

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
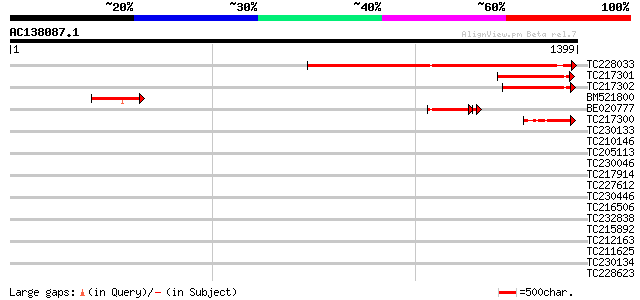
Query= AC138087.1 + phase: 0
(1399 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC228033 similar to UP|Q9FGK9 (Q9FGK9) Dbj|BAA90625.1, partial (... 984 0.0
TC217301 similar to UP|Q9FGK9 (Q9FGK9) Dbj|BAA90625.1, partial (5%) 251 1e-66
TC217302 234 2e-61
BM521800 201 1e-51
BE020777 171 1e-46
TC217300 113 5e-25
TC230133 weakly similar to UP|Q6VBJ3 (Q6VBJ3) Epa4p, partial (6%) 40 0.007
TC210146 homologue to GB|AAL32012.1|16930693|AF436830 AT3g07810/... 39 0.021
TC205113 37 0.047
TC230046 similar to UP|Q9LU79 (Q9LU79) Gb|AAF21150.1, partial (28%) 37 0.047
TC217914 similar to GB|AAN18208.1|23308477|BT000642 At2g29670/T2... 37 0.061
TC227612 weakly similar to UP|Q8L7S0 (Q8L7S0) At1g09730/F21M12_1... 35 0.18
TC230446 weakly similar to UP|Q6V9S3 (Q6V9S3) RPE1 protein, part... 34 0.39
TC216506 similar to GB|AAL15354.1|16323240|AY057724 AT5g14920/F2... 34 0.39
TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {A... 34 0.52
TC215892 weakly similar to UP|Q43522 (Q43522) Tfm5 protein, part... 33 0.67
TC212163 similar to UP|Q9LT26 (Q9LT26) Arabidopsis thaliana geno... 33 0.67
TC211625 33 0.67
TC230134 weakly similar to UP|NUCL_HUMAN (P19338) Nucleolin (Pro... 33 0.88
TC228623 similar to UP|Q9SA21 (Q9SA21) F3O9.2 protein, partial (... 33 1.1
>TC228033 similar to UP|Q9FGK9 (Q9FGK9) Dbj|BAA90625.1, partial (25%)
Length = 2651
Score = 984 bits (2545), Expect = 0.0
Identities = 511/669 (76%), Positives = 561/669 (83%), Gaps = 6/669 (0%)
Frame = +2
Query: 736 EGQLWGPALVLASQLGEKFYVDTVKQMALRQLVAGSPLRTLCLLIAGQPAEVFSSDSSNS 795
+GQLWGPALVLASQLGE+FYVDTVKQMALRQLV+GSPLRTLCLLIAGQ AE+FS+D+SNS
Sbjct: 5 QGQLWGPALVLASQLGEQFYVDTVKQMALRQLVSGSPLRTLCLLIAGQQAEIFSTDTSNS 184
Query: 796 GDPSAFNMPQNPAQFGSSGMLDDWEENLAVITSNRTKDDELVIIHLGDCLWKERSEITAA 855
G P A NM Q Q GS+GMLDDWEENLAVIT+NRTKDDELVIIHLGDCLWKERSEITAA
Sbjct: 185 GHPGASNMAQQSPQVGSNGMLDDWEENLAVITANRTKDDELVIIHLGDCLWKERSEITAA 364
Query: 856 HICYLIAEANFESYSDSARLCLIGADHWKFPRTYASPEAIQRTELYEYSKVLGNSQFILL 915
HICYL+AEANFESYSDSARLCLIGADHWK PRTYASPEAIQRTELYEYSKV+GNSQF L
Sbjct: 365 HICYLVAEANFESYSDSARLCLIGADHWKCPRTYASPEAIQRTELYEYSKVVGNSQFTLH 544
Query: 916 PFQPYKLIYAYMLAEVGKVSDSLKYCQAVLKSLKTGRAPEVETWKQRLSSLEERIRTHQQ 975
PFQPYKLIYA++LAEVGKVSDSLKYCQA+LKSLKTGRAPEVE+WKQ SLEERIR HQQ
Sbjct: 545 PFQPYKLIYAFLLAEVGKVSDSLKYCQALLKSLKTGRAPEVESWKQLALSLEERIRIHQQ 724
Query: 976 GGYAANLAPGKLVGKLLNFFDSTAHRVVGGLPPPAPSSQ-GNVHGNEQNYQSGAHRVSNS 1034
GGYAANLAP KLVGKLLNFFDSTAHRVVGGLPPPAPSS G VHG+E+ YQ+ A RVS+S
Sbjct: 725 GGYAANLAPAKLVGKLLNFFDSTAHRVVGGLPPPAPSSSAGTVHGSEKQYQNMAPRVSSS 904
Query: 1035 QSTMAMSSLVPSGSMEPNGEWTADNNRMTKSNRSVSEPDFGRSPRQE-TSHDAQGK--AS 1091
QSTM SL PS SMEP EWTADNNRM K NRSVS+PDFGR+PRQE TS DAQ K AS
Sbjct: 905 QSTM---SLAPSASMEPISEWTADNNRMGKPNRSVSDPDFGRTPRQETTSPDAQEKPQAS 1075
Query: 1092 EGTSRFSRFSFGSQLLQKTMGLVLKPRPGKQAKLGEKNKFYYDENLKRWVEEGAAPPAEE 1151
GTSRFSRF FGSQLLQKT+GLVLKPR G+QAKLG+KNKFYYDE LKRWVEEGA PAEE
Sbjct: 1076 GGTSRFSRFGFGSQLLQKTVGLVLKPRSGRQAKLGDKNKFYYDEKLKRWVEEGAEVPAEE 1255
Query: 1152 -TALPPPPTTAAFQNGLTEYNLQSALKTEGPPSKEGSDLKTSNPELTPGIPPIPPGTNHF 1210
AL PPPTTAAFQNG TEYNL+SALKTE P EGS ++TS+ EL+PG+P IPP N F
Sbjct: 1256 AAALTPPPTTAAFQNGSTEYNLRSALKTESSPPIEGSSIRTSSLELSPGMPLIPPSANQF 1435
Query: 1211 SARGRVGIRSRYVDTFNQGGGNSANLFQSPSVPSAKPVVAANAKFFIPTPAPSSNEQTME 1270
SARGR+G+RSRYVDTFNQGGG SANLFQSPSVPS KP VAANAKFFIP+ APSSNEQTME
Sbjct: 1436 SARGRLGVRSRYVDTFNQGGGTSANLFQSPSVPSVKPAVAANAKFFIPSAAPSSNEQTME 1615
Query: 1271 AIEENNQEDDLAYENPSTSYRND-WSFQSPKHASASTWQRCPSMGNFANHEAVVSGSNSR 1329
AI E+ QED E+PSTS N+ WS+QSPK S++T QR PS+GN +N A GSNS
Sbjct: 1616 AIVESKQEDSATNEDPSTSATNEWWSYQSPKQVSSTTIQRFPSLGNISNQRA-TEGSNSH 1792
Query: 1330 SPHSRRTVSWGGSTDVTYSPTKMREIMPLGEALGMPPSTYMSDDISSMRTSMKSGNFGED 1389
PHSRRT SW GS + +++P KM GMP S +M D+ S MRT +KS ++ ED
Sbjct: 1793 LPHSRRTSSWSGSFNDSFTPPKM----------GMPSSRFMPDE-SLMRTHVKSSSYAED 1939
Query: 1390 LHEVDLSTH 1398
L EV+L H
Sbjct: 1940 LQEVEL*AH 1966
>TC217301 similar to UP|Q9FGK9 (Q9FGK9) Dbj|BAA90625.1, partial (5%)
Length = 1110
Score = 251 bits (642), Expect = 1e-66
Identities = 124/190 (65%), Positives = 149/190 (78%)
Frame = +2
Query: 1205 PGTNHFSARGRVGIRSRYVDTFNQGGGNSANLFQSPSVPSAKPVVAANAKFFIPTPAPSS 1264
P +N FSARGR+G+RSRYVDTFNQGGG SANLFQSPSVPS KPV+AANAKFF+PTPAPSS
Sbjct: 8 PSSNQFSARGRLGVRSRYVDTFNQGGGTSANLFQSPSVPSVKPVLAANAKFFVPTPAPSS 187
Query: 1265 NEQTMEAIEENNQEDDLAYENPSTSYRNDWSFQSPKHASASTWQRCPSMGNFANHEAVVS 1324
NE+T+EAI E+ QED+ E PS S N+WS+QSPKH S++T QR PSMGN +N +
Sbjct: 188 NERTIEAIVESKQEDNATNEYPSISTTNEWSYQSPKHVSSTTIQRFPSMGNISN-QVAAD 364
Query: 1325 GSNSRSPHSRRTVSWGGSTDVTYSPTKMREIMPLGEALGMPPSTYMSDDISSMRTSMKSG 1384
G+NS PHSRRT SW GS + +++P KM I PLGEALGMPPS + S D S M +KS
Sbjct: 365 GNNSHLPHSRRTASWSGSFNDSFTPQKMGNIKPLGEALGMPPSRF-SPDESLMHKPVKSS 541
Query: 1385 NFGEDLHEVD 1394
++GEDLHEV+
Sbjct: 542 SYGEDLHEVE 571
>TC217302
Length = 590
Score = 234 bits (597), Expect = 2e-61
Identities = 117/181 (64%), Positives = 137/181 (75%)
Frame = +1
Query: 1215 RVGIRSRYVDTFNQGGGNSANLFQSPSVPSAKPVVAANAKFFIPTPAPSSNEQTMEAIEE 1274
R+G+RSRYVDTFNQGGG SANLFQSPSVPS KP +AANAKFF+PTPAPSSNEQ M+AI E
Sbjct: 1 RLGVRSRYVDTFNQGGGTSANLFQSPSVPSVKPALAANAKFFVPTPAPSSNEQAMDAIAE 180
Query: 1275 NNQEDDLAYENPSTSYRNDWSFQSPKHASASTWQRCPSMGNFANHEAVVSGSNSRSPHSR 1334
QED E PSTS NDWS++SPKH S++ QR PSMGN + + GSNS PHSR
Sbjct: 181 GKQEDSATNEYPSTSATNDWSYRSPKHVSSTAIQRFPSMGNISK-QGATEGSNSHLPHSR 357
Query: 1335 RTVSWGGSTDVTYSPTKMREIMPLGEALGMPPSTYMSDDISSMRTSMKSGNFGEDLHEVD 1394
RT SW GS + +++P KM + PLGEALGMP S Y S D SSM +KS ++GEDLHEV+
Sbjct: 358 RTASWSGSFNDSFTPQKMGNMKPLGEALGMPLSRY-SPDESSMHKPVKSSSYGEDLHEVE 534
Query: 1395 L 1395
L
Sbjct: 535 L 537
>BM521800
Length = 428
Score = 201 bits (512), Expect = 1e-51
Identities = 94/141 (66%), Positives = 112/141 (78%), Gaps = 11/141 (7%)
Frame = +1
Query: 203 DGMNASVDYVQYQEGQSYDASARNSTSGEDVNSSQYWESLYPGWKYDYNTGQWYQVDEHN 262
DG+NAS ++VQYQEG++Y AS+ +G+D++SSQYWE LYPGWKYD+NTGQWYQ+D +
Sbjct: 7 DGLNASANHVQYQEGETYVASSEEHPNGQDLSSSQYWEDLYPGWKYDHNTGQWYQIDGYI 186
Query: 263 ATAATQGSSEVNTA-----------EVSYMQQTAQSAVAGTLAESAATETVPSWNQVSQG 311
T+ TQ SSE NTA E+SYMQQTAQS VAGTLAES T+ V SW+QVS+G
Sbjct: 187 VTSTTQQSSEANTAADLSAASDGKTEISYMQQTAQS-VAGTLAESGTTKNVSSWSQVSEG 363
Query: 312 NNGYPEHMIFDPQYPGWYYDT 332
NNGYPEHMIFDPQYPGWYYDT
Sbjct: 364 NNGYPEHMIFDPQYPGWYYDT 426
>BE020777
Length = 413
Score = 171 bits (432), Expect(2) = 1e-46
Identities = 90/117 (76%), Positives = 97/117 (81%), Gaps = 3/117 (2%)
Frame = +3
Query: 1031 VSNSQSTMAMSSLVPSGSMEPNGEWTADNNRMTKSNRSVSEPDFGRSPRQET-SHDAQGK 1089
VS+SQSTM SL PS SMEP +WTADNN+M K NRS+SEPD GR+PRQET S D QGK
Sbjct: 6 VSSSQSTM---SLAPSASMEPISDWTADNNKMAKPNRSISEPDIGRTPRQETTSPDIQGK 176
Query: 1090 A--SEGTSRFSRFSFGSQLLQKTMGLVLKPRPGKQAKLGEKNKFYYDENLKRWVEEG 1144
A S GTSRFSRF FGSQLLQKT+GLVLKPR G+QAKLGEKNKFYYDE LKRWV EG
Sbjct: 177 AQASGGTSRFSRFGFGSQLLQKTVGLVLKPRSGRQAKLGEKNKFYYDEKLKRWVXEG 347
Score = 35.8 bits (81), Expect(2) = 1e-46
Identities = 18/23 (78%), Positives = 19/23 (82%), Gaps = 1/23 (4%)
Frame = +1
Query: 1143 EGAAPPAEETA-LPPPPTTAAFQ 1164
+GA PAEE A LPPPPTTAAFQ
Sbjct: 343 KGAELPAEEAAALPPPPTTAAFQ 411
>TC217300
Length = 793
Score = 113 bits (283), Expect = 5e-25
Identities = 65/127 (51%), Positives = 85/127 (66%)
Frame = +2
Query: 1269 MEAIEENNQEDDLAYENPSTSYRNDWSFQSPKHASASTWQRCPSMGNFANHEAVVSGSNS 1328
MEAI E+ QED S N+ S+QS K S++T QR PS+GN +N A G+NS
Sbjct: 2 MEAIAESKQED---------SAXNECSYQSXK--SSTTIQRFPSLGNISNQGAT-DGNNS 145
Query: 1329 RSPHSRRTVSWGGSTDVTYSPTKMREIMPLGEALGMPPSTYMSDDISSMRTSMKSGNFGE 1388
PHSRRT SW GS + +++P KM I PLGE+LGMPPS ++ D+ S MRT +KS ++GE
Sbjct: 146 HLPHSRRTASWSGSFNDSFTPRKMGNIKPLGESLGMPPSRFLPDE-SLMRTHVKSSSYGE 322
Query: 1389 DLHEVDL 1395
DL EV+L
Sbjct: 323 DLQEVEL 343
>TC230133 weakly similar to UP|Q6VBJ3 (Q6VBJ3) Epa4p, partial (6%)
Length = 850
Score = 40.0 bits (92), Expect = 0.007
Identities = 29/119 (24%), Positives = 49/119 (40%)
Frame = +3
Query: 12 QDDEDFFDKLVEDDVGNVNDEANDSDDVKAFSNLSIGGDDADVNASAFENSSGGGSGGEG 71
++++D D +DD NVND +D+DD + + GDD A + ++ G GG
Sbjct: 258 EENKDASDTEDDDDDDNVNDGEDDNDDEE---DEDFSGDDGGEEADSDDDPEANGGGGSD 428
Query: 72 KERKEEGDVKLDGGNVQEGSSSGCDGMMDRSDHGMESRNSSGSSADKSNRRSSLDVKEK 130
+ DG + ++ + DG D D G + + R+ L EK
Sbjct: 429 DD---------DGDDDEDDDNDEDDGDEDDEDEGDDDEDEESPQPPSKKRK*ELY*YEK 578
>TC210146 homologue to GB|AAL32012.1|16930693|AF436830 AT3g07810/F17A17_15
{Arabidopsis thaliana;} , partial (12%)
Length = 1163
Score = 38.5 bits (88), Expect = 0.021
Identities = 53/219 (24%), Positives = 84/219 (38%), Gaps = 17/219 (7%)
Frame = +3
Query: 421 SAYGGNQQFDHSFGSSNSVNKNQQNASSSFGSVPLYNKVNHGHGLVNGTVEVQRFAPSGN 480
+ +GGN F+ + +N +S+ FGS Y N G+ +V + GN
Sbjct: 231 AGFGGNANFNSNLSYDRGINPYFIGSSNRFGSPVGYESGNGGNNSFFSSVTRNLW---GN 401
Query: 481 FGQHYNYSNTQFDEQKNISNDYAESHQPFGYSNQSYQS---GHQQSYAPNVGRSS----- 532
G Y S+ + S + FG + ++ S QQ N+ +SS
Sbjct: 402 GGLSYGTSSANSNAYIG-SGSGSAGGNTFGNTGVNWSSSAISGQQGGGNNMSQSSGNLGY 578
Query: 533 -AGRPPHALVTFGFG--GKLIILKDSSLSSSTYGSQGAAQGSVSVLNL------MEAVSG 583
G + L T G+G I SS S+S G GA + ++ A S
Sbjct: 579 GGGDNNYGLGTGGYGRSSGAIFAPTSSYSASNGGVDGAFADFYNNSSVYGDPTWRSANSE 758
Query: 584 SIGSSSIGNGAGDYFRALGQQSIPGPLVGGSVGSKELNK 622
GS G G G + +S PG + G +V ++ N+
Sbjct: 759 RDGSGPFGYGLGGAASDVSAKSSPGYVGGYTVNKRQPNR 875
>TC205113
Length = 733
Score = 37.4 bits (85), Expect = 0.047
Identities = 41/149 (27%), Positives = 62/149 (41%), Gaps = 4/149 (2%)
Frame = +2
Query: 438 SVNKNQQNASSSFGSVPLYNKVNHGHGLVNGTVEVQRFAPSGNF---GQHYNYSNTQFDE 494
S K+ +A SF + NK++ G ++ TV+V S N G + N N+ F +
Sbjct: 131 SYYKSPASADVSFTT----NKISLPEGEISTTVDVADCVSSTNAMNKGNNVNVGNSNFTD 298
Query: 495 QKNISNDYAESHQPFGYSNQSYQSGHQQSYAPNVGRSSAGRPPHALVTFGFGGKLIILKD 554
+ ++ S +N SYQ +YAP G R H G +L + D
Sbjct: 299 KNGLNPFLTSSQHTSLNTNDSYQGASLPAYAPLSGYQGP-RSTH-------GTQLPVPSD 454
Query: 555 SSLSSSTYGSQGAAQG-SVSVLNLMEAVS 582
SL S GA G S SV+ + + S
Sbjct: 455 VSLVSDRQSKHGAKVGLSSSVVPVKDFTS 541
>TC230046 similar to UP|Q9LU79 (Q9LU79) Gb|AAF21150.1, partial (28%)
Length = 1798
Score = 37.4 bits (85), Expect = 0.047
Identities = 40/151 (26%), Positives = 59/151 (38%), Gaps = 10/151 (6%)
Frame = +3
Query: 1107 LQKTMGLVLKPRPGKQAKLGEKNKFYYDENLKRWVEEGAAPPAEETALPPPPTT------ 1160
+++ M ++ Q K +K K +EN ++ A T +PPPP
Sbjct: 1023 VKREMAMIWASVLSNQRKKKKKQKTKNNENQDPQYDDNADELTNNTTVPPPPPPPPPLPS 1202
Query: 1161 ---AAFQNGL-TEYNLQSALKTEGPPSKEGSDLKTSNPELTPGIPPIPPGTNHFSARGRV 1216
+ F+ GL + S PP S K+ P PP PP + GR
Sbjct: 1203 VFHSLFRKGLGKSKKIHSVSAPPPPPPPPPSKRKSQTPP-----PPEPPQRRN---SGRP 1358
Query: 1217 GIRSRYVDTFNQGGGNSANLFQSPSVPSAKP 1247
+ +R V TFN N+ N QSP +P P
Sbjct: 1359 PLPNRTV-TFNDETLNAGN--QSPLIPVPPP 1442
>TC217914 similar to GB|AAN18208.1|23308477|BT000642 At2g29670/T27A16.23
{Arabidopsis thaliana;} , partial (52%)
Length = 1970
Score = 37.0 bits (84), Expect = 0.061
Identities = 38/118 (32%), Positives = 50/118 (42%), Gaps = 7/118 (5%)
Frame = +3
Query: 24 DDVGNVNDEANDSDDVKAFSNLSIGGDDADVNASAFENSSGGGSGGEGKERKEEGDVKLD 83
D G++ DS +K FS +S G + A V GGG+GG GK R
Sbjct: 909 DQRGHIQSHNIDSSSIKTFS-VSNGKNTAYV---------GGGNGGGGKVRP-------- 1034
Query: 84 GGNVQEGSSSGCDGMMDRSDHGMESRNSSGSS-----ADKSNRRSSLDVKEK--DWNA 134
GN +G DG DRS HG + SS DK+ S + +E+ WNA
Sbjct: 1035 AGNGTDG-----DGRFDRSRHGTVFSDGGASSQVYKTGDKTESVSGQEEEEELNLWNA 1193
>TC227612 weakly similar to UP|Q8L7S0 (Q8L7S0) At1g09730/F21M12_12, partial
(19%)
Length = 1563
Score = 35.4 bits (80), Expect = 0.18
Identities = 33/93 (35%), Positives = 50/93 (53%), Gaps = 1/93 (1%)
Frame = +1
Query: 1264 SNEQTMEAIE-ENNQEDDLAYENPSTSYRNDWSFQSPKHASASTWQRCPSMGNFANHEAV 1322
S+E+T+ ++E EN+Q LAY+ P+T ND H ++ T+Q S+ NF EAV
Sbjct: 715 SSEETVLSLERENSQVGILAYDFPATYVSND-------HGASETFQVGFSV-NFV--EAV 864
Query: 1323 VSGSNSRSPHSRRTVSWGGSTDVTYSPTKMREI 1355
S S+SR+ VSW S T+ + +I
Sbjct: 865 ESHSHSRTSTG---VSWIPSNTATHEDQPLEKI 954
>TC230446 weakly similar to UP|Q6V9S3 (Q6V9S3) RPE1 protein, partial (12%)
Length = 803
Score = 34.3 bits (77), Expect = 0.39
Identities = 23/73 (31%), Positives = 36/73 (48%), Gaps = 4/73 (5%)
Frame = +2
Query: 1145 AAPPAEETALPPPPTTAAFQNGLTEYNLQSALKTEGPPSKEGSD-LKTSNPELTPGIPPI 1203
A PPA ++ PPP++ A + T + +S T P+ G+ +S+ TP PP
Sbjct: 23 APPPASSSSTTPPPSSTAGSS--TPPSSRSPASTSTSPASRGTTKTASSSTSATPSTPPS 196
Query: 1204 P---PGTNHFSAR 1213
P PG++ S R
Sbjct: 197 PTASPGSSSTSFR 235
>TC216506 similar to GB|AAL15354.1|16323240|AY057724 AT5g14920/F2G14_40
{Arabidopsis thaliana;} , partial (12%)
Length = 447
Score = 34.3 bits (77), Expect = 0.39
Identities = 21/75 (28%), Positives = 31/75 (41%)
Frame = +1
Query: 1131 FYYDENLKRWVEEGAAPPAEETALPPPPTTAAFQNGLTEYNLQSALKTEGPPSKEGSDLK 1190
F YDE+LK V APP + L PP + ++ G + PP + +K
Sbjct: 136 FSYDEDLKT-VVPAPAPPVKAPTLAPPVKSPSYPPGPVTTPTVPTPTVKVPPPPQSPVVK 312
Query: 1191 TSNPELTPGIPPIPP 1205
P + P +PP
Sbjct: 313 PPTPTVPPPTVKVPP 357
>TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;} , partial (46%)
Length = 902
Score = 33.9 bits (76), Expect = 0.52
Identities = 36/147 (24%), Positives = 56/147 (37%), Gaps = 5/147 (3%)
Frame = +2
Query: 9 VEDQDDEDFFDKLVEDDVGNVN-DEANDSDDVKAFSNLSIGGDDADVNASAFENSSGGGS 67
V+D +D++ ++ +DD G+ N DE +D+ D GGDD D
Sbjct: 320 VDDAEDDEDEEEDDDDDEGDDNDDEEDDAPD---------GGDDDD-------------- 430
Query: 68 GGEGKERKEEGDVKLDGGNVQEGSSSGCDGMMDRSDHGMESRNSSGSSADKSN----RRS 123
++ +EEGDV+ GG + + D D D E G D R
Sbjct: 431 ---DEDDEEEGDVQ-RGGEPDDDDNDSDDDSDDDEDEDEEDEEEQGEEEDLGTEYLIRPL 598
Query: 124 SLDVKEKDWNAFNVDSNGGAGSESYSD 150
+E+ + F + NG E D
Sbjct: 599 ETAEEEEASSDFEPEENGEEEEEDVDD 679
>TC215892 weakly similar to UP|Q43522 (Q43522) Tfm5 protein, partial (27%)
Length = 743
Score = 33.5 bits (75), Expect = 0.67
Identities = 34/116 (29%), Positives = 43/116 (36%), Gaps = 17/116 (14%)
Frame = +1
Query: 353 GHGNGHASS-------GTFSHNDNSLYRDYGQVGYYESQGVGSQAANNNWSGSY-GIN-- 402
G G+G+ + G +S ND Y YG Y G NN+SG Y G N
Sbjct: 1 GGGSGYRGNVSDGYGGGNYSRNDGGGY-GYGAGRYGSGGNYGDSGPGNNYSGGYSGSNSG 177
Query: 403 HQQD---LDRHTTDTATKSG---GSAYGGNQQFDHSFGSSNSV-NKNQQNASSSFG 451
H D + H + T GS G FGSS + +K N FG
Sbjct: 178 HFGDAGSVGNHESSTGFAGNGYDGSVVDGGVGAGSEFGSSGQLDSKTSSNGDEGFG 345
>TC212163 similar to UP|Q9LT26 (Q9LT26) Arabidopsis thaliana genomic DNA,
chromosome 3, P1 clone: MPN9, partial (30%)
Length = 739
Score = 33.5 bits (75), Expect = 0.67
Identities = 19/42 (45%), Positives = 23/42 (54%), Gaps = 2/42 (4%)
Frame = -3
Query: 1243 PSAKPVVAANAKFFIPTPAPSSN--EQTMEAIEENNQEDDLA 1282
P AKPV+AA A P P PS E + EE+ Q DD+A
Sbjct: 314 PHAKPVLAAAAVVLAPPPPPSPQLVEHDADEEEEDEQADDVA 189
>TC211625
Length = 293
Score = 33.5 bits (75), Expect = 0.67
Identities = 16/53 (30%), Positives = 25/53 (46%)
Frame = +1
Query: 55 NASAFENSSGGGSGGEGKERKEEGDVKLDGGNVQEGSSSGCDGMMDRSDHGME 107
N + +++ GGG G + KE ++ + DGG EGS D+ H E
Sbjct: 121 NQKSGKSNKGGGDGNKEKEDQKNSEPDADGGGSNEGSKDAPGEDSDKEGHSDE 279
>TC230134 weakly similar to UP|NUCL_HUMAN (P19338) Nucleolin (Protein C23),
partial (8%)
Length = 581
Score = 33.1 bits (74), Expect = 0.88
Identities = 23/85 (27%), Positives = 39/85 (45%)
Frame = +2
Query: 12 QDDEDFFDKLVEDDVGNVNDEANDSDDVKAFSNLSIGGDDADVNASAFENSSGGGSGGEG 71
++++D D +DD +VND + DD + + S GDD A + ++ G GGE
Sbjct: 254 EENKDASDTEDDDDDDDVNDGEDGDDDDEDDEDFS--GDDGGEEADSDDDPEANG-GGES 424
Query: 72 KERKEEGDVKLDGGNVQEGSSSGCD 96
+ E+ D D + +G D
Sbjct: 425 DDDDEDDDDDDDDNDEDDGDEDDDD 499
>TC228623 similar to UP|Q9SA21 (Q9SA21) F3O9.2 protein, partial (85%)
Length = 1082
Score = 32.7 bits (73), Expect = 1.1
Identities = 20/72 (27%), Positives = 30/72 (40%), Gaps = 6/72 (8%)
Frame = +2
Query: 48 GGDDADVNASAFENSSGGGSGGEGKERKEEGDVKL------DGGNVQEGSSSGCDGMMDR 101
GG A+ + + G G+G +E +EGD + DGG +E G G +
Sbjct: 290 GGSRAESGGATVGGAGGEGAGEVDEEGGQEGDTGVDPGAEGDGGGAEEAEGGGAGGG*EE 469
Query: 102 SDHGMESRNSSG 113
G E+ S G
Sbjct: 470 GGGG*EATESDG 505
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.309 0.128 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 58,323,499
Number of Sequences: 63676
Number of extensions: 835915
Number of successful extensions: 4716
Number of sequences better than 10.0: 107
Number of HSP's better than 10.0 without gapping: 4410
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4642
length of query: 1399
length of database: 12,639,632
effective HSP length: 109
effective length of query: 1290
effective length of database: 5,698,948
effective search space: 7351642920
effective search space used: 7351642920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC138087.1


