
Identities = 53/228 (23%), Positives = 104/228 (45%), Gaps = 11/228 (4%)
Frame = +1
Query: 46 DKDNNCPSDQDLEALKLENRRLRNLLDQNLKLLNNLSESNSFLSN-----CPPDLHVRLA 100
D + + D + L+++ + L +L Q + L ++ ++S ++ C + +
Sbjct: 205 DPNPQASVESDPDPLRVKRKTLEAVLLQCQRALELINATSSTAADVDEEDCEIEGEATAS 384
Query: 101 ATVRSDEYLTRLKVLQEETASGGNQFPFKEATEVDYQSADILINVDSKEPSWWV------ 154
A +DE LK E A +++ A + N+D + SW +
Sbjct: 385 ADPDADELCDLLKSRVECPAF---------LQQLECAQASVSQNIDEEGNSWDMVSKNDL 537
Query: 155 WVADETDPINFEECSGIDDESYLIISEEHVVDGVANFMARCIMSNPKARNMSPEELQNNL 214
W + D + E Y+++ +E +V+G+A FMA ++S + ++++P +LQ+ L
Sbjct: 538 WEGERADS---------EQEDYVLVRQEDIVEGIACFMAAYLLSLKQTKDLTPIQLQSAL 690
Query: 215 SKAFAGTSKLEKVLDIWAAGKLFYALSTWGLALAGLYQTRSLLKVAAK 262
SK F+ K K+ W K+ Y +++WG G+YQ +L A K
Sbjct: 691 SKTFSVKKKKGKLRKAWDGSKVIYNVASWGATAIGIYQNPVILGAATK 834
>TC216585 similar to UP|Q6K819 (Q6K819) MADS box interactor-like, partial
(31%)
Length = 839
Score = 42.4 bits (98), Expect = 2e-04
Identities = 24/87 (27%), Positives = 49/87 (55%)
Frame = +3
Query: 133 EVDYQSADILINVDSKEPSWWVWVADETDPINFEECSGIDDESYLIISEEHVVDGVANFM 192
+++ A + N+D + SW + E D E + E Y++ +E +++G+A FM
Sbjct: 585 QLECARASVSQNIDEEGNSWDM--VSENDLWEGERVDS-EQEDYVLGRQEDIMEGIAWFM 755
Query: 193 ARCIMSNPKARNMSPEELQNNLSKAFA 219
A ++S + ++++P +LQ+ LSK F+
Sbjct: 756 AAYLLSLKQTKDLTPIQLQSALSKTFS 836
>BE661545
Length = 676
Score = 36.6 bits (83), Expect = 0.013
Identities = 15/29 (51%), Positives = 18/29 (61%)
Frame = +3
Query: 26 YHHHHHRHHDPVVTHDDKYDDKDNNCPSD 54
+HHHHH HH HD +DD D+NC D
Sbjct: 219 HHHHHHNHH-----HD--FDDHDHNCGDD 284
Score = 28.5 bits (62), Expect = 3.6
Identities = 14/37 (37%), Positives = 19/37 (50%), Gaps = 5/37 (13%)
Frame = -2
Query: 25 EYHH----HHHRHHDPVVTHDDKYDDK-DNNCPSDQD 56
+YH+ HHH + HDD DD D++C D D
Sbjct: 306 DYHYDLDCHHHNYDRDHQNHDDDCDDDGDDDCGYDND 196
>TC220830 similar to UP|Q6PQQ5 (Q6PQQ5) AP2/EREBP transcription factor,
partial (10%)
Length = 759
Score = 29.3 bits (64), Expect(2) = 0.017
Identities = 10/20 (50%), Positives = 12/20 (60%)
Frame = +1
Query: 15 EVADVAWTAVEYHHHHHRHH 34
E D + T + HHHHH HH
Sbjct: 502 ETQDSSLTHIYDHHHHHHHH 561
Score = 25.8 bits (55), Expect(2) = 0.017
Identities = 9/22 (40%), Positives = 12/22 (53%)
Frame = +1
Query: 26 YHHHHHRHHDPVVTHDDKYDDK 47
+HHHHH H D++D K
Sbjct: 541 HHHHHHHHGSTSYFGGDQHDLK 606
>TC230667 weakly similar to UP|AGA1_YEAST (P32323) A-agglutinin attachment
subunit precursor, partial (5%)
Length = 865
Score = 35.0 bits (79), Expect = 0.038
Identities = 23/77 (29%), Positives = 34/77 (43%)
Frame = -1
Query: 22 TAVEYHHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLEALKLENRRLRNLLDQNLKLLNNL 81
+ +E HHH+H HHD D D+ D+ D +L + N+ R Q ++ L
Sbjct: 517 SGLENHHHYHYHHDDDDMGGDDNDEDDDGYGLDDELVPWNVSNKFGR----QRMRKLGKR 350
Query: 82 SESNSFLSNCPPDLHVR 98
+ S S P L VR
Sbjct: 349 AFSKMHGSKKSPYLFVR 299
>AW830557
Length = 454
Score = 33.9 bits (76), Expect = 0.085
Identities = 14/34 (41%), Positives = 18/34 (52%), Gaps = 4/34 (11%)
Frame = -3
Query: 11 KTVLEVADVAWTAVEY----HHHHHRHHDPVVTH 40
K E A + W+ + + HHHHH HH V TH
Sbjct: 407 KVSAEQAQLLWSLLSHYHVLHHHHHLHHQDVPTH 306
>TC204084 UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-binding protein, complete
Length = 1937
Score = 26.2 bits (56), Expect(2) = 0.12
Identities = 14/52 (26%), Positives = 22/52 (41%), Gaps = 8/52 (15%)
Frame = -1
Query: 26 YHHHHHRHHDP--------VVTHDDKYDDKDNNCPSDQDLEALKLENRRLRN 69
++HHHH HH + H ++ D++ D L L RR R+
Sbjct: 1637 HNHHHHHHHSHRHNHRHGYGIHHRHRHRHYDHHIHRHHDCILLLLRRRRRRH 1482
Score = 25.8 bits (55), Expect(2) = 0.12
Identities = 9/29 (31%), Positives = 17/29 (58%)
Frame = -1
Query: 3 THNAVEMAKTVLEVADVAWTAVEYHHHHH 31
T N K L + ++++++ +HHHHH
Sbjct: 1751 TKNTGHNHKP*LTTSSLSFSSLHHHHHHH 1665
>TC204100 homologue to UP|Q93WA5 (Q93WA5) Ribosomal protein L32, complete
Length = 1145
Score = 25.8 bits (55), Expect(2) = 0.16
Identities = 8/13 (61%), Positives = 10/13 (76%)
Frame = +3
Query: 26 YHHHHHRHHDPVV 38
Y HH HRHHD ++
Sbjct: 327 YDHHIHRHHDCIL 365
Score = 25.8 bits (55), Expect(2) = 0.16
Identities = 9/29 (31%), Positives = 17/29 (58%)
Frame = +3
Query: 3 THNAVEMAKTVLEVADVAWTAVEYHHHHH 31
T N K L + ++++++ +HHHHH
Sbjct: 126 TKNTGHNHKP*LTTSSLSFSSLHHHHHHH 212
>TC219773 similar to UP|Q9VQA9 (Q9VQA9) CG15388-PA, partial (3%)
Length = 662
Score = 32.7 bits (73), Expect = 0.19
Identities = 14/35 (40%), Positives = 19/35 (54%)
Frame = +1
Query: 11 KTVLEVADVAWTAVEYHHHHHRHHDPVVTHDDKYD 45
K +L+V D + + HHHHH HH HD +D
Sbjct: 433 KPMLDVDDEWYNNITNHHHHHHHH-----HDMTFD 522
>TC219050 similar to UP|AHM2_ARATH (O64474) Potential
cadmium/zinc-transporting ATPase 2 , partial (4%)
Length = 589
Score = 32.7 bits (73), Expect = 0.19
Identities = 11/28 (39%), Positives = 15/28 (53%)
Frame = +1
Query: 26 YHHHHHRHHDPVVTHDDKYDDKDNNCPS 53
+HHHHHR H + D++D C S
Sbjct: 355 HHHHHHRQHQHEYHNHDQHDQHHQKCAS 438
>TC229037 similar to GB|AAM16226.1|20334730|AY093965 At1g15730/F7H2_7
{Arabidopsis thaliana;} , partial (52%)
Length = 958
Score = 32.3 bits (72), Expect = 0.25
Identities = 11/26 (42%), Positives = 17/26 (65%)
Frame = +3
Query: 25 EYHHHHHRHHDPVVTHDDKYDDKDNN 50
E+H H H HHD +HD K++ D++
Sbjct: 372 EHHDHDHHHHDHDHSHDHKHEHHDHH 449
>TC227452 UP|Q9XGS2 (Q9XGS2) Ni-binding urease accessory protein UreG,
complete
Length = 1145
Score = 31.6 bits (70), Expect = 0.42
Identities = 13/34 (38%), Positives = 19/34 (55%)
Frame = +3
Query: 25 EYHHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLE 58
++HHHHH H D HD + D D++ D + E
Sbjct: 138 DHHHHHHHHQD----HDHHHHDHDHHHHHDGEGE 227
>TC219583 weakly similar to UP|GRP2_NICSY (P27484) Glycine-rich protein 2,
partial (45%)
Length = 995
Score = 31.6 bits (70), Expect = 0.42
Identities = 27/82 (32%), Positives = 33/82 (39%)
Frame = -1
Query: 26 YHHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLEALKLENRRLRNLLDQNLKLLNNLSESN 85
YHHHHH H PV +N PSD L + + LL + L + L
Sbjct: 503 YHHHHHHH--PV--------HGNNPLPSD-----LFFHSYNTQGLLLSDRTLCHGLPRPR 369
Query: 86 SFLSNCPPDLHVRLAATVRSDE 107
S L CP H R +R DE
Sbjct: 368 SVLLLCPS--HRRRREILREDE 309
>TC225618 similar to UP|Q9LSQ5 (Q9LSQ5) 1,4-benzoquinone reductase-like; Trp
repressor binding protein-like (1,4-benzoquinone
reductase-like protein), partial (75%)
Length = 1864
Score = 31.2 bits (69), Expect = 0.55
Identities = 9/16 (56%), Positives = 10/16 (62%)
Frame = +3
Query: 21 WTAVEYHHHHHRHHDP 36
W +HHHHH HH P
Sbjct: 654 WMNHHHHHHHHHHHHP 701
>AW278208
Length = 396
Score = 31.2 bits (69), Expect = 0.55
Identities = 20/62 (32%), Positives = 29/62 (46%)
Frame = +2
Query: 26 YHHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLEALKLENRRLRNLLDQNLKLLNNLSESN 85
+HHHHH HH H ++NNC SD + +L N + ++NL L +
Sbjct: 20 HHHHHHHHHHRHRRH------RNNNC-SDDVVSSLHHLNEWAIAVPERNLLWFPTLKLFS 178
Query: 86 SF 87
SF
Sbjct: 179SF 184
>BM732165 homologue to SP|Q9TV66|HCN4 Potassium/sodium
hyperpolarization-activated cyclic nucleotide-gated
channel 4, partial (2%)
Length = 423
Score = 30.4 bits (67), Expect = 0.94
Identities = 8/13 (61%), Positives = 10/13 (76%)
Frame = +1
Query: 24 VEYHHHHHRHHDP 36
+ +HHHHH HH P
Sbjct: 28 IHHHHHHHHHHHP 66
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.131 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,183,056
Number of Sequences: 63676
Number of extensions: 186986
Number of successful extensions: 4594
Number of sequences better than 10.0: 136
Number of HSP's better than 10.0 without gapping: 2585
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3709
length of query: 282
length of database: 12,639,632
effective HSP length: 96
effective length of query: 186
effective length of database: 6,526,736
effective search space: 1213972896
effective search space used: 1213972896
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC139354.3


