

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138017.8 - phase: 2 /pseudo
(600 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
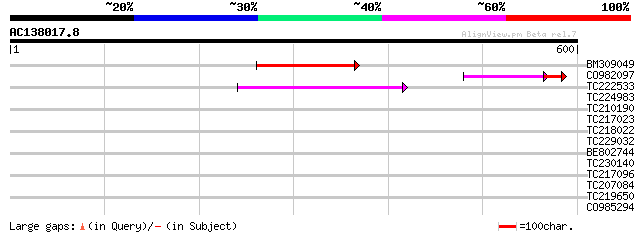
Score E
Sequences producing significant alignments: (bits) Value
BM309049 90 2e-18
CO982097 50 1e-08
TC222533 similar to UP|Q9SKV9 (Q9SKV9) F5J5.11, partial (11%) 55 1e-07
TC224983 similar to UP|Q8LUS1 (Q8LUS1) NADH dehydrogenase 1 (Fra... 30 3.1
TC210190 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, part... 30 4.0
TC217023 UP|Q9ZTZ2 (Q9ZTZ2) Late-embryogenesis abundant protein,... 29 5.2
TC218022 homologue to UP|Q9SL05 (Q9SL05) Expressed protein (At2g... 29 5.2
TC229032 29 5.2
BE802744 29 6.8
TC230140 similar to UP|O03296 (O03296) Cytochrome b (Fragment), ... 29 6.8
TC217096 homologue to UP|Q8W1S2 (Q8W1S2) ABC transporter-like pr... 29 6.8
TC207084 UP|Q9XES9 (Q9XES9) Seed maturation protein PM29, complete 29 6.8
TC219650 similar to GB|AAN18074.1|23308209|BT000505 At2g30580/T6... 28 8.9
CO985294 28 8.9
>BM309049
Length = 430
Score = 90.1 bits (222), Expect = 2e-18
Identities = 57/109 (52%), Positives = 71/109 (64%)
Frame = +1
Query: 262 SSLACIGLLVQLIALT*CFKTLENYLKLKRQFHMPQMLPSMYIIIVIHCI**GNLLMEER 321
S L+ IGL V AL *C+ TL N K R H+ Q LPS++ II+I I** N+ +E+
Sbjct: 4 SFLSYIGLHVLHSALI*CWMTL*N*RK*VRLCHLLQKLPSIFTIIIILYI**ENIQVEKI 183
Query: 322 YFVLLQLALPLISLLCRVFCLRKMHLEPW*HLKNGQLLLMQKMPRPNNL 370
Y V LQL+LPLISL C+V+ L KMH EPW*HL+ GQ LM K R +
Sbjct: 184 YLVQLQLSLPLISLPCKVYWLIKMH*EPW*HLEIGQAQLMPKNLRQKKI 330
Score = 33.1 bits (74), Expect = 0.36
Identities = 21/47 (44%), Positives = 27/47 (56%)
Frame = +2
Query: 357 QLLLMQKMPRPNNLWNKS*TLTFGLLVLT**NSQNHLYVCCVSWTVK 403
+L L Q++ LWNKS*T FG VL *+SQ+H + TVK
Sbjct: 290 KLSLCQRI*GKKKLWNKS*TPGFGKNVLIL*SSQSHWFTSYKLLTVK 430
>CO982097
Length = 805
Score = 50.4 bits (119), Expect(2) = 1e-08
Identities = 38/90 (42%), Positives = 45/90 (49%)
Frame = -3
Query: 481 MSLKSMLVIIMSCEQS*I*RQVYLEILRATLEGNLL*KLEIHHFQMNGANFTGVKHHICK 540
MSLK ML+ I+ C + YL++ L G LL I QMNG N V H K
Sbjct: 797 MSLKGMLMEILICNLIWQVK*EYLKMPS*ILGG*LLHVNGIR*CQMNGGNLMDVVHQTYK 618
Query: 541 NWRFGF*VKLVALLVARETGVCLSIFTKKK 570
+W F F* KL L V GV LSIF + K
Sbjct: 617 SWLFLF*AKLAVLQVVNGIGVYLSIFIQIK 528
Score = 26.9 bits (58), Expect(2) = 1e-08
Identities = 16/24 (66%), Positives = 19/24 (78%)
Frame = -1
Query: 566 FTKKKKK*VGASKA*RSSLCSLQL 589
++ K K+*V ASKA* S LCSLQL
Sbjct: 544 YSFK*KE*VRASKA**SCLCSLQL 473
>TC222533 similar to UP|Q9SKV9 (Q9SKV9) F5J5.11, partial (11%)
Length = 596
Score = 54.7 bits (130), Expect = 1e-07
Identities = 57/180 (31%), Positives = 79/180 (43%)
Frame = +1
Query: 242 FR**QIMLQTMLLLVGYWRKSSLACIGLLVQLIALT*CFKTLENYLKLKRQFHMPQMLPS 301
F+ * IM T+ V WR+ IGLLVQLI L *C K L ++ RQ +
Sbjct: 1 FKL*PIMGATIF*RVSCWRRKGNIFIGLLVQLIVLI*CLKILGSFP**GRQLEGQLI*LG 180
Query: 302 MYIIIVIHCI**GNLLMEERYFVLLQLALPLISLLCRVFCLRKMHLEPW*HLKNGQLLLM 361
+ + I++ + * L + +L L LPL+ + F RK LE L NG
Sbjct: 181 LSMSILVP*VC*EILQTRGNW*DMLLLDLPLLI*PWKGFTKRKPILERCLLLMNGP*TSY 360
Query: 362 QKMPRPNNLWNKS*TLTFGLLVLT**NSQNHLYVCCVSWTVKINLLWVFFTEICIRLERR 421
+ R L L FG++ T S HL C V W VK N W F + R +++
Sbjct: 361 LRSLREKKLQR*CSCLLFGIVWFTLLKSWLHL*KCFVLWMVKGNQPWTIFMKQWTRQKKQ 540
>TC224983 similar to UP|Q8LUS1 (Q8LUS1) NADH dehydrogenase 1 (Fragment),
partial (37%)
Length = 1412
Score = 30.0 bits (66), Expect = 3.1
Identities = 14/33 (42%), Positives = 20/33 (60%)
Frame = +3
Query: 55 KSKHKEDNLGQWLLHLKRERTMEALIITFCLEQ 87
KSKH++ GQW+ +KR+R + II EQ
Sbjct: 78 KSKHQQGPSGQWVDVMKRKRAIHENIINLVREQ 176
>TC210190 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, partial (65%)
Length = 866
Score = 29.6 bits (65), Expect = 4.0
Identities = 15/30 (50%), Positives = 18/30 (60%)
Frame = -2
Query: 317 LMEERYFVLLQLALPLISLLCRVFCLRKMH 346
L+ Y+VLL L P SLLC VF L+ H
Sbjct: 538 LLHTVYYVLLWLPSPSRSLLCAVFSLKGYH 449
>TC217023 UP|Q9ZTZ2 (Q9ZTZ2) Late-embryogenesis abundant protein, complete
Length = 677
Score = 29.3 bits (64), Expect = 5.2
Identities = 14/26 (53%), Positives = 17/26 (64%)
Frame = -1
Query: 328 LALPLISLLCRVFCLRKMHLEPW*HL 353
L LPL+SL R+F L + L PW HL
Sbjct: 155 LFLPLLSLPSRLFWLYRCRLVPWQHL 78
>TC218022 homologue to UP|Q9SL05 (Q9SL05) Expressed protein
(At2g05620/T20G20.3), partial (57%)
Length = 750
Score = 29.3 bits (64), Expect = 5.2
Identities = 11/32 (34%), Positives = 20/32 (62%)
Frame = +3
Query: 41 QGSRKEKLNMQ*VMKSKHKEDNLGQWLLHLKR 72
Q S +K N + +K K++++ QWLLH ++
Sbjct: 78 QASSSKKSNSTFIFTTKKKKESINQWLLHFQQ 173
>TC229032
Length = 668
Score = 29.3 bits (64), Expect = 5.2
Identities = 14/32 (43%), Positives = 21/32 (64%), Gaps = 1/32 (3%)
Frame = -1
Query: 55 KSKHKEDNLGQWL-LHLKRERTMEALIITFCL 85
+SK++ N QW+ LHLK+ +TME+ CL
Sbjct: 266 ESKYQNGNKNQWIHLHLKQVKTMESKPNRICL 171
>BE802744
Length = 450
Score = 28.9 bits (63), Expect = 6.8
Identities = 11/28 (39%), Positives = 20/28 (71%)
Frame = +1
Query: 124 MQQIHHIFSLRSMLFVAWEPDIKLLLYM 151
M++ HH+F+L S L++ W +KLL+ +
Sbjct: 337 MRRGHHLFNLESRLWIEWAMWMKLLMIL 420
>TC230140 similar to UP|O03296 (O03296) Cytochrome b (Fragment), partial (5%)
Length = 1462
Score = 28.9 bits (63), Expect = 6.8
Identities = 17/52 (32%), Positives = 29/52 (55%)
Frame = -1
Query: 20 SKHATKLLKKSTLR*NKIVKRQGSRKEKLNMQ*VMKSKHKEDNLGQWLLHLK 71
+ +A + KK+ LR N + + ++ K + + + K KE LG+WLLH K
Sbjct: 1018 NNNAR*MAKKTRLRNNDRSENKRFKRRKTDRK---RKKTKEIQLGRWLLHRK 872
>TC217096 homologue to UP|Q8W1S2 (Q8W1S2) ABC transporter-like protein,
complete
Length = 2478
Score = 28.9 bits (63), Expect = 6.8
Identities = 11/28 (39%), Positives = 20/28 (71%)
Frame = +1
Query: 124 MQQIHHIFSLRSMLFVAWEPDIKLLLYM 151
M++ HH+F+L S L++ W +KLL+ +
Sbjct: 1489 MRRGHHLFNLESRLWIEWAMWMKLLMIL 1572
>TC207084 UP|Q9XES9 (Q9XES9) Seed maturation protein PM29, complete
Length = 637
Score = 28.9 bits (63), Expect = 6.8
Identities = 13/25 (52%), Positives = 16/25 (64%)
Frame = -3
Query: 328 LALPLISLLCRVFCLRKMHLEPW*H 352
L LPL+SL R+F L + L PW H
Sbjct: 167 LFLPLLSLSSRLFLLYRCRLAPWQH 93
>TC219650 similar to GB|AAN18074.1|23308209|BT000505 At2g30580/T6B20.7
{Arabidopsis thaliana;} , partial (39%)
Length = 1218
Score = 28.5 bits (62), Expect = 8.9
Identities = 16/44 (36%), Positives = 23/44 (51%)
Frame = -2
Query: 139 VAWEPDIKLLLYMIYVVLC*ISGLMKQRKR*RNTVRFGRLLVVL 182
V W P I L M++ VLC GLM R +R GR+ +++
Sbjct: 368 VQWCPKINLPFCMLHTVLCIFMGLMLLR-----MLRHGRIHIII 252
>CO985294
Length = 836
Score = 28.5 bits (62), Expect = 8.9
Identities = 19/67 (28%), Positives = 30/67 (44%), Gaps = 7/67 (10%)
Frame = +3
Query: 92 NLL*KVCCKPRKL*KSVILHLQNGSLLHLFPSMQQIHHI-------FSLRSMLFVAWEPD 144
NL CC R +V++ + S FP+ Q+HH+ F + L++ E D
Sbjct: 222 NLKKIACCDGRVFLPAVVIKHLDSSTWMFFPASHQVHHLD*Q*LVQFWMELDLYLKCEED 401
Query: 145 IKLLLYM 151
+ LL M
Sbjct: 402 TQKLLPM 422
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.360 0.161 0.561
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,768,559
Number of Sequences: 63676
Number of extensions: 466219
Number of successful extensions: 5634
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 3106
Number of HSP's successfully gapped in prelim test: 242
Number of HSP's that attempted gapping in prelim test: 2447
Number of HSP's gapped (non-prelim): 3491
length of query: 600
length of database: 12,639,632
effective HSP length: 103
effective length of query: 497
effective length of database: 6,081,004
effective search space: 3022258988
effective search space used: 3022258988
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC138017.8


