

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138453.15 + phase: 0 /pseudo/partial
(195 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
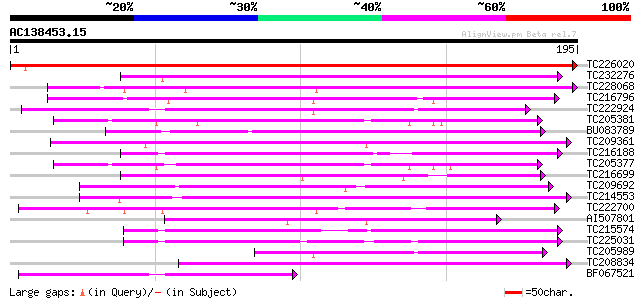
Score E
Sequences producing significant alignments: (bits) Value
TC226020 similar to UP|Q9SP53 (Q9SP53) Flavanone 3-hydroxylase, ... 323 2e-89
TC232276 similar to UP|Q9FFF6 (Q9FFF6) Leucoanthocyanidin dioxyg... 100 4e-22
TC228068 anthocyanidin synthase [Glycine max] 100 7e-22
TC216796 similar to GB|AAQ65160.1|34365697|BT010537 At4g10500 {A... 99 1e-21
TC222924 weakly similar to UP|Q9FLV0 (Q9FLV0) Flavanone 3-hydrox... 98 3e-21
TC205381 weakly similar to UP|Q86B83 (Q86B83) CG33099-PA, partia... 82 1e-16
BU083789 82 2e-16
TC209361 weakly similar to UP|Q7XJ89 (Q7XJ89) Flavonol synthase ... 82 2e-16
TC216188 similar to UP|Q84L58 (Q84L58) 1-aminocyclopropane-1-car... 79 1e-15
TC205377 79 1e-15
TC216699 weakly similar to UP|Q84MB3 (Q84MB3) At1g06620, partial... 77 4e-15
TC209692 weakly similar to UP|Q39224 (Q39224) SRG1 protein (F6I1... 75 1e-14
TC214553 similar to UP|Q39224 (Q39224) SRG1 protein (F6I1.30/F6I... 75 1e-14
TC222700 similar to GB|AAS76251.1|45752706|BT012156 At1g55290 {A... 70 6e-13
AI507801 weakly similar to GP|20260150|gb| strong similarity to ... 70 8e-13
TC215574 similar to UP|O04076 (O04076) ACC-oxidase, partial (98%) 69 1e-12
TC225031 homologue to UP|Q8W3Y3 (Q8W3Y3) 1-aminocyclopropane-1-c... 67 7e-12
TC205989 similar to UP|Q9FLV0 (Q9FLV0) Flavanone 3-hydroxylase-l... 66 9e-12
TC208834 similar to UP|Q39224 (Q39224) SRG1 protein (F6I1.30/F6I... 65 2e-11
BF067521 similar to GP|10177853|dbj flavanone 3-hydroxylase-like... 65 2e-11
>TC226020 similar to UP|Q9SP53 (Q9SP53) Flavanone 3-hydroxylase, partial
(93%)
Length = 1441
Score = 323 bits (829), Expect = 2e-89
Identities = 153/196 (78%), Positives = 176/196 (89%), Gaps = 1/196 (0%)
Frame = +3
Query: 1 MAPV-KTLKYLAQEKTLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCN 59
MAP KTL YLAQEKTLESSF+RD ERPKVAYN FS+EIP+ISLAGID+VDG R E C
Sbjct: 63 MAPTAKTLTYLAQEKTLESSFVRDEEERPKVAYNEFSDEIPVISLAGIDEVDGRRREICE 242
Query: 60 KIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSH 119
KIVEACENWGIFQ+VDHGVDQ+L++EMTRLAK FF LPP+EKLRFD+SG KKGGFIVSSH
Sbjct: 243 KIVEACENWGIFQVVDHGVDQQLVAEMTRLAKEFFALPPDEKLRFDMSGAKKGGFIVSSH 422
Query: 120 LQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEA 179
LQGE V+DWRE++ YFSYP ++RDYSRWP+ PEGW+ VTE+YS+K+M L+CKL+EVLSEA
Sbjct: 423 LQGESVQDWREIVTYFSYPKRERDYSRWPDTPEGWRSVTEEYSDKVMGLACKLMEVLSEA 602
Query: 180 MGLEKEALTKACVDMD 195
MGLEKE L+KACVDMD
Sbjct: 603 MGLEKEGLSKACVDMD 650
>TC232276 similar to UP|Q9FFF6 (Q9FFF6) Leucoanthocyanidin dioxygenase-like
protein (AT5g05600/MOP10_14), partial (38%)
Length = 463
Score = 100 bits (249), Expect = 4e-22
Identities = 55/153 (35%), Positives = 85/153 (54%), Gaps = 1/153 (0%)
Frame = +1
Query: 39 IPIISLAGIDDVD-GLRIETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLP 97
IPII LAG+ D R T KI EAC WG FQIV+HGV +L+ + FF +P
Sbjct: 4 IPIIDLAGLYGGDPDARASTLKKISEACNEWGFFQIVNHGVSPELMDMARETWRQFFHMP 183
Query: 98 PEEKLRFDLSGGKKGGFIVSSHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEV 157
E K ++ S G+ ++ + DW + P+ +D ++WP++P +EV
Sbjct: 184 LEVKQQYANSPKTYEGYGSRLGIEKGAILDWSDYYYLHYLPLSLKDNNKWPSQPPSCREV 363
Query: 158 TEQYSEKLMSLSCKLLEVLSEAMGLEKEALTKA 190
++Y +L+ L +L++VLS +GLE++AL KA
Sbjct: 364 CDEYGRELVKLCGRLMKVLSINLGLEEDALQKA 462
>TC228068 anthocyanidin synthase [Glycine max]
Length = 1231
Score = 99.8 bits (247), Expect = 7e-22
Identities = 60/191 (31%), Positives = 94/191 (48%), Gaps = 9/191 (4%)
Frame = +3
Query: 14 KTLESSFIRDVSERPKVAYNNFSNE------IPIISLAGIDDVDGLRIETCN-KIVEACE 66
K + ++R E + N F E +P I L ID D + C K+ +A E
Sbjct: 51 KCIPKEYVRPQEELKSIG-NVFEEEKKEGLQVPTIDLREIDSEDEVVRGKCREKLKKAAE 227
Query: 67 NWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRF--DLSGGKKGGFIVSSHLQGEP 124
WG+ +V+HG+ +LI + + + FF L EEK ++ DL GK G+
Sbjct: 228 EWGVMHLVNHGIPDELIERVKKAGETFFGLAVEEKEKYANDLESGKIQGYGSKLANNASG 407
Query: 125 VRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEK 184
+W + + ++P +RD S WP KP + EVT +Y+++L L+ K+LE LS +GLE
Sbjct: 408 QLEWEDYFFHLAFPEDKRDLSFWPKKPADYIEVTSEYAKRLRGLATKILEALSIGLGLEG 587
Query: 185 EALTKACVDMD 195
L K M+
Sbjct: 588 RRLEKEVGGME 620
>TC216796 similar to GB|AAQ65160.1|34365697|BT010537 At4g10500 {Arabidopsis
thaliana;} , partial (72%)
Length = 1454
Score = 99.0 bits (245), Expect = 1e-21
Identities = 56/179 (31%), Positives = 100/179 (55%), Gaps = 3/179 (1%)
Frame = +1
Query: 14 KTLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGL-RIETCNKIVEACENWGIFQ 72
K + S+FIR + +RP + S+++ I L + D+ G R +I +AC+N+G FQ
Sbjct: 133 KQVPSNFIRPLGDRPNLQGVVQSSDV-CIPLIDLQDLHGPNRSHIIQQIDQACQNYGFFQ 309
Query: 73 IVDHGVDQKLISEMTRLAKGFFDLPPEEKLR-FDLSGGKKGGFIVSSHLQGEPVRDWREM 131
+ +HGV + +I ++ ++ + FF LP EKL+ + K S ++ E V WR+
Sbjct: 310 VTNHGVPEGVIEKIMKVTREFFGLPESEKLKSYSTDPFKASRLSTSFNVNSEKVSSWRDF 489
Query: 132 MIYFSYPIKQRDY-SRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEALTK 189
+ +PI+ DY WP+ P +E +Y K+ +S KL+E +SE++GLE++ + +
Sbjct: 490 LRLHCHPIE--DYIKEWPSNPPSLREDVAEYCRKMRGVSLKLVEAISESLGLERDYINR 660
>TC222924 weakly similar to UP|Q9FLV0 (Q9FLV0) Flavanone 3-hydroxylase-like
protein, partial (44%)
Length = 590
Score = 97.8 bits (242), Expect = 3e-21
Identities = 57/176 (32%), Positives = 92/176 (51%), Gaps = 1/176 (0%)
Frame = +3
Query: 5 KTLKYLAQEKTLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVEA 64
K L Q L S+IR SERP+++ + ++PII L + R + ++I EA
Sbjct: 81 KVLSSGVQYSNLPESYIRPESERPRLSEVSECEDVPIIDLGCQN-----RAQIVHQIGEA 245
Query: 65 CENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLR-FDLSGGKKGGFIVSSHLQGE 123
C N+G FQ+++HGV + EM +A GFF LP EEKL+ + K S +++ E
Sbjct: 246 CRNYGFFQVINHGVALEAAKEMAEVAHGFFKLPVEEKLKLYSEDPSKTMRLSTSFNVKKE 425
Query: 124 PVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEA 179
V +WR+ + YP+ + WP+ P +KE +Y + L ++ E +SE+
Sbjct: 426 TVHNWRDYLRLHCYPL-DKYAPEWPSNPPSFKETVTEYCTLVRELGLRIQEYISES 590
>TC205381 weakly similar to UP|Q86B83 (Q86B83) CG33099-PA, partial (7%)
Length = 739
Score = 82.4 bits (202), Expect = 1e-16
Identities = 54/185 (29%), Positives = 99/185 (53%), Gaps = 17/185 (9%)
Frame = +3
Query: 16 LESSFIRDVSERPKVAYNNFSNEIPIISLAGIDD---VDGLRIETCNKIVE-ACENWGIF 71
++++FI++ RPK++ N + IPII L+ I + D IE K +E AC+ WG F
Sbjct: 27 VDTAFIQEPEHRPKLSPNQ-AEGIPIIDLSPITNHAVSDPSAIENLVKEIESACKEWGFF 203
Query: 72 QIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEPVRDWREM 131
Q+ +HGV L + + ++ FF+ EEK + G+ + H + VRDW+E+
Sbjct: 204 QVTNHGVPLTLRQNIEKASRLFFEQTQEEKKKVSRDEKSTTGYYDTEHTKN--VRDWKEV 377
Query: 132 MIYFS-----YPIKQRDY----SRW----PNKPEGWKEVTEQYSEKLMSLSCKLLEVLSE 178
+ + P+ ++ + W P P ++++ E+Y +++ LS KLLE+++
Sbjct: 378 FDFLAKDPTFIPVTSDEHDDRVTHWTNVSPEYPPNFRDIMEEYIQEVEKLSFKLLELIAL 557
Query: 179 AMGLE 183
++GLE
Sbjct: 558 SLGLE 572
>BU083789
Length = 449
Score = 82.0 bits (201), Expect = 2e-16
Identities = 44/151 (29%), Positives = 81/151 (53%)
Frame = +3
Query: 34 NFSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGF 93
+ S+E+P+I LA + + + E K+ AC+ WG FQIV+HGV + L +M + F
Sbjct: 3 HLSSEVPVIDLALLSNGNK---EELLKLDVACKEWGFFQIVNHGVQEHL-QKMKDASSEF 170
Query: 94 FDLPPEEKLRFDLSGGKKGGFIVSSHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEG 153
F LP + ++ + G+ + + E DW + ++ +YP + R WP PEG
Sbjct: 171 FKLPIDLINKYASASNDTHGYGQAYVVSEEQTLDWSDALMLITYPTRYRKLQFWPKTPEG 350
Query: 154 WKEVTEQYSEKLMSLSCKLLEVLSEAMGLEK 184
+ ++ + Y+ ++ + +L+ LS MG++K
Sbjct: 351 FMDIIDAYASEVRRVGEELISSLSVIMGMQK 443
>TC209361 weakly similar to UP|Q7XJ89 (Q7XJ89) Flavonol synthase (Fragment),
partial (19%)
Length = 784
Score = 82.0 bits (201), Expect = 2e-16
Identities = 54/184 (29%), Positives = 93/184 (50%), Gaps = 5/184 (2%)
Frame = +2
Query: 15 TLESSFIRDVSERPKVAYNNFSNEIPIISLA--GIDDVDGLRIETCNKIVEACENWGIFQ 72
+L FI ERP+++ + IPII L+ D + KI +ACE +G FQ
Sbjct: 80 SLSPKFILPEDERPQLSEVTSLDSIPIIDLSDHSYDGNNHSSSLVVQKISQACEEYGFFQ 259
Query: 73 IVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQ---GEPVRDWR 129
IV+HG+ +++ ++M F+LPPE+ + + K + + +L GE V+ W
Sbjct: 260 IVNHGIPEQVCNKMMTAITDIFNLPPEQTGQLYTTDHTKNTKLYNYYLNVEGGEKVKMWS 439
Query: 130 EMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEALTK 189
E ++ YPI+ + + E +Y+ ++ SL +LL +LS +G+E++ L K
Sbjct: 440 ECFSHYWYPIEDIIHLLPQEIGTQYGEAFSEYAREIGSLVRRLLGLLSIGLGIEEDFLLK 619
Query: 190 ACVD 193
C D
Sbjct: 620 ICGD 631
>TC216188 similar to UP|Q84L58 (Q84L58) 1-aminocyclopropane-1-carboxylic acid
oxidase, complete
Length = 1362
Score = 79.0 bits (193), Expect = 1e-15
Identities = 44/152 (28%), Positives = 78/152 (50%)
Frame = +3
Query: 39 IPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPP 98
+P+I + ++ + R +T +I CE WG FQ+++HG+ ++L+ + ++A F+ L
Sbjct: 147 VPVIDFSKLNGEE--RTKTMAQIANGCEEWGFFQLINHGIPEELLERVKKVASEFYKLER 320
Query: 99 EEKLRFDLSGGKKGGFIVSSHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVT 158
EE + S + + E V DW ++ I D + WP K G++E
Sbjct: 321 EENFKNSTSVKLLSDSVEKKSSEMEHV-DWEDV-------ITLLDDNEWPEKTPGFRETM 476
Query: 159 EQYSEKLMSLSCKLLEVLSEAMGLEKEALTKA 190
+Y +L L+ KL+EV+ E +GL K + KA
Sbjct: 477 AEYRAELKKLAEKLMEVMDENLGLTKGYIKKA 572
>TC205377
Length = 1345
Score = 79.0 bits (193), Expect = 1e-15
Identities = 53/189 (28%), Positives = 99/189 (52%), Gaps = 21/189 (11%)
Frame = +3
Query: 16 LESSFIRDVSERPKVAYNNFSNEIPIISLAGIDD--------VDGLRIETCNKIVEACEN 67
++++FI+D RPK++ + IPII L+ I + ++GL +I AC
Sbjct: 99 VDAAFIQDPPHRPKLSTIQ-AEGIPIIDLSPITNHRVSDPSAIEGL----VKEIGSACNE 263
Query: 68 WGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEPVRD 127
WG FQ+ +HGV L + + +K FF EEK + + G+ + H + VRD
Sbjct: 264 WGFFQVTNHGVPLTLRQNIEKASKLFFAQSAEEKRKVSRNESSPAGYYDTEHTKN--VRD 437
Query: 128 WREMMIYFS-----YPIKQRDY----SRWPNK----PEGWKEVTEQYSEKLMSLSCKLLE 174
W+E+ + + P+ ++ ++W N+ P ++ VT++Y +++ LS K+LE
Sbjct: 438 WKEVFDFLAKEPTFIPVTSDEHDDRVNQWTNQSPEYPLNFRVVTQEYIQEMEKLSFKILE 617
Query: 175 VLSEAMGLE 183
+++ ++GLE
Sbjct: 618 LIALSLGLE 644
>TC216699 weakly similar to UP|Q84MB3 (Q84MB3) At1g06620, partial (53%)
Length = 1436
Score = 77.4 bits (189), Expect = 4e-15
Identities = 45/149 (30%), Positives = 78/149 (52%), Gaps = 3/149 (2%)
Frame = +1
Query: 39 IPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPP 98
IPII L GI D LR K+ ACE WG FQ+++HG+ ++ EM + F
Sbjct: 439 IPIIDLTGIHDDPILRDHVVGKVRYACEKWGFFQVINHGIPTHVLDEMIKGTCRFHQQDA 618
Query: 99 E-EKLRFDLSGGKKGGFIVSSHLQGEPVRDWREMMIY--FSYPIKQRDYSRWPNKPEGWK 155
+ K + +K ++ + L +P DWR+ + + +P + ++ P +
Sbjct: 619 KVRKEYYTREVSRKVAYLSNYTLFEDPSADWRDTLAFSLAPHPPEAEEF------PAVCR 780
Query: 156 EVTEQYSEKLMSLSCKLLEVLSEAMGLEK 184
++ +YS+K+M+L+ L E+LSEA+GL +
Sbjct: 781 DIVNEYSKKIMALAYALFELLSEALGLNR 867
>TC209692 weakly similar to UP|Q39224 (Q39224) SRG1 protein (F6I1.30/F6I1.30)
(At1g17020/F6I1.30), partial (36%)
Length = 1229
Score = 75.5 bits (184), Expect = 1e-14
Identities = 44/165 (26%), Positives = 84/165 (50%), Gaps = 2/165 (1%)
Frame = +2
Query: 25 SERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKLIS 84
+E +A FS+ +P I+L + + + +E K+ AC +WG FQ+V+HG+ ++
Sbjct: 95 NEPSLLAGETFSHALPTINLKKLIHGEDIELEL-EKLTSACRDWGFFQLVEHGISSVVMK 271
Query: 85 EMTRLAKGFFDLPPEEKLRFDLSGGKKGGF--IVSSHLQGEPVRDWREMMIYFSYPIKQR 142
+ +GFF LP EEK+++ + G G+ ++ S + DW + + R
Sbjct: 272 TLEDEVEGFFMLPMEEKMKYKVRPGDVEGYGTVIGSE---DQKLDWGDRLFMKINXRSIR 442
Query: 143 DYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEAL 187
+ +P P + + E Y E+L +L+ L+ +L + + +EK L
Sbjct: 443 NPHLFPELPSSLRNILELYIEELQNLAMILMGLLGKTLKIEKREL 577
>TC214553 similar to UP|Q39224 (Q39224) SRG1 protein (F6I1.30/F6I1.30)
(At1g17020/F6I1.30), partial (52%)
Length = 1808
Score = 75.5 bits (184), Expect = 1e-14
Identities = 45/170 (26%), Positives = 85/170 (49%), Gaps = 1/170 (0%)
Frame = +1
Query: 25 SERPKVAYNNFS-NEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKLI 83
++ P + N S ++PII L + D +E K+ AC+ WG FQ+++HGVD ++
Sbjct: 121 NQDPHILSNTISLPQVPIIDLHQLLSEDPSELE---KLDHACKEWGFFQLINHGVDPPVV 291
Query: 84 SEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEPVRDWREMMIYFSYPIKQRD 143
M + FF+LP EEK +F + GF + E +W +M ++P+ R+
Sbjct: 292 ENMKIGVQEFFNLPMEEKQKFWQTPEDMQGFGQLFVVSEEQKLEWADMFYAHTFPLHSRN 471
Query: 144 YSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEALTKACVD 193
P P+ ++E E Y +L + ++ ++ +A+ ++ L++ D
Sbjct: 472 PHLIPKIPQPFRENLENYCLELRKMCITIIGLMKKALKIKTNELSELFED 621
>TC222700 similar to GB|AAS76251.1|45752706|BT012156 At1g55290 {Arabidopsis
thaliana;} , partial (8%)
Length = 677
Score = 70.1 bits (170), Expect = 6e-13
Identities = 52/192 (27%), Positives = 98/192 (50%), Gaps = 6/192 (3%)
Frame = +2
Query: 4 VKTLKYLAQEKTLESSFIRDVS-ERPKVAYNNFSNE-IPIISLAGIDDVD-GLRIETCNK 60
+ ++K L + K L S + E P+ + N+ + IP I + + + +R + +
Sbjct: 38 MSSVKELVESKCLSSVPSNYICLENPEDSILNYETDNIPTIDFSQLTSSNPSVRSKAIKQ 217
Query: 61 IVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRF---DLSGGKKGGFIVS 117
+ +AC +WG F +++HGV + L E+ R ++GFFDL +EK+ +L + G S
Sbjct: 218 LGDACRDWGFFMLINHGVSEILRDEVIRTSQGFFDLTEKEKMEHAGRNLFDPIRYG--TS 391
Query: 118 SHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLS 177
++ + WR+ + +P + P+KP G+ + E+Y K L +LLE +S
Sbjct: 392 FNVTVDKTLFWRDYLKCHVHP-----HFNAPSKPPGFSQTLEEYITKGRELVEELLEGIS 556
Query: 178 EAMGLEKEALTK 189
++GLE+ + K
Sbjct: 557 LSLGLEENFIHK 592
>AI507801 weakly similar to GP|20260150|gb| strong similarity to naringenin
3-dioxygenase {Arabidopsis thaliana}, partial (28%)
Length = 419
Score = 69.7 bits (169), Expect = 8e-13
Identities = 37/129 (28%), Positives = 61/129 (46%), Gaps = 13/129 (10%)
Frame = +2
Query: 54 RIETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFF-DLPPEEKLRFDLSGGKKG 112
R T + I AC +WG F + +HGV L++ + R FF D P +KLR+ S
Sbjct: 29 RASTRDSIARACRHWGAFHVTNHGVPPSLLASLRRAGLSFFSDTPMPDKLRYSCSAAAAS 208
Query: 113 GFIVSSHLQ------------GEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQ 160
S L V DWR+ + + P+ +R+ +RWP P ++E+ +
Sbjct: 209 EGYGSKMLATTTTTTTSDQNGAVQVLDWRDYFDHHTLPLSRRNPTRWPESPSDYRELVAR 388
Query: 161 YSEKLMSLS 169
YS+++ L+
Sbjct: 389 YSDEMNVLA 415
>TC215574 similar to UP|O04076 (O04076) ACC-oxidase, partial (98%)
Length = 1169
Score = 68.9 bits (167), Expect = 1e-12
Identities = 43/151 (28%), Positives = 83/151 (54%)
Frame = +3
Query: 40 PIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPE 99
P+I+L ++ + R T N+I +AC+NWG F++V+HG+ +L+ + RL K + E
Sbjct: 51 PVINLENLNGEE--RKATLNQIEDACQNWGFFELVNHGIPLELLDTVERLTKEHYRKCME 224
Query: 100 EKLRFDLSGGKKGGFIVSSHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTE 159
++ + +S S L+ E V+D +F + + S P+ + +++ +
Sbjct: 225 KRFKEAVS---------SKGLEAE-VKDMDWESTFFLRHLPTSNISEIPDLSQEYRDAMK 374
Query: 160 QYSEKLMSLSCKLLEVLSEAMGLEKEALTKA 190
++++KL L+ +LL++L E +GLEK L A
Sbjct: 375 EFAQKLEKLAEELLDLLCENLGLEKGYLKNA 467
>TC225031 homologue to UP|Q8W3Y3 (Q8W3Y3) 1-aminocyclopropane-1-carboxylic
acid oxidase, complete
Length = 1244
Score = 66.6 bits (161), Expect = 7e-12
Identities = 46/151 (30%), Positives = 79/151 (51%)
Frame = +1
Query: 40 PIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPE 99
P+I+L + + R +T KI +ACENWG F++V+HG+ ++ + RL K + E
Sbjct: 124 PLINLEKLSGEE--RNDTMEKIKDACENWGFFELVNHGIPHDILDTVERLTKEHYRKCME 297
Query: 100 EKLRFDLSGGKKGGFIVSSHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTE 159
E RF KG V + ++ DW P + + S P+ + +++V +
Sbjct: 298 E--RFKELVASKGLDAVQTEVKD---MDWESTFHLRHLP--ESNISEIPDLIDEYRKVMK 456
Query: 160 QYSEKLMSLSCKLLEVLSEAMGLEKEALTKA 190
++ +L L+ +LL++L E +GLEK L KA
Sbjct: 457 DFALRLEKLAEQLLDLLCENLGLEKGYLKKA 549
>TC205989 similar to UP|Q9FLV0 (Q9FLV0) Flavanone 3-hydroxylase-like protein,
partial (74%)
Length = 1192
Score = 66.2 bits (160), Expect = 9e-12
Identities = 35/102 (34%), Positives = 57/102 (55%), Gaps = 1/102 (0%)
Frame = +2
Query: 85 EMTRLAKGFFDLPPEEKLR-FDLSGGKKGGFIVSSHLQGEPVRDWREMMIYFSYPIKQRD 143
EM +A GFF LP EEKL+ + K S +++ E VR+WR+ + YP+ ++
Sbjct: 8 EMEEVAHGFFKLPVEEKLKLYSEDTSKTMRLSTSFNVKKETVRNWRDYLRLHCYPL-EKY 184
Query: 144 YSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKE 185
WP+ P +KE +Y + L ++ E +SE++GLEK+
Sbjct: 185 APEWPSNPPSFKETVTEYCTIIRELGLRIQEYISESLGLEKD 310
>TC208834 similar to UP|Q39224 (Q39224) SRG1 protein (F6I1.30/F6I1.30)
(At1g17020/F6I1.30), partial (54%)
Length = 1070
Score = 65.5 bits (158), Expect = 2e-11
Identities = 33/135 (24%), Positives = 70/135 (51%)
Frame = +2
Query: 59 NKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSS 118
+K+ AC+ WG FQ+V+HGV+ L+ ++ + FF+LP EK +F + GF +
Sbjct: 29 DKLHLACKEWGFFQLVNHGVNSSLVEKVRLETQDFFNLPMSEKKKFWQTPQHMEGFGQAF 208
Query: 119 HLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSE 178
+ + DW ++ + P R +P P +++ E YS ++ L+ ++ ++ +
Sbjct: 209 VVSEDQKLDWADLYYMTTLPKHSRMPHLFPQLPLPFRDTLEAYSREIKDLAIVIIGLMGK 388
Query: 179 AMGLEKEALTKACVD 193
A+ +++ + + D
Sbjct: 389 ALKIQEREIRELFED 433
>BF067521 similar to GP|10177853|dbj flavanone 3-hydroxylase-like protein
{Arabidopsis thaliana}, partial (14%)
Length = 412
Score = 65.1 bits (157), Expect = 2e-11
Identities = 35/96 (36%), Positives = 53/96 (54%)
Frame = +3
Query: 4 VKTLKYLAQEKTLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVE 63
+K L Q L S+IR SERP+++ + ++PII L + R + ++I E
Sbjct: 138 IKVLSSGVQYSNLPESYIRPESERPRLSEVSECEDVPIIDLGSQN-----RAQIVHQIGE 302
Query: 64 ACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPE 99
AC N+G F +++HGV + EM +A GFF LP E
Sbjct: 303 ACRNYGFFHVINHGVALEAAEEMEEVAHGFFKLPVE 410
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,520,069
Number of Sequences: 63676
Number of extensions: 113447
Number of successful extensions: 591
Number of sequences better than 10.0: 138
Number of HSP's better than 10.0 without gapping: 567
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 567
length of query: 195
length of database: 12,639,632
effective HSP length: 92
effective length of query: 103
effective length of database: 6,781,440
effective search space: 698488320
effective search space used: 698488320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC138453.15


