

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138130.17 + phase: 0 /pseudo
(626 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
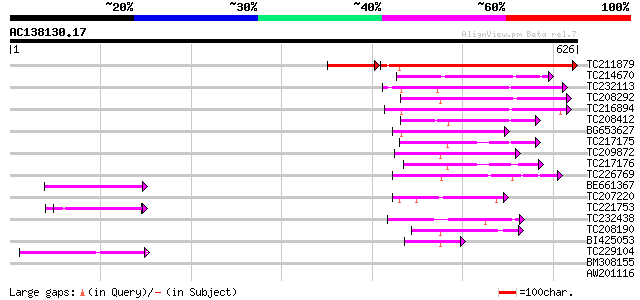
Score E
Sequences producing significant alignments: (bits) Value
TC211879 weakly similar to UP|Q9SY76 (Q9SY76) F14N23.22 (At1g103... 322 e-100
TC214670 similar to UP|Q8LED2 (Q8LED2) Ankyrin-like protein, par... 100 3e-21
TC232113 similar to UP|Q9LVG7 (Q9LVG7) Ankyrin-like protein, par... 99 7e-21
TC208292 90 3e-18
TC216894 similar to UP|Q9ZU96 (Q9ZU96) Expressed protein, partia... 84 2e-16
TC208412 similar to UP|Q9SKB8 (Q9SKB8) Ankyrin-like protein, par... 83 4e-16
BG653627 76 5e-14
TC217175 weakly similar to UP|Q9FG97 (Q9FG97) Ankyrin-like prote... 65 1e-10
TC209872 60 4e-09
TC217176 similar to GB|AAO42978.1|28416895|BT004732 At3g13950 {A... 60 4e-09
TC226769 54 2e-07
BE661367 similar to PIR|E84725|E84 ankyrin-like protein [importe... 51 1e-06
TC207220 weakly similar to UP|Q9LSB0 (Q9LSB0) Emb|CAB70981.1, pa... 51 2e-06
TC221753 similar to UP|Q9LVG7 (Q9LVG7) Ankyrin-like protein, par... 50 2e-06
TC232438 weakly similar to UP|Q6ZJG2 (Q6ZJG2) Ankyrin-like prote... 50 2e-06
TC208190 49 5e-06
BI425053 45 1e-04
TC229104 weakly similar to UP|Q8LLW2 (Q8LLW2) Receptor-like kina... 44 2e-04
BM308155 weakly similar to GP|14279688|gb receptor-like kinase X... 42 0.001
AW201116 42 0.001
>TC211879 weakly similar to UP|Q9SY76 (Q9SY76) F14N23.22
(At1g10340/F14N23_22), partial (25%)
Length = 895
Score = 322 bits (826), Expect(2) = e-100
Identities = 167/222 (75%), Positives = 186/222 (83%), Gaps = 5/222 (2%)
Frame = +3
Query: 410 NLVKQKHHHNKGKIENVNH-----TKRKHYHEMHKEALLNARNTIVLVAVLIATVTFAAG 464
NL K K NK K EN+N ++R ++EMHKEA+LNARNTI +VAVLIATVTFAAG
Sbjct: 189 NLGKHKQQ-NKTKAENLNQLYYTQSRRNKHYEMHKEAILNARNTITIVAVLIATVTFAAG 365
Query: 465 ISPPGGVYQEGPKKGISMAGETSAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTIL 524
I+PPGGVYQEGP +G SM G+T+AFKVFAISN IALFTSLS+VIVLVSIIPFRRKPQ L
Sbjct: 366 INPPGGVYQEGPMRGKSMVGKTTAFKVFAISNNIALFTSLSIVIVLVSIIPFRRKPQIRL 545
Query: 525 LTIAHKVMWVAVAFMGTGYVAATWVILPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLV 584
LTI HKVMWVAVAFM TGYVA TWVILPH+ EMQWLSVVLLA+GGG LGTIFI LSVMLV
Sbjct: 546 LTITHKVMWVAVAFMATGYVAGTWVILPHSPEMQWLSVVLLAVGGGSLGTIFIGLSVMLV 725
Query: 585 EHWLRKSSWRKKRKESGDGTAESDKESEDSDFQSSYLQGYHS 626
+HWLRKS W+K KES D A+ KESE+SDF+SSYLQGYHS
Sbjct: 726 DHWLRKSRWKKTMKESVDVAADYQKESENSDFESSYLQGYHS 851
Score = 60.8 bits (146), Expect(2) = e-100
Identities = 33/60 (55%), Positives = 40/60 (66%), Gaps = 3/60 (5%)
Frame = +2
Query: 352 SLSRRYISKEMEV--LTEMVSYDCISPPPVSESTESISP-QPQVSERFENGTYNPYYFSP 408
S+S RY + +E+ EMV+YDC SPP + ST S SP QPQVSER E+ TY YY SP
Sbjct: 5 SMSWRYTTNPVELPNQNEMVAYDCTSPPQLGRSTNSRSPSQPQVSERIEDTTYKSYYCSP 184
>TC214670 similar to UP|Q8LED2 (Q8LED2) Ankyrin-like protein, partial (50%)
Length = 1352
Score = 100 bits (248), Expect = 3e-21
Identities = 53/173 (30%), Positives = 101/173 (57%)
Frame = +3
Query: 428 HTKRKHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPKKGISMAGETS 487
H K ++H+E + NA N++ +VAVL ATV FAA + PGG +G ++ +
Sbjct: 285 HNISKELRKLHREGINNATNSVTVVAVLFATVAFAAIFTVPGGDDDDGS----AVVAAYA 452
Query: 488 AFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVAAT 547
AFK+F + N IALFTSL+VV+V ++++ K + ++ + +K+MW+A ++A++
Sbjct: 453 AFKIFFVFNAIALFTSLAVVVVQITLVRGETKAEKRVVEVINKLMWLASVCTSVAFIASS 632
Query: 548 WVILPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEHWLRKSSWRKKRKES 600
++++ E W ++++ +GG + + +++ +V R S RKK K++
Sbjct: 633 YIVVGRKNE--WAAILVTLVGGVIISGVIGTMTYYVVRS-KRSRSMRKKEKQA 782
>TC232113 similar to UP|Q9LVG7 (Q9LVG7) Ankyrin-like protein, partial (51%)
Length = 1086
Score = 98.6 bits (244), Expect = 7e-21
Identities = 65/216 (30%), Positives = 114/216 (52%), Gaps = 12/216 (5%)
Frame = +2
Query: 412 VKQKHHHNKGKIENVNHTKR------KHYHEMHKEALLNARNTIVLVAVLIATVTFAAGI 465
+K + H+ ++E+ T+R K ++MH E L NA N+ +VAVLIATV FAA
Sbjct: 326 IKHEVHY---QLEHTRQTRRGVQGIAKRINKMHAEGLNNAINSTTVVAVLIATVAFAAIF 496
Query: 466 SPPGG------VYQEGPKKGISMAGETSAFKVFAISNIIALFTSLSVVIVLVSIIPFRRK 519
+ PG V G G + +AF +F + + IALF SL+VV+V S++ K
Sbjct: 497 TVPGQFADDPKVLPAGMTIGEANIAPQAAFLIFFVFDSIALFISLAVVVVQTSVVIIESK 676
Query: 520 PQTILLTIAHKVMWVAVAFMGTGYVAATWVILPHNQEMQWLSVVLLALGGGFLGTIFISL 579
+ ++ I +K+MW+A + ++A ++V++ ++ +WL++ + +G + T ++
Sbjct: 677 AKKQMMAIINKLMWLACVLISVAFLALSFVVV--GKDQKWLAIGVTIIGTTIMATTLGTM 850
Query: 580 SVMLVEHWLRKSSWRKKRKESGDGTAESDKESEDSD 615
S ++ H + S+ R RK S + S S SD
Sbjct: 851 SYWVIRHRIEASNLRSIRKSSMGSRSRSFSVSVMSD 958
>TC208292
Length = 873
Score = 89.7 bits (221), Expect = 3e-18
Identities = 59/197 (29%), Positives = 100/197 (49%), Gaps = 8/197 (4%)
Frame = +3
Query: 432 KHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQE------GPKKGISMAGE 485
K ++MH E L NA N+ +VAVLIATV FAA + PG ++ G G +
Sbjct: 51 KRINKMHTEGLNNAINSNTIVAVLIATVAFAAIFNVPGQYPEKQNELSPGMSPGEAYIAP 230
Query: 486 TSAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVA 545
FK+F I + ALF SL+VVIV S++ RK + ++ + +K+MWVA + ++A
Sbjct: 231 DIGFKIFIIFDSTALFISLAVVIVQTSVVVIERKAKRQMMAVINKLMWVACVLISVAFIA 410
Query: 546 ATWVILPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEHWLRKSSWRKKR--KESGDG 603
+++I+ ++E L++ LG + +L ++ H L S R R S
Sbjct: 411 MSYIIVGDHKE---LAIAATVLGTVIMAATLGTLCYWVITHHLEASRLRSLRTTMSSRQS 581
Query: 604 TAESDKESEDSDFQSSY 620
+ S ++++++ Y
Sbjct: 582 MSMSMMSGSENEYKTVY 632
>TC216894 similar to UP|Q9ZU96 (Q9ZU96) Expressed protein, partial (54%)
Length = 1224
Score = 83.6 bits (205), Expect = 2e-16
Identities = 59/222 (26%), Positives = 113/222 (50%), Gaps = 16/222 (7%)
Frame = +3
Query: 415 KHHHNKGKIENVNHTKR-----KHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPG 469
KH I+N +R K ++H+EA+ N N++ LVAVL A++ F A + PG
Sbjct: 267 KHEVQSQLIQNETTRRRVSGIAKELKKLHREAVQNTINSVTLVAVLFASIAFLAIFNLPG 446
Query: 470 G-VYQEGPKKGISMAGETSAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIA 528
+ EG + G + + +F+VF + N +LF SL+VV+V ++++ + + Q ++++
Sbjct: 447 QYITDEGKEIGKAKIADHVSFQVFCLLNSTSLFISLAVVVVQITLVAWDTRAQKQIVSVV 626
Query: 529 HKVMWVAVAFMGTGYVAATWVILPHNQEMQWLSVVLLALGGGFL-GTIFISLSVMLVEHW 587
+K+MW A A ++A + ++ + W+++ + LG L GT+ + +H+
Sbjct: 627 NKLMWAACACTCGAFLAIAFEVV---GKKTWMAITITLLGVPVLVGTLASMCYFVFRQHF 797
Query: 588 --LRKSSWRKKRKESGDGTAE-------SDKESEDSDFQSSY 620
R S R+ ++ SG + SD + +SD + Y
Sbjct: 798 GIFRSDSQRRIKRASGSKSFSWSYSANISDLDEYNSDIEKIY 923
>TC208412 similar to UP|Q9SKB8 (Q9SKB8) Ankyrin-like protein, partial (39%)
Length = 779
Score = 82.8 bits (203), Expect = 4e-16
Identities = 49/159 (30%), Positives = 87/159 (53%), Gaps = 4/159 (2%)
Frame = +3
Query: 432 KHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPKKGISMA----GETS 487
K ++H L NA N+ +VAVLIATV FAA + PG Y E G S+ +
Sbjct: 312 KKLKKLHISGLNNAINSATVVAVLIATVAFAAIFTVPGQ-YVEDKTHGFSLGQANIANNA 488
Query: 488 AFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVAAT 547
AF +F + + +ALF SL+VV+V S++ +K + L+ + +K+MW+A F+ +++ T
Sbjct: 489 AFLIFFVFDSLALFISLAVVVVQTSVVVIEQKAKKQLVFVINKLMWMACLFISIAFISLT 668
Query: 548 WVILPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEH 586
+V++ +WL++ +G + + S+ ++ H
Sbjct: 669 YVVV--GSHSRWLAIYATVIGSLIMLSTIGSMCYCVILH 779
>BG653627
Length = 413
Score = 75.9 bits (185), Expect = 5e-14
Identities = 40/134 (29%), Positives = 77/134 (56%), Gaps = 5/134 (3%)
Frame = +3
Query: 423 IENVNHTKR-----KHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPK 477
I+N KR K ++H+EA+ N N++ +VAVL ++ F A S PG ++ P+
Sbjct: 3 IQNEKTRKRVSCIAKELKKIHREAVQNTINSVTVVAVLFGSIAFMALFSLPGQYRKKQPE 182
Query: 478 KGISMAGETSAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVA 537
G + + +AF F + N ALF SL+VV+ ++++ + + Q ++++ +K+MW A A
Sbjct: 183 AGKANIADDAAFSAFCLLNATALFLSLAVVVAQITLVAWDTRSQRQVVSVINKLMWAACA 362
Query: 538 FMGTGYVAATWVIL 551
++A ++V++
Sbjct: 363 CTCGAFLAISFVVV 404
>TC217175 weakly similar to UP|Q9FG97 (Q9FG97) Ankyrin-like protein, partial
(4%)
Length = 792
Score = 64.7 bits (156), Expect = 1e-10
Identities = 43/160 (26%), Positives = 81/160 (49%), Gaps = 4/160 (2%)
Frame = +2
Query: 431 RKHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPKKGIS----MAGET 486
R +E ++ + RN ++++ L+A VTF AG++PPGGV+QE + I+ A +
Sbjct: 44 RYFQYEEERDTPSDTRNILLIIFTLVAAVTFQAGVNPPGGVWQETNGEHIAGRAIYASDK 223
Query: 487 SAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVAA 546
A+ VF I N +A S+ V++ L + PF H + VA M Y +A
Sbjct: 224 QAYYVFLIFNTLAFSNSILVILSLTNKFPF------------HFEICVATVSMAVTYGSA 367
Query: 547 TWVILPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEH 586
+ + P ++ +++ V++ A G + + +++L +H
Sbjct: 368 IFAVSP-DESVRFRYVLITAAGPFVFRFLVLIFNLLLRKH 484
>TC209872
Length = 818
Score = 59.7 bits (143), Expect = 4e-09
Identities = 40/145 (27%), Positives = 75/145 (51%), Gaps = 6/145 (4%)
Frame = +2
Query: 426 VNHTKRKHYHEMHK--EALLNARNTIVLVAVLIATVTFAAGISPPGGVYQE---GPK-KG 479
VN+ R + ++ K +L + NT ++VA L+ TVTFAA + PGGVY PK +G
Sbjct: 74 VNNMLRSQHQQVSKTNSSLKDLINTFLVVATLMVTVTFAAAFTVPGGVYSSDDTNPKNRG 253
Query: 480 ISMAGETSAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFM 539
+++ F VF N+ A+++S+ +++ + F K T +A + +A +
Sbjct: 254 MAVLAHKRFFWVFTTFNMTAMYSSVLACGLMLMALIFDHKLATRTTILAMSCLILAFVTV 433
Query: 540 GTGYVAATWVILPHNQEMQWLSVVL 564
++AA +++ +N + L V+
Sbjct: 434 PVAFMAAVRLVVANNSALSLLITVI 508
>TC217176 similar to GB|AAO42978.1|28416895|BT004732 At3g13950 {Arabidopsis
thaliana;} , partial (12%)
Length = 594
Score = 59.7 bits (143), Expect = 4e-09
Identities = 42/159 (26%), Positives = 73/159 (45%), Gaps = 4/159 (2%)
Frame = +1
Query: 435 HEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPKKGIS----MAGETSAFK 490
+E ++ RN ++++ L+A VTF AG++PPGGV+QE ++ A +T A+
Sbjct: 94 YEEERDTPSETRNILLIIFTLVAAVTFQAGVNPPGGVWQEDKDGHVAGRAIYASDTQAYY 273
Query: 491 VFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVAATWVI 550
F I N +A S+ V++ L PF H + VA M Y ++ + +
Sbjct: 274 TFLIFNTLAFSNSILVILSLTHKFPF------------HFEICVATISMAVTYGSSIFAV 417
Query: 551 LPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEHWLR 589
P S+ L + G + + L V++ +LR
Sbjct: 418 SPKK------SINLRFIPVYAAGPVLVRLLVLIFNLYLR 516
>TC226769
Length = 1023
Score = 53.9 bits (128), Expect = 2e-07
Identities = 49/210 (23%), Positives = 89/210 (42%), Gaps = 22/210 (10%)
Frame = +2
Query: 423 IENVNHTKRKHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEG------- 475
+ N K+ H+ E L + R + L+A +IAT+TF + I+PPGG+
Sbjct: 59 MSNSKQKKKLKAHKKKDEWLKDMRGNLSLLATVIATMTFQSAINPPGGIRPASETGEITC 238
Query: 476 -----------PKKGISMAGETSAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTIL 524
P + + + + F N I +SL+V ++LVS +P +
Sbjct: 239 PDTSKNITVPCPGEAVLSVLKADTYNSFLYCNTICFASSLAVCLLLVSGLPLNNRFFIWF 418
Query: 525 LTIAHKVMWVAVAFMGTGYVAATWVILPH----NQEMQWLSVVLLALGGGFLGTIFISLS 580
+I M + + + Y+ ++ P+ N + VV+ + LG + I LS
Sbjct: 419 FSIC---MCITLTALTLTYLYGLQMVTPNDVWDNSLFSMVGVVIF-IWIILLGIVVIFLS 586
Query: 581 VMLVEHWLRKSSWRKKRKESGDGTAESDKE 610
+ L+ W+ KK+ E G+ +S +E
Sbjct: 587 LRLL-FWIVTKCRNKKQTEQGEDQNQSKQE 673
>BE661367 similar to PIR|E84725|E84 ankyrin-like protein [imported] -
Arabidopsis thaliana, partial (29%)
Length = 791
Score = 51.2 bits (121), Expect = 1e-06
Identities = 39/115 (33%), Positives = 55/115 (46%), Gaps = 1/115 (0%)
Frame = +3
Query: 39 PLHLASKYGCIEMVSEIVK-LCPDMVSAENKNMETPIHEACRQENVKVLMLLLEVNPTAA 97
PL++AS+ G +VSEI+K L S KN P H A +Q +++VL LL P A
Sbjct: 327 PLYVASENGHALVVSEILKYLDLQTASIAAKNGYDPFHIAAKQGHLEVLRELLHSFPNLA 506
Query: 98 CKLNPTCKSAFLVACSHGHLDLVNLLLNLSEIVGQEVAGFDQACFHVAAVRGHTE 152
+ + +A A + GH+D+ NLLL + H AA GH E
Sbjct: 507 MTTDLSNSTALHTAATQGHIDVGNLLLESDSXLAXIARN*W*TVLHSAARMGHLE 671
>TC207220 weakly similar to UP|Q9LSB0 (Q9LSB0) Emb|CAB70981.1, partial (16%)
Length = 880
Score = 50.8 bits (120), Expect = 2e-06
Identities = 45/153 (29%), Positives = 69/153 (44%), Gaps = 25/153 (16%)
Frame = +2
Query: 423 IENVNH---TKRKHYHEMHKEALLNARN-------TIVLVAVLIATVTFAAGISPPGGVY 472
IE NH R+ + E HKE L + + ++V+ LIAT F+A S PGG
Sbjct: 251 IERPNHEGIVPRELFTEKHKELLKKGESWMKRTASSCMVVSTLIATGVFSAAFSVPGGTK 430
Query: 473 QEGPKKGISMAGETSAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVM 532
+ G + F VFAIS+ +AL S + ++ +SI+ R + L ++ K++
Sbjct: 431 DD---SGSPNYLKKHLFTVFAISDALALTLSTASTLIFLSILISRYAEEDFLRSLPFKLI 601
Query: 533 WVAV---------------AFMGTGYVAATWVI 550
+ V AF T Y A TWV+
Sbjct: 602 FGLVSLFLSIVSMMGAFSSAFFITYYHAKTWVV 700
>TC221753 similar to UP|Q9LVG7 (Q9LVG7) Ankyrin-like protein, partial (31%)
Length = 762
Score = 50.4 bits (119), Expect = 2e-06
Identities = 31/112 (27%), Positives = 55/112 (48%)
Frame = +3
Query: 40 LHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVLMLLLEVNPTAACK 99
LH+A+K G +++V +++ P++ + + T +H A Q + +++ LLLE A
Sbjct: 93 LHIAAKQGDLDIVKILMEAHPELSMTVDPSNTTAVHTAALQGHTEIVKLLLEAGSNLATI 272
Query: 100 LNPTCKSAFLVACSHGHLDLVNLLLNLSEIVGQEVAGFDQACFHVAAVRGHT 151
K+A A +GHL++V LL V Q H+ AV+G +
Sbjct: 273 SRSNGKTALHSAARNGHLEVVKALLGKEPSVATRTDKKGQTAIHM-AVKGQS 425
Score = 44.3 bits (103), Expect = 2e-04
Identities = 27/106 (25%), Positives = 53/106 (49%), Gaps = 2/106 (1%)
Frame = +3
Query: 49 IEMVSEIVKLCPDMVSA--ENKNMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKS 106
++MV E+++ D+ A + +N +H A +Q ++ ++ +L+E +P + ++P+ +
Sbjct: 15 VDMVRELIQYY-DLAGAGIKARNGFDALHIAAKQGDLDIVKILMEAHPELSMTVDPSNTT 191
Query: 107 AFLVACSHGHLDLVNLLLNLSEIVGQEVAGFDQACFHVAAVRGHTE 152
A A GH ++V LLL + + H AA GH E
Sbjct: 192 AVHTAALQGHTEIVKLLLEAGSNLATISRSNGKTALHSAARNGHLE 329
Score = 32.7 bits (73), Expect = 0.50
Identities = 16/62 (25%), Positives = 31/62 (49%)
Frame = +3
Query: 24 REGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENV 83
+E + +TD +H+A K +E+V E++K P ++ + T +H A R+
Sbjct: 351 KEPSVATRTDKKGQTAIHMAVKGQSLEVVEELIKADPSTINMVDNKGNTALHIATRKGRA 530
Query: 84 KV 85
+V
Sbjct: 531 RV 536
>TC232438 weakly similar to UP|Q6ZJG2 (Q6ZJG2) Ankyrin-like protein, partial
(4%)
Length = 919
Score = 50.4 bits (119), Expect = 2e-06
Identities = 42/160 (26%), Positives = 78/160 (48%), Gaps = 9/160 (5%)
Frame = +1
Query: 418 HNKGKI-ENVNHTKRKHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGP 476
++KGK E V + + + + K+ N+ ++VA+L+ATV FAA ++ PG
Sbjct: 205 NDKGKTPEEVFYDQHEDLSDKIKDDSKEIANSGMIVAILVATVAFAAALTVPG------- 363
Query: 477 KKGISMAGETSA-FKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTIL-------LTIA 528
+T+A F VF +N +ALF S + ++ +S R Q LT
Sbjct: 364 -------EKTNAWFVVFIFTNAVALFASSASILSFLSNFTSLRFGQREFVKSLHPSLTFG 522
Query: 529 HKVMWVAVAFMGTGYVAATWVILPHNQEMQWLSVVLLALG 568
+++++V M + AA+++I H +W+S + ++G
Sbjct: 523 PVLLFISVVAMVVAFTAASFLIFDHTS--KWVSYAVASMG 636
>TC208190
Length = 1117
Score = 49.3 bits (116), Expect = 5e-06
Identities = 36/129 (27%), Positives = 70/129 (53%), Gaps = 5/129 (3%)
Frame = +2
Query: 444 NARNTIVLVAVLIATVTFAAGISPPGGVYQE---GPK-KGISMAGETSAFKVFAISNIIA 499
+ R ++VA L+ TV+FAAG + PGGVY PK +G ++ S F +F I N I
Sbjct: 473 DTREAFLIVAALLMTVSFAAGFTVPGGVYSSDDPNPKIRGTAVFAGNSVFWIFIIFNTIT 652
Query: 500 LFTS-LSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVAATWVILPHNQEMQ 558
+++S ++ ++ V I+ + + L + + +VAF+ AA +++ +N+ +
Sbjct: 653 MYSSAMACGLLSVGIVNRSKLSRFSDLFLTCAFLAASVAFL-----AAVLLVVANNRLLA 817
Query: 559 WLSVVLLAL 567
++++ AL
Sbjct: 818 GATILIGAL 844
>BI425053
Length = 412
Score = 45.1 bits (105), Expect = 1e-04
Identities = 25/72 (34%), Positives = 40/72 (54%), Gaps = 4/72 (5%)
Frame = +3
Query: 436 EMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQE---GPK-KGISMAGETSAFKV 491
+ H + + R ++VA L+ TV+FAA + PGGVY PK +G ++ F +
Sbjct: 117 QSHVQPGKDIREAFLIVAALLVTVSFAAAFTVPGGVYSSDDPNPKIRGTAVFARKPLFWI 296
Query: 492 FAISNIIALFTS 503
F I NII +++S
Sbjct: 297 FTIFNIITMYSS 332
>TC229104 weakly similar to UP|Q8LLW2 (Q8LLW2) Receptor-like kinase
Xa21-binding protein 3, partial (28%)
Length = 578
Score = 44.3 bits (103), Expect = 2e-04
Identities = 37/149 (24%), Positives = 69/149 (45%), Gaps = 5/149 (3%)
Frame = +1
Query: 11 KNDMITFSSIVKEREGILNQKTD--DTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENK 68
KN+ I SS + ++ Q + L A ++G +E+V+ ++ P ++
Sbjct: 97 KNNNINHSSPLLSPTIVMGQSLSCSGNYDHGLFTAVQHGDLEIVTTLLDSDPSLLHQTTL 276
Query: 69 -NMETPIHEACRQENVKVLMLLLE--VNPTAACKLNPTCKSAFLVACSHGHLDLVNLLLN 125
+ +P+H A + +++L LL+ +NP LN ++ ++A HG++ V LL
Sbjct: 277 YDRHSPLHIAATNDQIEILSKLLDGSLNPDV---LNRHKQTPLMLAAMHGNIACVEKLLQ 447
Query: 126 LSEIVGQEVAGFDQACFHVAAVRGHTESC 154
V + + C H AA GH+ SC
Sbjct: 448 AGANVLMFDTSYGRTCLHYAAYYGHS-SC 531
>BM308155 weakly similar to GP|14279688|gb receptor-like kinase Xa21-binding
protein 3 {Oryza sativa}, partial (28%)
Length = 433
Score = 41.6 bits (96), Expect = 0.001
Identities = 33/118 (27%), Positives = 58/118 (48%), Gaps = 3/118 (2%)
Frame = +2
Query: 40 LHLASKYGCIEMVSEIVKLCPDMVSAENK-NMETPIHEACRQENVKVLMLLLE--VNPTA 96
L A ++G ++ V+ +++ P +++ + +P+H A ++VL LL+ VNP
Sbjct: 53 LFRAVQHGDLDTVAALLQTHPSLMNHTTVYDHHSPLHIAAANGQIQVLSWLLDGSVNPDV 232
Query: 97 ACKLNPTCKSAFLVACSHGHLDLVNLLLNLSEIVGQEVAGFDQACFHVAAVRGHTESC 154
LN ++ ++A HG + V LL V A + + C H AA GH+ SC
Sbjct: 233 ---LNRQKQTPLMLAAMHGKIACVEKLLEAGANVLMFDACYGRTCLHYAAYYGHS-SC 394
>AW201116
Length = 364
Score = 41.6 bits (96), Expect = 0.001
Identities = 23/65 (35%), Positives = 41/65 (62%)
Frame = +2
Query: 450 VLVAVLIATVTFAAGISPPGGVYQEGPKKGISMAGETSAFKVFAISNIIALFTSLSVVIV 509
+L++ +IAT FAA I+ PGG+ +G K + ++F+VFAIS+ A S + +++
Sbjct: 152 MLISTVIATAVFAAAINIPGGI-DDGTNKPNYL--NKASFQVFAISDAAAFVFSATAILI 322
Query: 510 LVSII 514
+SI+
Sbjct: 323 FLSIL 337
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.330 0.140 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,989,934
Number of Sequences: 63676
Number of extensions: 405370
Number of successful extensions: 3303
Number of sequences better than 10.0: 76
Number of HSP's better than 10.0 without gapping: 3207
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3285
length of query: 626
length of database: 12,639,632
effective HSP length: 103
effective length of query: 523
effective length of database: 6,081,004
effective search space: 3180365092
effective search space used: 3180365092
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC138130.17


