

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138130.15 + phase: 0
(174 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
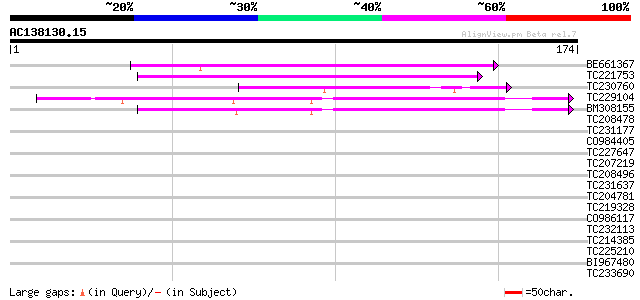
Score E
Sequences producing significant alignments: (bits) Value
BE661367 similar to PIR|E84725|E84 ankyrin-like protein [importe... 49 1e-06
TC221753 similar to UP|Q9LVG7 (Q9LVG7) Ankyrin-like protein, par... 49 1e-06
TC230760 weakly similar to UP|Q9LVG7 (Q9LVG7) Ankyrin-like prote... 46 1e-05
TC229104 weakly similar to UP|Q8LLW2 (Q8LLW2) Receptor-like kina... 42 1e-04
BM308155 weakly similar to GP|14279688|gb receptor-like kinase X... 40 6e-04
TC208478 weakly similar to UP|Q7X6P5 (Q7X6P5) Ankyrin, partial (... 38 0.003
TC231177 similar to UP|Q9SF90 (Q9SF90) F8A24.6 protein, partial ... 35 0.018
CO984405 33 0.052
TC227647 32 0.15
TC207219 similar to UP|Q8S9J9 (Q8S9J9) At1g14000/F7A19_9, partia... 32 0.15
TC208496 weakly similar to UP|Q7PZI6 (Q7PZI6) EbiP8728 (Fragment... 32 0.20
TC231637 31 0.26
TC204781 homologue to UP|UBC7_ARATH (Q42540) Ubiquitin-conjugati... 30 0.58
TC219328 similar to UP|Q7EZ44 (Q7EZ44) Receptor-like kinase Xa21... 28 1.7
CO986117 28 2.2
TC232113 similar to UP|Q9LVG7 (Q9LVG7) Ankyrin-like protein, par... 28 2.2
TC214385 homologue to UP|PRP2_SOYBN (P13993) Repetitive proline-... 27 3.7
TC225210 UP|PRP2_SOYBN (P13993) Repetitive proline-rich cell wal... 27 3.7
BI967480 27 3.7
TC233690 weakly similar to UP|Q6K393 (Q6K393) Ankyrin 3, epithel... 27 3.7
>BE661367 similar to PIR|E84725|E84 ankyrin-like protein [imported] -
Arabidopsis thaliana, partial (29%)
Length = 791
Score = 48.9 bits (115), Expect = 1e-06
Identities = 37/114 (32%), Positives = 55/114 (47%), Gaps = 1/114 (0%)
Frame = +3
Query: 38 TPLHLASKYGCIEMVSEIVR-LCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTA 96
TPL++AS+ G +VSEI++ L S +N P H A +Q +++VL LL P
Sbjct: 324 TPLYVASENGHALVVSEILKYLDLQTASIAAKNGYDPFHIAAKQGHLEVLRELLHSFPNL 503
Query: 97 ACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGH 150
A + + +A A + GH+D+ NLLL + H AA GH
Sbjct: 504 AMTTDLSNSTALHTAATQGHIDVGNLLLESDSXLAXIARN*W*TVLHSAARMGH 665
>TC221753 similar to UP|Q9LVG7 (Q9LVG7) Ankyrin-like protein, partial (31%)
Length = 762
Score = 48.9 bits (115), Expect = 1e-06
Identities = 29/106 (27%), Positives = 50/106 (46%)
Frame = +3
Query: 40 LHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAACK 99
LH+A+K G +++V ++ P++ + + T +H A Q + +++ LLLE A
Sbjct: 93 LHIAAKQGDLDIVKILMEAHPELSMTVDPSNTTAVHTAALQGHTEIVKLLLEAGSNLATI 272
Query: 100 LNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIA 145
K+A A +GHL++V LL V Q H+A
Sbjct: 273 SRSNGKTALHSAARNGHLEVVKALLGKEPSVATRTDKKGQTAIHMA 410
Score = 35.4 bits (80), Expect = 0.014
Identities = 19/79 (24%), Positives = 40/79 (50%)
Frame = +3
Query: 7 NAIKNNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAE 66
+A +N + +++ +E + RTD T +H+A K +E+V E+++ P ++
Sbjct: 303 SAARNGHLEVVKALLG-KEPSVATRTDKKGQTAIHMAVKGQSLEVVEELIKADPSTINMV 479
Query: 67 NENMETPIHEACRQENVKV 85
+ T +H A R+ +V
Sbjct: 480 DNKGNTALHIATRKGRARV 536
>TC230760 weakly similar to UP|Q9LVG7 (Q9LVG7) Ankyrin-like protein, partial
(11%)
Length = 976
Score = 45.8 bits (107), Expect = 1e-05
Identities = 30/95 (31%), Positives = 52/95 (54%), Gaps = 11/95 (11%)
Frame = +2
Query: 71 ETPIHEACRQENVKVLMLLLEVNPT-----AACKLNPTCKSAFFVACSHGHLDLVNLLLN 125
++P+ A R N+++++ ++ +P K N + ++A +VA +GHLD++ L+
Sbjct: 521 DSPLQSAIRVGNLELVLEIISQSPEDELKELLSKQNNSFETALYVAAENGHLDILKELIR 700
Query: 126 LSEIVEPGLA------GFDQACFHIAASRGHTGEN 154
+I GLA GFD FHIAA GH G++
Sbjct: 701 YHDI---GLASFKARNGFDP--FHIAAKNGHLGKS 790
>TC229104 weakly similar to UP|Q8LLW2 (Q8LLW2) Receptor-like kinase
Xa21-binding protein 3, partial (28%)
Length = 578
Score = 42.0 bits (97), Expect = 1e-04
Identities = 39/170 (22%), Positives = 76/170 (43%), Gaps = 5/170 (2%)
Frame = +1
Query: 9 IKNNDISTFSSIVKVREGILNQRTD--DTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAE 66
+KNN+I+ S ++ ++ Q ++ L A ++G +E+V+ ++ P ++
Sbjct: 94 VKNNNINHSSPLLSPTI-VMGQSLSCSGNYDHGLFTAVQHGDLEIVTTLLDSDPSLLHQT 270
Query: 67 N-ENMETPIHEACRQENVKVLMLLLE--VNPTAACKLNPTCKSAFFVACSHGHLDLVNLL 123
+ +P+H A + +++L LL+ +NP LN ++ +A HG++ V L
Sbjct: 271 TLYDRHSPLHIAATNDQIEILSKLLDGSLNPDV---LNRHKQTPLMLAAMHGNIACVEKL 441
Query: 124 LNLSEIVEPGLAGFDQACFHIAASRGHTGENKEFSLLCLHVFLTLLSTLP 173
L V + + C H AA GH+ CL L+ + P
Sbjct: 442 LQAGANVLMFDTSYGRTCLHYAAYYGHSS--------CLKAILSSAQSSP 567
>BM308155 weakly similar to GP|14279688|gb receptor-like kinase Xa21-binding
protein 3 {Oryza sativa}, partial (28%)
Length = 433
Score = 40.0 bits (92), Expect = 6e-04
Identities = 35/137 (25%), Positives = 61/137 (43%), Gaps = 3/137 (2%)
Frame = +2
Query: 40 LHLASKYGCIEMVSEIVRLCPDMVSAENE-NMETPIHEACRQENVKVLMLLLE--VNPTA 96
L A ++G ++ V+ +++ P +++ + +P+H A ++VL LL+ VNP
Sbjct: 53 LFRAVQHGDLDTVAALLQTHPSLMNHTTVYDHHSPLHIAAANGQIQVLSWLLDGSVNPDV 232
Query: 97 ACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGHTGENKE 156
LN ++ +A HG + V LL V A + + C H AA GH+
Sbjct: 233 ---LNRQKQTPLMLAAMHGKIACVEKLLEAGANVLMFDACYGRTCLHYAAYYGHSS---- 391
Query: 157 FSLLCLHVFLTLLSTLP 173
CL L+ + P
Sbjct: 392 ----CLKAILSAAQSSP 430
>TC208478 weakly similar to UP|Q7X6P5 (Q7X6P5) Ankyrin, partial (51%)
Length = 980
Score = 37.7 bits (86), Expect = 0.003
Identities = 33/123 (26%), Positives = 61/123 (48%), Gaps = 3/123 (2%)
Frame = +1
Query: 4 EFFNAIKNNDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMV 63
E A ++ D+ +SI+ +N R D TPLHLA+ G E+V+ + + D+
Sbjct: 106 ELHTAARSGDLIAVNSILASNPLAVNSR-DKHSRTPLHLAAFSGQAEVVTYLCKQKADVG 282
Query: 64 SAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCK---SAFFVACSHGHLDLV 120
++ ++M IH A ++ +++V+ LL +A L T + ++ A H++LV
Sbjct: 283 ASAMDDM-AAIHFASQKGHLEVVRALL----SAGASLKATTRKGMTSLHYAVQGSHMELV 447
Query: 121 NLL 123
L
Sbjct: 448 KYL 456
>TC231177 similar to UP|Q9SF90 (Q9SF90) F8A24.6 protein, partial (78%)
Length = 938
Score = 35.0 bits (79), Expect = 0.018
Identities = 24/83 (28%), Positives = 40/83 (47%), Gaps = 8/83 (9%)
Frame = +2
Query: 23 VREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPD------MVSAENENMETPIHE 76
+ G + D+ PLH A G E+V ++ D M+ + + +TP+H
Sbjct: 338 IERGANIEAKDEEGAIPLHDACAGGFTEIVQLLLNRANDAEHIKRMLESVDSEGDTPLHH 517
Query: 77 ACRQENVKVLMLLLE--VNPTAA 97
A R E+++V+ LLL +PT A
Sbjct: 518 AARGEHIEVIRLLLSNGASPTKA 586
Score = 26.6 bits (57), Expect = 6.4
Identities = 26/104 (25%), Positives = 44/104 (42%), Gaps = 5/104 (4%)
Frame = +2
Query: 33 DDTFNTPLHL-----ASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLM 87
DDT +TP HL A++ G + + + E+ +T +H C ++ +
Sbjct: 155 DDT-DTPPHLRDLSAAAQIGDAHALRIALDNLTGSIDEPVEDGDTALHLTCLYGHLACVQ 331
Query: 88 LLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVE 131
LL+E K + AC+ G ++V LLLN + E
Sbjct: 332 LLIERGANIEAK-DEEGAIPLHDACAGGFTEIVQLLLNRANDAE 460
>CO984405
Length = 677
Score = 33.5 bits (75), Expect = 0.052
Identities = 18/66 (27%), Positives = 34/66 (51%)
Frame = +1
Query: 61 DMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLV 120
++ NE+ +TP+HEA +V ++ + + A LN +S A G ++++
Sbjct: 322 EITRERNEHGDTPLHEAVYSGHVDLVKEIFGADMAAVHCLNKPKRSPQCAAVESGKVEIL 501
Query: 121 NLLLNL 126
NLLL +
Sbjct: 502 NLLLQI 519
Score = 32.0 bits (71), Expect = 0.15
Identities = 23/96 (23%), Positives = 47/96 (48%)
Frame = +1
Query: 22 KVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQE 81
++++ + + ++ +TPLH A G +++V EI V N+ +P A
Sbjct: 307 EMKDKEITRERNEHGDTPLHEAVYSGHVDLVKEIFGADMAAVHCLNKPKRSPQCAAVESG 486
Query: 82 NVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHL 117
V++L LLL++ A L+ + K + +CS+ +
Sbjct: 487 KVEILNLLLQIPFPADQPLSVSWKFSTPCSCSNSEV 594
Score = 29.6 bits (65), Expect = 0.75
Identities = 14/43 (32%), Positives = 23/43 (52%)
Frame = +1
Query: 40 LHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQEN 82
LH+A+ G E+ I P+++ N +TP+H A R +N
Sbjct: 115 LHVAADLGKEEITELIAHHFPELLIRRNVRGDTPLHVAVRSKN 243
>TC227647
Length = 1313
Score = 32.0 bits (71), Expect = 0.15
Identities = 26/108 (24%), Positives = 49/108 (45%), Gaps = 4/108 (3%)
Frame = +1
Query: 32 TDDTFNTPLHLASKYGCIEMVSEIVRLCPDM-VSAENENME--TPIHEACRQENVKVLML 88
TDD T LH+ ++ G +++V +++ D+ VSA + TP+H A ++ V+ +
Sbjct: 565 TDDRGWTSLHVFARKGDLKLVKKLLNEGMDVNVSAWGPKSKGVTPLHLAAEGGHIGVMDV 744
Query: 89 LLEVNPTAACKLNPTCK-SAFFVACSHGHLDLVNLLLNLSEIVEPGLA 135
LLE + C + +A D V L+ + P ++
Sbjct: 745 LLECGADIDARTKGACGWTPLHIAAKERRRDAVKFLIENGAFMPPDIS 888
>TC207219 similar to UP|Q8S9J9 (Q8S9J9) At1g14000/F7A19_9, partial (82%)
Length = 1189
Score = 32.0 bits (71), Expect = 0.15
Identities = 21/78 (26%), Positives = 41/78 (51%)
Frame = +2
Query: 12 NDISTFSSIVKVREGILNQRTDDTFNTPLHLASKYGCIEMVSEIVRLCPDMVSAENENME 71
ND + +++ ++ R D TPLH+AS +G I++ + ++ D V+A++
Sbjct: 176 NDAAAVRKLLQEDPSLVKARDYDN-RTPLHVASLHGWIDVATCLIEFGAD-VNAQDRWKN 349
Query: 72 TPIHEACRQENVKVLMLL 89
TP+ +A + V+ LL
Sbjct: 350 TPLADAEGAKKSNVIELL 403
>TC208496 weakly similar to UP|Q7PZI6 (Q7PZI6) EbiP8728 (Fragment), partial
(3%)
Length = 588
Score = 31.6 bits (70), Expect = 0.20
Identities = 12/30 (40%), Positives = 21/30 (70%)
Frame = +2
Query: 65 AENENMETPIHEACRQENVKVLMLLLEVNP 94
A N+ +T +HEA R +++V+ LLE++P
Sbjct: 149 ARNDEKDTALHEAVRNHHIEVVKTLLEMDP 238
>TC231637
Length = 649
Score = 31.2 bits (69), Expect = 0.26
Identities = 25/102 (24%), Positives = 45/102 (43%), Gaps = 12/102 (11%)
Frame = +1
Query: 37 NTPLHLASKYGCIEMVSEIVRLCP------------DMVSAENENMETPIHEACRQENVK 84
+TPLH+A++ E V I+ ++ NE TP+HEA +V
Sbjct: 130 DTPLHVAARSKKYETVKLILSQYATKQSTYDEMKDKEITRETNECGNTPLHEAVYSGDVD 309
Query: 85 VLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNL 126
V+ + + + LN + +S A +G+ ++ LLL +
Sbjct: 310 VVKEIFDQDKAVVHCLNKSKRSPLCSAVVNGNEQILELLLQI 435
Score = 31.2 bits (69), Expect = 0.26
Identities = 16/51 (31%), Positives = 27/51 (52%)
Frame = +1
Query: 40 LHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEACRQENVKVLMLLL 90
LH+A+ G +V I L P ++ N +TP+H A R + + + L+L
Sbjct: 37 LHVAADLGKETIVGRICDLFPLLLIRRNVRGDTPLHVAARSKKYETVKLIL 189
Score = 29.6 bits (65), Expect = 0.75
Identities = 13/41 (31%), Positives = 24/41 (57%)
Frame = +1
Query: 37 NTPLHLASKYGCIEMVSEIVRLCPDMVSAENENMETPIHEA 77
++PLH A ++ M+ I+ P++V +E+ TP+H A
Sbjct: 475 SSPLHTAIQHQRRVMIQAIIETRPELVYLRDEDGNTPLHYA 597
>TC204781 homologue to UP|UBC7_ARATH (Q42540) Ubiquitin-conjugating enzyme E2
7 (Ubiquitin-protein ligase 7) (Ubiquitin carrier
protein 7) , complete
Length = 894
Score = 30.0 bits (66), Expect = 0.58
Identities = 19/63 (30%), Positives = 29/63 (45%)
Frame = -2
Query: 101 NPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGHTGENKEFSLL 160
NP+CKS F+ H NLLL + P L F+ I +R H + ++L
Sbjct: 590 NPSCKSQHFL*LPH---TSANLLLEFISPISPLLRSFNICRRFIIRTRKHRYDT*YYTLY 420
Query: 161 CLH 163
C++
Sbjct: 419 CMN 411
>TC219328 similar to UP|Q7EZ44 (Q7EZ44) Receptor-like kinase Xa21-binding
protein 3-like, partial (51%)
Length = 1424
Score = 28.5 bits (62), Expect = 1.7
Identities = 14/46 (30%), Positives = 21/46 (45%)
Frame = +3
Query: 106 SAFFVACSHGHLDLVNLLLNLSEIVEPGLAGFDQACFHIAASRGHT 151
+A AC HGH ++V L+ + + H+AA GHT
Sbjct: 3 TALMQACQHGHWEVVQTLIIFNANIHKADYLNGGTVLHLAALNGHT 140
>CO986117
Length = 639
Score = 28.1 bits (61), Expect = 2.2
Identities = 20/70 (28%), Positives = 31/70 (43%), Gaps = 5/70 (7%)
Frame = -1
Query: 59 CPDMVSAENENMETPIHEACRQENVKVLMLLLEVNPTAAC-----KLNPTCKSAFFVACS 113
C ++ + N T +H A R K++ L+ +A +PT K+A +A S
Sbjct: 624 CGVNINFRDINGWTALHWAARFGREKMVASLIASGASAGAVTDPSSQDPTGKTAASIAAS 445
Query: 114 HGHLDLVNLL 123
HGH L L
Sbjct: 444 HGHKGLAGYL 415
>TC232113 similar to UP|Q9LVG7 (Q9LVG7) Ankyrin-like protein, partial (51%)
Length = 1086
Score = 28.1 bits (61), Expect = 2.2
Identities = 15/47 (31%), Positives = 23/47 (48%)
Frame = +2
Query: 84 KVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIV 130
+++ LLLE A K+A A +GHL++V LL +V
Sbjct: 146 QIVKLLLEAGSNLATIARSNGKTALHSAARNGHLEVVKALLGKEPVV 286
>TC214385 homologue to UP|PRP2_SOYBN (P13993) Repetitive proline-rich cell
wall protein 2 precursor, partial (61%)
Length = 439
Score = 27.3 bits (59), Expect = 3.7
Identities = 20/59 (33%), Positives = 30/59 (49%), Gaps = 6/59 (10%)
Frame = -3
Query: 77 ACRQENVKVLMLLLEVNPTAACKLNPT----CK--SAFFVACSHGHLDLVNLLLNLSEI 129
ACR E + + EV+P ACKL + CK ++ VAC+ LV +L S++
Sbjct: 374 ACRPEASQQVACK*EVSPLGACKLEVSLLVACKLEASLLVACTLEASQLVACILVASQL 198
>TC225210 UP|PRP2_SOYBN (P13993) Repetitive proline-rich cell wall protein 2
precursor, complete
Length = 964
Score = 27.3 bits (59), Expect = 3.7
Identities = 20/59 (33%), Positives = 30/59 (49%), Gaps = 6/59 (10%)
Frame = -3
Query: 77 ACRQENVKVLMLLLEVNPTAACKLNPT----CK--SAFFVACSHGHLDLVNLLLNLSEI 129
ACR E + + EV+P ACKL + CK ++ VAC+ LV +L S++
Sbjct: 665 ACRPEASQQVACK*EVSPLGACKLEVSLLVACKLEASLLVACTLEASQLVACILVASQL 489
>BI967480
Length = 713
Score = 27.3 bits (59), Expect = 3.7
Identities = 20/59 (33%), Positives = 30/59 (49%), Gaps = 6/59 (10%)
Frame = +3
Query: 77 ACRQENVKVLMLLLEVNPTAACKLNPT----CK--SAFFVACSHGHLDLVNLLLNLSEI 129
ACR E + + EV+P ACKL + CK ++ VAC+ LV +L S++
Sbjct: 261 ACRPEASQQVACK*EVSPLGACKLEVSLLVACKLEASLLVACTLEASQLVACILVASQL 437
>TC233690 weakly similar to UP|Q6K393 (Q6K393) Ankyrin 3, epithelial isoform
a-like, partial (13%)
Length = 1024
Score = 27.3 bits (59), Expect = 3.7
Identities = 16/59 (27%), Positives = 29/59 (49%)
Frame = +3
Query: 77 ACRQENVKVLMLLLEVNPTAACKLNPTCKSAFFVACSHGHLDLVNLLLNLSEIVEPGLA 135
A R+ + +L +LL+ + P C+ A A HG + LL++ S+ + P +A
Sbjct: 576 AVREGHFNILEILLKAGAS-----QPACEEALIEASCHGQAGCLELLMS-SDFIRPHVA 734
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.137 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,541,113
Number of Sequences: 63676
Number of extensions: 127734
Number of successful extensions: 606
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 590
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 602
length of query: 174
length of database: 12,639,632
effective HSP length: 91
effective length of query: 83
effective length of database: 6,845,116
effective search space: 568144628
effective search space used: 568144628
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 55 (25.8 bits)
Medicago: description of AC138130.15


