

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139343.10 + phase: 0 /pseudo
(1612 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
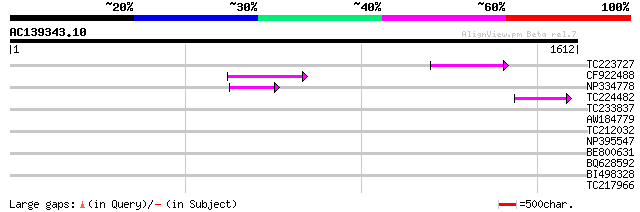
Sequences producing significant alignments: (bits) Value
TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 87 5e-17
CF922488 86 1e-16
NP334778 reverse transcriptase [Glycine max] 65 2e-10
TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 44 6e-04
TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 43 0.001
AW184779 43 0.001
TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 42 0.003
NP395547 reverse transcriptase [Glycine max] 40 0.011
BE800631 weakly similar to GP|9294065|dbj| contains similarity t... 33 1.0
BQ628592 28 1.4
BI498328 31 3.8
TC217966 homologue to UP|Q9LX13 (Q9LX13) (3R)-hydroxymyristoyl-[... 30 6.6
>TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (9%)
Length = 843
Score = 87.4 bits (215), Expect = 5e-17
Identities = 82/222 (36%), Positives = 99/222 (43%)
Frame = +2
Query: 1196 TINPGTTTSSSSC*AVSISRVLPNKIRRP*EDWPVDSC*MEILCTRETMTWYC*DVLMNM 1255
T++ G +TSS *A + + LP IR *E W S * E C RET T V M
Sbjct: 128 TVSLGISTSSDMS*AKNTCQRLPTMIRGH*EGWRPVSS*AEAYCIRETTT*NLCGVWMPG 307
Query: 1256 KRSS*CMMYMTVPSGPMLQGILCQGSCYEQVTTGWPWSMIATSTPESATNVKSMLIRFMC 1315
++ +* M G+L G EQV TG PW +I S +ATNVK I M
Sbjct: 308 RQIT*SRKSMRARLERTPTGMLWPGRS*EQVITGLPWKVIVVSM*GNATNVKRSQIMSMP 487
Query: 1316 LRTLSMLFHPHSRSQCGAST*LEELNRRLQMVIVSS*WQLTTSPSGLKQHLIPM*PSKW* 1375
R L M P S CG L+ R +MVI SS + SPSG +Q IP W
Sbjct: 488 HRIL*MSCPPLGLSPCGE*MSSGPLSPRPRMVIASSS*R*IISPSGSRQLPIPTS*GVWW 667
Query: 1376 LSSSRTTSFVDMVFPARSLPTMVQT*TTMWCKLFVKNSKLSI 1417
S R S DMV R T T* T F ++ K SI
Sbjct: 668 SGSLRKRSSADMVCQGRLSRTTAPT*ITR*WGKFARSLKSSI 793
>CF922488
Length = 741
Score = 85.9 bits (211), Expect = 1e-16
Identities = 86/227 (37%), Positives = 109/227 (47%)
Frame = +1
Query: 619 RRMVKSGCVLTSET*TKPVQKTTFHYLILMCLLIILLSLRCSPSWMVSPVTIRSKCLLKI 678
+RM + C T E *TKPVQ+ +F Y I I SPSWM V R + +I
Sbjct: 10 KRMGRCECAWTIEI*TKPVQRISFLYRISTFSWITRPVFPNSPSWMDFQVITR*R*HQRI 189
Query: 679 EKRRLLSLHGVLSATK*CCSA**MLVLPTKEE*LLCFMT*FTKKSKYMWMI*L*SQQMRS 738
KR+L L+G SA + C * ML T T* T++ + WM * *+Q+ R
Sbjct: 190 WKRQLSLLYGEPSAIRLCRLG*RMLGQHTSGPWWHYSRT*CTRR*RSTWMT*S*NQERRR 369
Query: 739 NMLSI*QRCLKG*ENTSFD*IPTNVHSASDPGSY*ASLSAKRALKSILIKSGPSEKCQLH 798
N LSI + CL NT D*IP +V +P S L A+ + I + S + H
Sbjct: 370 NTLSICESCLGDYVNTG*D*IPQSVCLR*NPESCSTLLIAREE*RWIRTR*K*SLRWPSH 549
Query: 799 RPRSKSEVSSGV*ITSPDLYLT*PQPAGRSSSYSERISLLYGMMNAK 845
RSKS+VS G * TS D Y +* A S RISL G M K
Sbjct: 550 IQRSKSKVSWGG*TTS*DSYHS*LPLASLFSYCCARISLSNGTMIVK 690
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 65.1 bits (157), Expect = 2e-10
Identities = 57/141 (40%), Positives = 73/141 (51%)
Frame = +1
Query: 625 GCVLTSET*TKPVQKTTFHYLILMCLLIILLSLRCSPSWMVSPVTIRSKCLLKIEKRRLL 684
GCV T E *T+PVQ+ F Y + L S SWM V IR + +I KR+L
Sbjct: 4 GCVWTIEI*TEPVQRIIFLYRTSTFSWLTWPVLPYSLSWMDFRVIIR*RWHQRIWKRQLS 183
Query: 685 SLHGVLSATK*CCSA**MLVLPTKEE*LLCFMT*FTKKSKYMWMI*L*SQQMRSNMLSI* 744
L+G SA +*C * +L PT T* T++S+ MWM * +Q+ R N SI
Sbjct: 184 LLYGEPSAIR*CRLG*RILGQPTIGPWWHYSRT*CTRRSRPMWMK*SRNQEWRRNT*SIC 363
Query: 745 QRCLKG*ENTSFD*IPTNVHS 765
+ CL NT D*IP +V S
Sbjct: 364 KICLGNYVNTG*D*IPGSVCS 426
>TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 669
Score = 43.9 bits (102), Expect = 6e-04
Identities = 56/166 (33%), Positives = 77/166 (45%), Gaps = 2/166 (1%)
Frame = +2
Query: 1434 PTRISRELSRRW*PLTRTGMRCYPMLCMATVLQCAVQPRQPLSLLYMVWKQFFLWKWRSH 1493
P RIS+ LS+R TR G RC T LQC Q Q S YM W+ + + +S
Sbjct: 8 PIRISKRLSKR*PCHTRIGTRCSHSRYTVTGLQCERQLGQRRSHWYMGWRLCYRLR*KSR 187
Query: 1494 LSV*SWKQSYLR-LNGARAGMIS*I*LRKNVWIPWLVDSHIKQE*RLLLTRKSILENSR* 1552
SW+ R +G + MIS LR + P ++ + +E*R+ TR+ +S
Sbjct: 188 -H*GSWQNPD*RNQSGLKRAMISSTSLRVSA*RP*VMGACTSKE*RVHSTRRYACASSMR 364
Query: 1553 GNLY*-KGG*ANNPIQGASGRLTMKVLMLSRRPSPVVL*SLHTWMV 1597
L * K + I+G +G T K L+L R P L TWMV
Sbjct: 365 ETLC*RKCPMLSRTIEG-NGPRTTKGLLL*RGLFPEEPWCLPTWMV 499
>TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 402
Score = 42.7 bits (99), Expect = 0.001
Identities = 40/109 (36%), Positives = 54/109 (48%)
Frame = +3
Query: 663 WMVSPVTIRSKCLLKIEKRRLLSLHGVLSATK*CCSA**MLVLPTKEE*LLCFMT*FTKK 722
WMVS I+ + +K+ +R L S +G SA +* S * +L P C M * +K
Sbjct: 12 WMVSRGIIKYRWHVKM*RRPLSSPYGGHSAIE*WPSG*KILGQPISVPWWRCSMI*CIRK 191
Query: 723 SKYMWMI*L*SQQMRSNMLSI*QRCLKG*ENTSFD*IPTNVHSASDPGS 771
+ M *L S +R N LSI CL+G NT+ +* N H GS
Sbjct: 192 *RST*MT*LPSLGLRPNTLSICVSCLEGCRNTN*N*TQPNAHLG*SRGS 338
>AW184779
Length = 432
Score = 42.7 bits (99), Expect = 0.001
Identities = 27/46 (58%), Positives = 31/46 (66%)
Frame = +1
Query: 1331 CGAST*LEELNRRLQMVIVSS*WQLTTSPSGLKQHLIPM*PSKW*L 1376
CGA T* E L+ RLQM I S * QLTTSP+G KQ + +* W*L
Sbjct: 292 CGA*T*SEPLSPRLQMDITSF*SQLTTSPNGSKQFRMLV*LGVW*L 429
>TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (3%)
Length = 803
Score = 41.6 bits (96), Expect = 0.003
Identities = 37/121 (30%), Positives = 48/121 (39%), Gaps = 4/121 (3%)
Frame = +3
Query: 1132 LQRLSCTIFLVMRTKWLMLLLLYPPCFE*TIGMMCQ*SKCNALKDLHMCLLLGM*SIRLV 1191
L R L + KW M L L PC * + L LG+ + +
Sbjct: 354 LMRSPSITLLERKIKWQMRLPL*CPCSS*H------------RMGTYRTLSLGVVADPHI 497
Query: 1192 KMWL----TINPGTTTSSSSC*AVSISRVLPNKIRRP*EDWPVDSC*MEILCTRETMTWY 1247
+W T++ G SS + A S LP + DW S * E CTRETMTW+
Sbjct: 498 VVWWKRNGTVSLGILISSDTLKAKSTHWRLPTTTKGERGDWQPASS*AEAYCTRETMTWF 677
Query: 1248 C 1248
C
Sbjct: 678 C 680
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 39.7 bits (91), Expect = 0.011
Identities = 33/104 (31%), Positives = 50/104 (47%)
Frame = +2
Query: 626 CVLTSET*TKPVQKTTFHYLILMCLLIILLSLRCSPSWMVSPVTIRSKCLLKIEKRRLLS 685
CVL + KP +KT H+ + L L + SW + VTIR + +L+I+K++LL
Sbjct: 167 CVLIIGSSMKPQEKTITHFPSWIKCLRDLQGNPSTVSWTDTQVTIRLQWILRIKKKQLLH 346
Query: 686 LHGVLSATK*CCSA**MLVLPTKEE*LLCFMT*FTKKSKYMWMI 729
+ V C S M +L ++ * MT K +WMI
Sbjct: 347 VLSVFLLIAACRSVYVMPLLLFRDV*WQFLMTW*RNVLKSLWMI 478
>BE800631 weakly similar to GP|9294065|dbj| contains similarity to myb
proteins~gene_id:MRC8.8 {Arabidopsis thaliana}, partial
(8%)
Length = 413
Score = 33.1 bits (74), Expect = 1.0
Identities = 16/37 (43%), Positives = 22/37 (59%)
Frame = +3
Query: 954 LLLLGRLHAGRCFCLNMILCSKLKRQSKVAFLPIILL 990
+L+L GRC+ LNM LC K + +SK L + LL
Sbjct: 117 VLILNAGRDGRCYVLNMFLCFKKQERSKGLLLVVTLL 227
>BQ628592
Length = 423
Score = 28.1 bits (61), Expect(2) = 1.4
Identities = 21/60 (35%), Positives = 26/60 (43%)
Frame = -1
Query: 913 QCLRRLVVLWPGPLNACVIIW*VIQLG*YPEWIR*SISLRKLLLLGRLHAGRCFCLNMIL 972
QC + V W G VI G +P+WI *+ SLR GR + LN IL
Sbjct: 180 QCWKGRAVPWYGRHIVLGSTCSVIPRGLFPKWIL*NTSLRSRPSRDESLGGRYYYLNSIL 1
Score = 23.1 bits (48), Expect(2) = 1.4
Identities = 12/26 (46%), Positives = 15/26 (57%)
Frame = -2
Query: 881 WDAYLVNRMRLERKNMLSTI*ARSSP 906
WDA + L ++N TI*ARS P
Sbjct: 275 WDACWFSTTTLGKRNKPFTI*ARSLP 198
>BI498328
Length = 335
Score = 31.2 bits (69), Expect = 3.8
Identities = 22/57 (38%), Positives = 27/57 (46%)
Frame = +2
Query: 1031 SGV*SLMVLSMLMVKELGQSLYPHRGITSLLPPEFCSNVQTIWLSMKHVSLGSRKQL 1087
+G+ + M ML E GQSLYP L + V TIW S KH G R+ L
Sbjct: 149 NGLFASMGHPMLWATE*GQSLYPRMISVFLSRLD*VLIVPTIWPSTKHAPSGFRRPL 319
>TC217966 homologue to UP|Q9LX13 (Q9LX13) (3R)-hydroxymyristoyl-[acyl carrier
protein] dehydratase-like protein (At5g10160), partial
(71%)
Length = 910
Score = 30.4 bits (67), Expect = 6.6
Identities = 27/100 (27%), Positives = 44/100 (44%)
Frame = -2
Query: 595 NRLMRVSSLQLSTLNGLPILYLCQRRMVKSGCVLTSET*TKPVQKTTFHYLILMCLLIIL 654
NR++R+ SL++ TLN + ++ C+ K V+K T M +
Sbjct: 903 NRMLRIISLKMHTLN*MFTMF----------CIKKQNNLIKDVRK*TLPIGRSMSTYSLP 754
Query: 655 LSLRCSPSWMVSPVTIRSKCLLKIEKRRLLSLHGVLSATK 694
++++ SPS SP T + I R SL V+ TK
Sbjct: 753 IAIKNSPSHTTSPPTYAFPSIFAI-PNRFCSLVSVILITK 637
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.352 0.154 0.547
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 85,313,102
Number of Sequences: 63676
Number of extensions: 1453651
Number of successful extensions: 16578
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 8971
Number of HSP's successfully gapped in prelim test: 651
Number of HSP's that attempted gapping in prelim test: 7045
Number of HSP's gapped (non-prelim): 10730
length of query: 1612
length of database: 12,639,632
effective HSP length: 110
effective length of query: 1502
effective length of database: 5,635,272
effective search space: 8464178544
effective search space used: 8464178544
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.5 bits)
S2: 66 (30.0 bits)
Medicago: description of AC139343.10


