

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138130.19 - phase: 0
(881 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
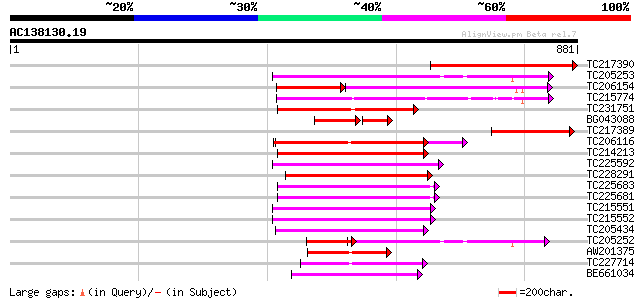
Score E
Sequences producing significant alignments: (bits) Value
TC217390 similar to UP|Q9LU17 (Q9LU17) Cell division protein Fts... 336 2e-92
TC205253 homologue to UP|O99018 (O99018) Chloroplast protease pr... 299 4e-81
TC206154 homologue to UP|FTSH_MEDSA (Q9BAE0) Cell division prote... 171 2e-77
TC215774 similar to UP|Q94ES0 (Q94ES0) AAA-metalloprotease FtsH,... 265 9e-71
TC231751 homologue to GB|BAB08420.1|9757998|AB025622 cell divisi... 229 5e-60
BG043088 homologue to GP|2062173|gb| cell division protein FtsH ... 134 6e-52
TC217389 similar to UP|Q9LU17 (Q9LU17) Cell division protein Fts... 191 1e-48
TC206116 UP|CC48_SOYBN (P54774) Cell division cycle protein 48 h... 189 3e-48
TC214213 homologue to UP|PRSA_BRACM (O23894) 26S protease regula... 189 5e-48
TC225592 homologue to GB|AAP78936.1|32306505|BT009668 At5g19990 ... 181 1e-45
TC228291 homologue to UP|Q9FXT8 (Q9FXT8) 26S proteasome regulato... 181 2e-45
TC225683 homologue to UP|Q9SZD4 (Q9SZD4) 26S proteasome subunit ... 178 8e-45
TC225681 homologue to UP|Q9SZD4 (Q9SZD4) 26S proteasome subunit ... 178 8e-45
TC215551 homologue to UP|PRS7_SPIOL (Q41365) 26S protease regula... 170 3e-42
TC215552 UP|PRS7_ARATH (Q9SSB5) 26S protease regulatory subunit ... 170 3e-42
TC205434 homologue to UP|PRS6_ARATH (Q9SEI4) 26S protease regula... 168 1e-41
TC205252 homologue to UP|Q9ZP50 (Q9ZP50) FtsH-like protein Pftf ... 164 2e-40
AW201375 homologue to GP|17065470|gb cell division protein FtsH-... 156 4e-38
TC227714 similar to UP|C48C_ARATH (Q9SS94) Cell division control... 153 4e-37
BE661034 similar to SP|Q9ZPR1|C48B Cell division control protein... 152 6e-37
>TC217390 similar to UP|Q9LU17 (Q9LU17) Cell division protein FtsH-like,
partial (21%)
Length = 886
Score = 336 bits (862), Expect = 2e-92
Identities = 164/227 (72%), Positives = 194/227 (85%)
Frame = +1
Query: 655 AQMEERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGYVRT 714
AQMEERGMLDRKERS E W+QVAINEAAMAV A+N P+ NIE++TIAPRAGRELGYVR
Sbjct: 1 AQMEERGMLDRKERSTETWKQVAINEAAMAVVAVNFPDLKNIEFVTIAPRAGRELGYVRV 180
Query: 715 MLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNARVAARMYMI 774
++S+ FN GMLTRQSL DHITVQLAPRAADE+WFG QLSTIWAETADNAR AAR +++
Sbjct: 181 KMDSVKFNQGMLTRQSLLDHITVQLAPRAADELWFGSGQLSTIWAETADNARSAARTFVL 360
Query: 775 GGLSDKYRGVSNFWVTDRINEIDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKT 834
GGLS+KY G+SNFWV+DRINEID EAM+I+N CYERAKEIL+QN+TLMD LVNELV KK+
Sbjct: 361 GGLSEKYHGMSNFWVSDRINEIDSEAMQIVNSCYERAKEILEQNRTLMDALVNELVEKKS 540
Query: 835 LTKEDIVRLVQLHGHAKPIPISVLDIRDAKHKELQEIASNGKEINES 881
LTK++ LV+LHG KP+P S+LDIR AK +E Q++ +GKE + S
Sbjct: 541 LTKQEFFHLVELHGSLKPMPPSILDIRVAKCREFQKLIGSGKETSLS 681
>TC205253 homologue to UP|O99018 (O99018) Chloroplast protease precursor,
partial (69%)
Length = 1678
Score = 299 bits (765), Expect = 4e-81
Identities = 186/459 (40%), Positives = 269/459 (58%), Gaps = 23/459 (5%)
Frame = +2
Query: 409 LERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLA 468
+E V F DVAG+ + + + E+V+F E + G +IP G+LL GPPG GKTLLA
Sbjct: 32 MEPNTGVTFDDVAGVDEAKQDFMEVVEFLKKPERFTAVGARIPKGVLLVGPPGTGKTLLA 211
Query: 469 KAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGL 528
KA+AGEAGV FFSIS S+FVE++VGVGASRVR L+++AKENAP +VF+DE+DAVGR+RG
Sbjct: 212 KAIAGEAGVPFFSISGSEFVEMFVGVGASRVRDLFRKAKENAPCIVFVDEIDAVGRQRGT 391
Query: 529 IKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKP 588
G G ER+ TLNQLL +DGFEG +I IA+TNR DILD AL+RPGRFDR++ + P
Sbjct: 392 GIGGGNDEREQTLNQLLTEMDGFEGNTGIIVIAATNRVDILDSALLRPGRFDRQVTVDVP 571
Query: 589 GFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVST 648
GR EILKVH K DV E++A T G GA+LAN++ AAI R +T +S+
Sbjct: 572 DIRGRTEILKVHGSNKKFEADVSLEVIAMRTPGFSGADLANLLNEAAILAGRRGKTAISS 751
Query: 649 ---DDLLQ--AAQMEERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAP 703
DD + A ME M D K +S VA +E A+ P D ++ +T+ P
Sbjct: 752 KEIDDSIDRIVAGMEGTVMTDGKSKS-----LVAYHEVGHAICGTLTPGHDPVQKVTLVP 916
Query: 704 RAGRELGYVRTMLESINFND-GMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWA-ET 761
R G R + I +D ++++Q LF I L RAA+E+ FG+ +++T A +
Sbjct: 917 R-----GQARGLTWFIPADDPTLISKQQLFARIVGGLGGRAAEEVIFGEPEVTTGAAGDL 1081
Query: 762 ADNARVAARMYMIGGLSD---------------KYRGVSNFWVTDRINE-IDLEAMKILN 805
+A +M G+SD R ++ +++++ E ID ++ +
Sbjct: 1082QQITSLAKQMVTTFGMSDIGPWSLMDSSAQSDVIMRMMARNSMSEKLAEDIDAAVKRLSD 1261
Query: 806 LCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLV 844
YE A ++ N+ +D +V L+ K+T++ ++ L+
Sbjct: 1262EAYEIALSQIRSNREAIDKIVEVLLEKETMSGDEFRALL 1378
>TC206154 homologue to UP|FTSH_MEDSA (Q9BAE0) Cell division protein ftsH
homolog, chloroplast precursor , partial (88%)
Length = 2059
Score = 171 bits (433), Expect(2) = 2e-77
Identities = 113/341 (33%), Positives = 182/341 (53%), Gaps = 20/341 (5%)
Frame = +1
Query: 523 GRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRK 582
G + G G G ER+ T+NQLL +DGF G VI +A+TNRPD+LD AL+RPGRFDR+
Sbjct: 838 GSREGRGFGGGNDEREQTINQLLTEMDGFSGNSGVIVLAATNRPDVLDSALLRPGRFDRQ 1017
Query: 583 IFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDS 642
+ + +P GR++IL+VH+R K +A+DVD+E +A T G GA+L N++ AAI R
Sbjct: 1018 VTVDRPDVAGRVKILQVHSRGKALAKDVDFEKIARRTPGFTGADLQNLMNEAAILAARRD 1197
Query: 643 RTEVSTDDLLQAAQMEERGMLDRKER--SKEKWEQVAINEAAMAVAAMNLPNFDNIEYIT 700
E+S D++ A + G ++K S EK + VA +EA A+ +P +D + I+
Sbjct: 1198 LKEISKDEISDALERIIAGP-EKKNAVVSDEKKKLVAYHEAGHALVGALMPEYDPVAKIS 1374
Query: 701 IAPRAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLST-IWA 759
I PR G+ G G+ +R L + + V L R A+E+ FG++ ++T
Sbjct: 1375 IIPR-GQAGGLTFFAPSEERLESGLYSRSYLENQMAVALGGRVAEEVIFGQENVTTGASN 1551
Query: 760 ETADNARVAARMYMIGGLSDKYRGVS------NFWVTDRINE-----------IDLEAMK 802
+ +RVA +M G S K V+ N ++ +++ +D E +
Sbjct: 1552 DFMQVSRVARQMVERFGFSKKIGQVAIGGPGGNPFLGQQMSSQKDYSMATADVVDAEVRE 1731
Query: 803 ILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRL 843
++ Y RA I+ + ++ L L+ K+T+ E+ + L
Sbjct: 1732 LVERAYSRATHIISTHIDILHKLAQLLIEKETVDGEEFMSL 1854
Score = 137 bits (346), Expect(2) = 2e-77
Identities = 68/108 (62%), Positives = 84/108 (76%)
Frame = +2
Query: 415 VKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGE 474
V F DVAG + +LEL+E+V F + + Y G KIP G LL GPPG GKTLLA+AVAGE
Sbjct: 512 VSFADVAGADQAKLELQEVVDFLKNPDKYTALGAKIPKGCLLVGPPGTGKTLLARAVAGE 691
Query: 475 AGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAV 522
AGV FFS +AS+FVE++VGVGASRVR L+++AK AP +VFIDE+DAV
Sbjct: 692 AGVPFFSCAASEFVELFVGVGASRVRDLFEKAKGKAPCIVFIDEIDAV 835
>TC215774 similar to UP|Q94ES0 (Q94ES0) AAA-metalloprotease FtsH, partial
(79%)
Length = 2084
Score = 265 bits (676), Expect = 9e-71
Identities = 172/450 (38%), Positives = 264/450 (58%), Gaps = 20/450 (4%)
Frame = +3
Query: 415 VKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGE 474
+ F DVAG + + E+ E V F + + Y G KIP G LL GPPG GKTLLAKA AGE
Sbjct: 489 IYFKDVAGCDEAKQEIMEFVHFLKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGE 668
Query: 475 AGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGR-KRGLIKGSG 533
+GV F SIS S F+E++VGVG SRVR+L+QEA++ +PS+VFIDE+DA+GR +RG G+
Sbjct: 669 SGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCSPSIVFIDEIDAIGRARRGSFSGA- 845
Query: 534 GQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGR 593
ER++TLNQLLV +DGF V+ +A TNRP+ILD AL+RPGRFDR+I I KP GR
Sbjct: 846 NDERESTLNQLLVEMDGFGTTSGVVVLAGTNRPEILDKALLRPGRFDRQITIDKPDIKGR 1025
Query: 594 IEILKVHARKKPIAEDVDY--EIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDL 651
+I +++ +K + + Y +A++T G GA++AN+ AA+ R T+V T +
Sbjct: 1026DQIFQIYLKKIKLDHEPSYYSPRLAALTPGFAGADIANVCNEAALIAARGEGTQV-TMEH 1202
Query: 652 LQAAQMEERGMLDRKERSKEKWEQ--VAINEAAMAVAAMNLPNFDNIEYITIAPRAGREL 709
+AA G L+++ + K E+ VA +EA AV+ L + + + +TI PR L
Sbjct: 1203FEAAIDRIIGGLEKRNKVISKLERRTVAYHEAGHAVSGWFLEHVEPLLKVTIVPRGTAAL 1382
Query: 710 GYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNARVAA 769
G+ + + + ++T++ LFD + L RAA+++ G+ +ST D +V
Sbjct: 1383GFA----QYVPNENLLMTKEQLFDMTCMTLGGRAAEQVLIGR--IST--GAQNDLEKVTK 1538
Query: 770 RMY---MIGGLSDKYRGVSNFWVTDRINE------------IDLEAMKILNLCYERAKEI 814
Y + G SDK G+ +F T+ E ID E +N YE ++
Sbjct: 1539MTYAQVAVYGFSDKV-GLLSFPPTEGSYEISKPYSSKTAAIIDNEVRDWVNKAYEHTVQL 1715
Query: 815 LQQNKTLMDTLVNELVVKKTLTKEDIVRLV 844
++++K + + L+ K+ L ++D++R++
Sbjct: 1716IKEHKEQVAQIAELLLEKEVLHQDDLLRVL 1805
>TC231751 homologue to GB|BAB08420.1|9757998|AB025622 cell division protein
FtsH protease-like {Arabidopsis thaliana;} , partial
(30%)
Length = 733
Score = 229 bits (583), Expect = 5e-60
Identities = 120/219 (54%), Positives = 154/219 (69%)
Frame = +1
Query: 417 FTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAG 476
F DV G + ELEE+V++ + + R G K+P GILL GPPG GKTLLAKA+AGEAG
Sbjct: 88 FKDVKGCDDAKQELEEVVEYLKNPAKFTRLGGKLPKGILLTGPPGTGKTLLAKAIAGEAG 267
Query: 477 VNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQE 536
V FF + S+F E+YVGVGA RVRSL+Q AK+ AP ++FIDE+DAVG R +G
Sbjct: 268 VPFFYRAGSEFEEMYVGVGARRVRSLFQAAKKKAPCIIFIDEIDAVGSTRKQWEG----H 435
Query: 537 RDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIEI 596
TL+QLLV +DGFE +I IA+TN PDILDPAL RPGRFDR I +P P GR EI
Sbjct: 436 TKKTLHQLLVEMDGFEQNEGIIVIAATNLPDILDPALTRPGRFDRHIVVPNPDLRGRQEI 615
Query: 597 LKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAA 635
L+++ + KP+A+D+D + +A T G GA+LAN+V +AA
Sbjct: 616 LELYLQDKPLADDIDIKSIARGTPGFNGADLANLVNIAA 732
>BG043088 homologue to GP|2062173|gb| cell division protein FtsH isolog
{Arabidopsis thaliana}, partial (11%)
Length = 349
Score = 134 bits (336), Expect(2) = 6e-52
Identities = 67/71 (94%), Positives = 71/71 (99%)
Frame = +2
Query: 474 EAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSG 533
EAGVNFFSISASQFVEIYVGVGASRVR+LYQEA+ENAPSVVFIDELDAVGR+RGLIKGSG
Sbjct: 2 EAGVNFFSISASQFVEIYVGVGASRVRALYQEARENAPSVVFIDELDAVGRERGLIKGSG 181
Query: 534 GQERDATLNQL 544
GQERDATLNQ+
Sbjct: 182 GQERDATLNQM 214
Score = 90.1 bits (222), Expect(2) = 6e-52
Identities = 42/46 (91%), Positives = 44/46 (95%)
Frame = +1
Query: 549 DGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRI 594
DGFEGRGEVITIASTNRP ILDPALVRPGRFDRKI++PKPG IGRI
Sbjct: 211 DGFEGRGEVITIASTNRPXILDPALVRPGRFDRKIYMPKPGLIGRI 348
>TC217389 similar to UP|Q9LU17 (Q9LU17) Cell division protein FtsH-like,
partial (11%)
Length = 755
Score = 191 bits (485), Expect = 1e-48
Identities = 91/129 (70%), Positives = 112/129 (86%)
Frame = +3
Query: 749 FGKDQLSTIWAETADNARVAARMYMIGGLSDKYRGVSNFWVTDRINEIDLEAMKILNLCY 808
FG QLSTIWAETADNAR AAR +++GGLS+KY G+SNFWV+DRINEID EAM+I+N CY
Sbjct: 3 FGSGQLSTIWAETADNARSAARTFVLGGLSEKYHGMSNFWVSDRINEIDSEAMRIVNSCY 182
Query: 809 ERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLVQLHGHAKPIPISVLDIRDAKHKEL 868
ERAKEIL+QN+TLMD LVNELV KK+LTK++ VRLV+LHG KP+P+S+LDIR AK +E
Sbjct: 183 ERAKEILEQNRTLMDALVNELVEKKSLTKQEFVRLVELHGFLKPMPLSILDIRVAKCREF 362
Query: 869 QEIASNGKE 877
Q++ +GKE
Sbjct: 363 QKLIDSGKE 389
>TC206116 UP|CC48_SOYBN (P54774) Cell division cycle protein 48 homolog
(Valosin containing protein homolog) (VCP), complete
Length = 2583
Score = 189 bits (481), Expect = 3e-48
Identities = 99/242 (40%), Positives = 153/242 (62%), Gaps = 1/242 (0%)
Frame = +1
Query: 410 ERGVDVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLLA 468
ER +V + DV G+ K ++ E+V+ H ++++ GVK P GILL GPPG GKTL+A
Sbjct: 619 ERLDEVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPPKGILLYGPPGSGKTLIA 798
Query: 469 KAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGL 528
+AVA E G FF I+ + + G S +R ++EA++NAPS++FIDE+D++ KR
Sbjct: 799 RAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAPSIIFIDEIDSIAPKR-- 972
Query: 529 IKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKP 588
+ + G+ ++QLL +DG + R VI I +TNRP+ +DPAL R GRFDR+I I P
Sbjct: 973 -EKTHGEVERRIVSQLLTLMDGLKSRAHVIVIGATNRPNSIDPALRRFGRFDREIDIGVP 1149
Query: 589 GFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVST 648
+GR+E+L++H + +++DVD E +A T G VGA+LA + AA+ +R+ +
Sbjct: 1150DEVGRLEVLRIHTKNMKLSDDVDLERIAKDTHGYVGADLAALCTEAALQCIREKMDVIDL 1329
Query: 649 DD 650
+D
Sbjct: 1330ED 1335
Score = 183 bits (464), Expect = 3e-46
Identities = 106/300 (35%), Positives = 163/300 (54%), Gaps = 2/300 (0%)
Frame = +1
Query: 414 DVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVA 472
+V + D+ GL ++ EL+E V++ H E + + G+ G+L GPPG GKTLLAKA+A
Sbjct: 1450 NVSWEDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPGCGKTLLAKAIA 1629
Query: 473 GEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGS 532
E NF S+ + + ++ G + VR ++ +A+++AP V+F DELD++ +RG G
Sbjct: 1630 NECQANFISVKGPELLTMWFGESEANVREIFDKARQSAPCVLFFDELDSIATQRGSSVGD 1809
Query: 533 GGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIG 592
G D LNQLL +DG + V I +TNRPDI+DPAL+RPGR D+ I+IP P
Sbjct: 1810 AGGAADRVLNQLLTEMDGMSAKKTVFIIGATNRPDIIDPALLRPGRLDQLIYIPLPDEDS 1989
Query: 593 RIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLL 652
R +I K RK PIA++VD +A T G GA++ I + A +R++ + D
Sbjct: 1990 RHQIFKACLRKSPIAKNVDLRALARHTQGFSGADITEICQRACKYAIREN---IEKDIER 2160
Query: 653 QAAQMEERGMLDRKERSKEKWE-QVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGY 711
+ E +D E E + A E +M A ++ + D +Y A + G+
Sbjct: 2161 ERKSRENPEAMDEDTVDDEVAEIKAAHFEESMKFARRSVSDADIRKYQAFAQTLQQSRGF 2340
>TC214213 homologue to UP|PRSA_BRACM (O23894) 26S protease regulatory subunit
6A homolog (TAT-binding protein homolog 1) (TBP-1),
partial (64%)
Length = 1086
Score = 189 bits (480), Expect = 5e-48
Identities = 103/235 (43%), Positives = 145/235 (60%), Gaps = 1/235 (0%)
Frame = +1
Query: 417 FTDVAGLGKIRLEL-EEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEA 475
+ D+ GL K EL E IV TH E +++ GV+ P G+LL GPPG GKTL+A+A A +
Sbjct: 46 YNDIGGLEKQIQELVEAIVLPMTHKERFQKLGVRPPKGVLLYGPPGTGKTLMARACAAQT 225
Query: 476 GVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQ 535
F ++ Q V++++G GA VR +Q AKE +P ++FIDE+DA+G KR + SG +
Sbjct: 226 NATFLKLAGPQLVQMFIGDGAKLVRDAFQLAKEKSPCIIFIDEIDAIGTKRFDSEVSGDR 405
Query: 536 ERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIE 595
E T+ +LL LDGF + IA+TNR DILDPAL+R GR DRKI P P R
Sbjct: 406 EVQRTMLELLNQLDGFSSDDRIKVIAATNRADILDPALMRSGRLDRKIEFPHPSEEARAR 585
Query: 596 ILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDD 650
IL++H+RK + DV++E +A TD GA+L + A + +R TEV+ +D
Sbjct: 586 ILQIHSRKMNVHPDVNFEELARSTDDFNGAQLKAVCVEAGMLALRRDATEVNHED 750
>TC225592 homologue to GB|AAP78936.1|32306505|BT009668 At5g19990 {Arabidopsis
thaliana;} , partial (94%)
Length = 1612
Score = 181 bits (460), Expect = 1e-45
Identities = 97/268 (36%), Positives = 156/268 (58%), Gaps = 2/268 (0%)
Frame = +3
Query: 409 LERGVDVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLL 467
+E+ D + + GL + E++E+++ H E++ G+ P G+LL GPPG GKTLL
Sbjct: 552 VEKVPDSTYDMIGGLDQQIKEIKEVIELPIKHPELFESLGIAQPKGVLLYGPPGTGKTLL 731
Query: 468 AKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRG 527
A+AVA F +S S+ V+ Y+G G+ VR L+ A+E+APS++F+DE+D++G R
Sbjct: 732 ARAVAHHTDCTFIRVSGSELVQKYIGEGSRMVRELFVMAREHAPSIIFMDEIDSIGSARM 911
Query: 528 LI-KGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIP 586
G+G E T+ +LL LDGFE ++ + +TNR DILD AL+RPGR DRKI P
Sbjct: 912 ESGSGNGDSEVQRTMLELLNQLDGFEASNKIKVLMATNRIDILDQALLRPGRIDRKIEFP 1091
Query: 587 KPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEV 646
P R++ILK+H+R+ + +D + +A +G GAEL + A + +R+ R V
Sbjct: 1092NPNEESRLDILKIHSRRMNLMRGIDLKKIAEKMNGASGAELKAVCTEAGMFALRERRVHV 1271
Query: 647 STDDLLQAAQMEERGMLDRKERSKEKWE 674
+ +D A + ++ ++ W+
Sbjct: 1272TQEDFEMAVAKVMKKETEKNMSLRKLWK 1355
>TC228291 homologue to UP|Q9FXT8 (Q9FXT8) 26S proteasome regulatory particle
triple-A ATPase subunit4, partial (62%)
Length = 1079
Score = 181 bits (458), Expect = 2e-45
Identities = 95/229 (41%), Positives = 142/229 (61%), Gaps = 1/229 (0%)
Frame = +3
Query: 429 ELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQF 487
EL E ++ + E++ R G+K P G+LL GPPG GKTLLA+A+A NF + +S
Sbjct: 9 ELRESIELPLMNPELFIRVGIKPPKGVLLYGPPGTGKTLLARAIASNIEANFLKVVSSAI 188
Query: 488 VEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVC 547
++ Y+G A +R ++ A+++ P ++F+DE+DA+G +R S +E TL +LL
Sbjct: 189 IDKYIGESARLIREMFGYARDHQPCIIFMDEIDAIGGRRFSEGTSADREIQRTLMELLNQ 368
Query: 548 LDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIA 607
LDGF+ G+V I +TNRPD+LDPAL+RPGR DRKI IP P R+EILK+HA
Sbjct: 369 LDGFDQLGKVKMIMATNRPDVLDPALLRPGRLDRKIEIPLPNEQSRMEILKIHAAGIAKH 548
Query: 608 EDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQAAQ 656
++DYE V + +G GA+L N+ A + +R R V +D ++A +
Sbjct: 549 GEIDYEAVVKLAEGFNGADLRNVCTEAGMAAIRAERDYVIHEDFMKAVR 695
>TC225683 homologue to UP|Q9SZD4 (Q9SZD4) 26S proteasome subunit 4-like
protein (26S proteasome subunit AtRPT2a), partial (97%)
Length = 1513
Score = 178 bits (452), Expect = 8e-45
Identities = 101/252 (40%), Positives = 147/252 (58%), Gaps = 1/252 (0%)
Frame = +2
Query: 417 FTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEA 475
+ D+ GL E++E V+ TH E+Y G+K P G++L G PG GKTLLAKAVA
Sbjct: 569 YADIGGLDAQIQEIKEAVELPLTHPELYEDIGIKPPKGVILYGEPGTGKTLLAKAVANST 748
Query: 476 GVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQ 535
F + S+ ++ Y+G G VR L++ A + +PS+VFIDE+DAVG KR G +
Sbjct: 749 SATFLRVVGSELIQKYLGDGPKLVRELFRVADDLSPSIVFIDEIDAVGTKRYDAHSGGER 928
Query: 536 ERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIE 595
E T+ +LL LDGF+ RG+V I +TNR + LDPAL+RPGR DRKI P P R
Sbjct: 929 EIQRTMLELLNQLDGFDSRGDVKVILATNRIESLDPALLRPGRIDRKIEFPLPDIKTRRR 1108
Query: 596 ILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQAA 655
I ++H + +A+DV+ E D GA++ I A + +R+ R +V+ D +A
Sbjct: 1109IFQIHTSRMTLADDVNLEEFVMTKDEFSGADIKAICTEAGLLALRERRMKVTHADFKKA- 1285
Query: 656 QMEERGMLDRKE 667
+++ M +KE
Sbjct: 1286--KDKVMFKKKE 1315
>TC225681 homologue to UP|Q9SZD4 (Q9SZD4) 26S proteasome subunit 4-like
protein (26S proteasome subunit AtRPT2a), partial (93%)
Length = 1433
Score = 178 bits (452), Expect = 8e-45
Identities = 101/252 (40%), Positives = 147/252 (58%), Gaps = 1/252 (0%)
Frame = +1
Query: 417 FTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEA 475
+ D+ GL E++E V+ TH E+Y G+K P G++L G PG GKTLLAKAVA
Sbjct: 466 YADIGGLDAQIQEIKEAVELPLTHPELYEDIGIKPPKGVILYGEPGTGKTLLAKAVANST 645
Query: 476 GVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQ 535
F + S+ ++ Y+G G VR L++ A + +PS+VFIDE+DAVG KR G +
Sbjct: 646 SATFLRVVGSELIQKYLGDGPKLVRELFRVADDLSPSIVFIDEIDAVGTKRYDAHSGGER 825
Query: 536 ERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIE 595
E T+ +LL LDGF+ RG+V I +TNR + LDPAL+RPGR DRKI P P R
Sbjct: 826 EIQRTMLELLNQLDGFDSRGDVKVILATNRIESLDPALLRPGRIDRKIEFPLPDIKTRRR 1005
Query: 596 ILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQAA 655
I ++H + +A+DV+ E D GA++ I A + +R+ R +V+ D +A
Sbjct: 1006IFQIHTSRMTLADDVNLEEFVMTKDEFSGADIKAICTEAGLLALRERRMKVTHADFKKA- 1182
Query: 656 QMEERGMLDRKE 667
+++ M +KE
Sbjct: 1183--KDKVMFKKKE 1212
>TC215551 homologue to UP|PRS7_SPIOL (Q41365) 26S protease regulatory subunit
7 (26S proteasome subunit 7) (26S proteasome AAA-ATPase
subunit RPT1) (Regulatory particle triple-A ATPase
subunit 1), complete
Length = 1561
Score = 170 bits (430), Expect = 3e-42
Identities = 93/254 (36%), Positives = 138/254 (53%), Gaps = 1/254 (0%)
Frame = +3
Query: 409 LERGVDVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLL 467
+E DV + DV G + ++ E+V+ H E + + G+ P G+L GPPG GKTLL
Sbjct: 531 VEEKPDVTYNDVGGCKEQIEKMREVVELPMLHPEKFVKLGIDPPKGVLCYGPPGTGKTLL 710
Query: 468 AKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRG 527
A+AVA F + S+ V+ YVG GA VR L+Q A+ +VF DE+DA+G R
Sbjct: 711 ARAVANRTDACFIRVIGSELVQKYVGEGARMVRELFQMARSKKACIVFFDEVDAIGGARF 890
Query: 528 LIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPK 587
G E T+ +++ LDGF+ RG + + +TNRPD LDPAL+RPGR DRK+
Sbjct: 891 DDGVGGDNEVQRTMLEIVNQLDGFDARGNIKVLMATNRPDTLDPALLRPGRLDRKVEFGL 1070
Query: 588 PGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVS 647
P R +I K+H R D+ +E++A + GA++ ++ A + +R R V+
Sbjct: 1071PDLESRTQIFKIHTRTMNCERDIRFELLARLCPNSTGADIRSVCTEAGMYAIRARRKTVT 1250
Query: 648 TDDLLQAAQMEERG 661
D L A +G
Sbjct: 1251EKDFLDAVNKVIKG 1292
>TC215552 UP|PRS7_ARATH (Q9SSB5) 26S protease regulatory subunit 7 (26S
proteasome subunit 7) (26S proteasome AAA-ATPase subunit
RPT1a) (Regulatory particle triple-A ATPase subunit 1a),
partial (73%)
Length = 1082
Score = 170 bits (430), Expect = 3e-42
Identities = 93/254 (36%), Positives = 138/254 (53%), Gaps = 1/254 (0%)
Frame = +1
Query: 409 LERGVDVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLL 467
+E DV + DV G + ++ E+V+ H E + + G+ P G+L GPPG GKTLL
Sbjct: 124 VEEKPDVTYNDVGGCKEQIEKMREVVELPMLHPEKFVKLGIDPPKGVLCYGPPGTGKTLL 303
Query: 468 AKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRG 527
A+AVA F + S+ V+ YVG GA VR L+Q A+ +VF DE+DA+G R
Sbjct: 304 ARAVANRTDACFIRVIGSELVQKYVGEGARMVRELFQMARSKKACIVFFDEVDAIGGARF 483
Query: 528 LIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPK 587
G E T+ +++ LDGF+ RG + + +TNRPD LDPAL+RPGR DRK+
Sbjct: 484 DDGVGGDNEVQRTMLEIVNQLDGFDARGNIKVLMATNRPDTLDPALLRPGRLDRKVEFGL 663
Query: 588 PGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVS 647
P R +I K+H R D+ +E++A + GA++ ++ A + +R R V+
Sbjct: 664 PDLESRTQIFKIHTRTMNCERDIRFELLARLCPNSTGADIRSVCTEAGMYAIRARRKTVT 843
Query: 648 TDDLLQAAQMEERG 661
D L A +G
Sbjct: 844 EKDFLDAVNKVIKG 885
>TC205434 homologue to UP|PRS6_ARATH (Q9SEI4) 26S protease regulatory subunit
6B homolog (26S proteasome AAA-ATPase subunit RPT3)
(Regulatory particle triple-A ATPase subunit 3), partial
(96%)
Length = 1573
Score = 168 bits (425), Expect = 1e-41
Identities = 94/238 (39%), Positives = 139/238 (57%), Gaps = 1/238 (0%)
Frame = +1
Query: 414 DVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVA 472
DV + D+ G + E+ E V+ TH E+Y++ G+ P G+LL GPPG GKT+LAKAVA
Sbjct: 565 DVTYNDIGGCDIQKQEIREAVELPLTHHELYKQIGIDPPRGVLLYGPPGTGKTMLAKAVA 744
Query: 473 GEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGS 532
F + S+FV+ Y+G G VR +++ AKENAP+++FIDE+DA+ R +
Sbjct: 745 NHTTAAFIRVVGSEFVQKYLGEGPRMVRDVFRLAKENAPAIIFIDEVDAIATARFDAQTG 924
Query: 533 GGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIG 592
+E L +LL +DGF+ V I +TNR D LDPAL+RPGR DRKI P P
Sbjct: 925 ADREVQRILMELLNQMDGFDQTVNVKVIMATNRADTLDPALLRPGRLDRKIEFPLPDRRQ 1104
Query: 593 RIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDD 650
+ + +V K ++++VD E S D + AE+A I + A ++ +R +R + D
Sbjct: 1105KRLVFQVCTAKMNLSDEVDLEDYVSRPDKISAAEIAAICQEAGMHAVRKNRYVILPKD 1278
>TC205252 homologue to UP|Q9ZP50 (Q9ZP50) FtsH-like protein Pftf precursor,
partial (61%)
Length = 1482
Score = 164 bits (415), Expect = 2e-40
Identities = 120/339 (35%), Positives = 179/339 (52%), Gaps = 24/339 (7%)
Frame = +3
Query: 525 KRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIF 584
KR G G ER+ TLNQLL +DGFEG +I +A+TNR DILD AL+RPGRFDR++
Sbjct: 195 KRDWKWGEGNDEREQTLNQLLTEMDGFEGNTGIIVVAATNRADILDSALLRPGRFDRQVT 374
Query: 585 IPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRT 644
+ P GR EILKVHA K DV E++A T G GA+LAN++ AAI R +T
Sbjct: 375 VDVPDIRGRTEILKVHASNKKFDADVSLEVIAMRTPGFSGADLANLLNEAAILAGRRGKT 554
Query: 645 EVST---DDLLQ--AAQMEERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYI 699
+S+ DD + A ME M D K +S VA +E A+ P D ++ +
Sbjct: 555 AISSKEIDDSIDRIVAGMEGTVMTDGKSKS-----LVAYHEVGHAICGTLTPGHDAVQKV 719
Query: 700 TIAPRAGRELGYVRTMLESI-NFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIW 758
T+ PR G R + I N + ++++Q LF I L RAA+E+ FG+ +++T
Sbjct: 720 TLVPR-----GQARGLTWFIPNDDPTLISKQQLFARIVGGLGGRAAEEIIFGEPEVTTGA 884
Query: 759 A-ETADNARVAARMYMIGGLSD----------------KYRGVSNFWVTDRINE-IDLEA 800
A + +A +M G+SD R ++ +++R+ E ID
Sbjct: 885 AGDLQQITGLAKQMVTTFGMSDIGPWSLMEASAQSGDVIMRMMARNSMSERLAEDIDAAI 1064
Query: 801 MKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTKED 839
+I + YE A + ++ N+ +D +V L+ K+TLT ++
Sbjct: 1065KRISDEAYEIALDHIRNNREAIDKIVEVLLEKETLTGDE 1181
Score = 111 bits (277), Expect = 2e-24
Identities = 53/79 (67%), Positives = 67/79 (84%)
Frame = +1
Query: 461 GVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELD 520
G GKTLLAKA+AGEAGV FFSIS S+FVE++VGVGASRVR L+++AKENAP +VF+DE+D
Sbjct: 1 GTGKTLLAKAIAGEAGVPFFSISGSEFVEMFVGVGASRVRDLFKKAKENAPCIVFVDEID 180
Query: 521 AVGRKRGLIKGSGGQERDA 539
AVGR+RG G G +++
Sbjct: 181 AVGRQRGTGNGGKGMMKES 237
>AW201375 homologue to GP|17065470|gb cell division protein FtsH-like protein
{Arabidopsis thaliana}, partial (20%)
Length = 385
Score = 156 bits (394), Expect = 4e-38
Identities = 80/130 (61%), Positives = 100/130 (76%)
Frame = +3
Query: 464 KTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVG 523
KTLLA+AVAGEAGV FF++SAS+FVE+ VG GA+R+R L+ A++ APS++FIDELDAVG
Sbjct: 3 KTLLARAVAGEAGVPFFTVSASEFVELVVGRGAARIRDLFNAARKFAPSIIFIDELDAVG 182
Query: 524 RKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKI 583
KRG S ERD TLNQLL +DGFE V+ IA+TNRP+ LDPAL RPGRF RK+
Sbjct: 183 GKRG---RSFNDERDQTLNQLLTEMDGFESEMRVVVIAATNRPEALDPALCRPGRFSRKV 353
Query: 584 FIPKPGFIGR 593
++ +P GR
Sbjct: 354 YVGEPDEEGR 383
>TC227714 similar to UP|C48C_ARATH (Q9SS94) Cell division control protein 48
homolog C (AtCDC48c), partial (22%)
Length = 918
Score = 153 bits (386), Expect = 4e-37
Identities = 85/198 (42%), Positives = 119/198 (59%), Gaps = 2/198 (1%)
Frame = +1
Query: 453 GILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPS 512
G LL GPPG GKTL+AKAVA EAG F I + + YVG VR+++ A+ AP
Sbjct: 10 GFLLYGPPGCGKTLIAKAVANEAGATFIHIKGPELLNKYVGESELAVRTMFSRARTCAPC 189
Query: 513 VVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPA 572
++F DE+DA+ KRG GG + LNQLLV LDG E R V I +TNRP+++D A
Sbjct: 190 ILFFDEIDALTTKRG---KEGGWVVERLLNQLLVELDGAEQRKGVFVIGATNRPEVMDRA 360
Query: 573 LVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASM--TDGMVGAELANI 630
++RPGRF + +++P P R+ ILK ARKK + VD +A M + + GA+LA +
Sbjct: 361 VLRPGRFGKLLYVPLPSPDERVLILKALARKKAVDASVDLSAIAKMEACENLSGADLAAL 540
Query: 631 VEVAAINMMRDSRTEVST 648
+ AA+ + + T + T
Sbjct: 541 MNEAAMAALEERLTSIET 594
>BE661034 similar to SP|Q9ZPR1|C48B Cell division control protein 48 homolog
B (AtCDC48b). [Mouse-ear cress] {Arabidopsis thaliana},
partial (38%)
Length = 882
Score = 152 bits (384), Expect = 6e-37
Identities = 82/203 (40%), Positives = 114/203 (55%)
Frame = +1
Query: 439 HGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASR 498
H + G+ GILL GPPG KT LAKA A A +FFS+S ++ +YVG G +
Sbjct: 4 HSAAFSXMGISPVRGILLHGPPGCSKTTLAKAAAHAAQASFFSLSGAELYSMYVGEGEAL 183
Query: 499 VRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVI 558
+R +Q A+ APS++F DE D V KRG + + L+ LL +DG E ++
Sbjct: 184 LRKTFQRARLAAPSIIFFDEADVVAAKRGDSSSNSATVGERLLSTLLTEIDGLEEAKGIL 363
Query: 559 TIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASM 618
+A+TNRP +D AL+RPGRFD +++P P R EIL VH RK DVD +A
Sbjct: 364 VLAATNRPYAIDAALMRPGRFDLVLYVPPPDLEARHEILCVHTRKMKTGNDVDLRRIAED 543
Query: 619 TDGMVGAELANIVEVAAINMMRD 641
T+ GAEL + + A I +R+
Sbjct: 544 TELFTGAELEGLCKEAGIVALRE 612
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.134 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,134,412
Number of Sequences: 63676
Number of extensions: 376069
Number of successful extensions: 2769
Number of sequences better than 10.0: 168
Number of HSP's better than 10.0 without gapping: 2705
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2733
length of query: 881
length of database: 12,639,632
effective HSP length: 106
effective length of query: 775
effective length of database: 5,889,976
effective search space: 4564731400
effective search space used: 4564731400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC138130.19


