

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138012.9 - phase: 0
(672 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
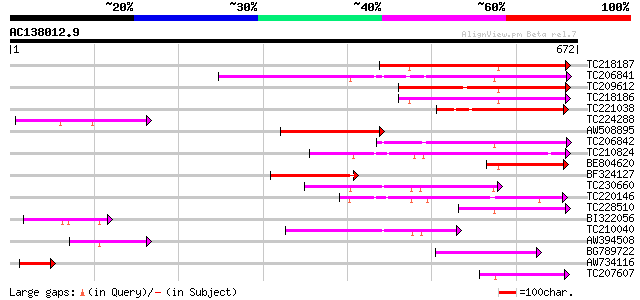
Score E
Sequences producing significant alignments: (bits) Value
TC218187 similar to UP|Q9FGH9 (Q9FGH9) Leucine zipper protein (A... 181 7e-46
TC206841 similar to UP|Q9FHC6 (Q9FHC6) Emb|CAB83315.1, partial (... 171 7e-43
TC209612 166 4e-41
TC218186 weakly similar to UP|Q9FGH9 (Q9FGH9) Leucine zipper pro... 159 3e-39
TC221038 similar to UP|Q40156 (Q40156) L.esculentum protein with... 156 3e-38
TC224288 135 8e-32
AW508895 weakly similar to GP|10177020|dbj leucine zipper protei... 132 4e-31
TC206842 similar to UP|LORI_MOUSE (P18165) Loricrin, partial (4%) 110 2e-24
TC210824 weakly similar to UP|Q9FNR3 (Q9FNR3) Leucine zipper pro... 107 1e-23
BE804620 102 4e-22
BF324127 similar to GP|10177020|db leucine zipper protein {Arabi... 98 1e-20
TC230660 weakly similar to UP|Q9FNR3 (Q9FNR3) Leucine zipper pro... 97 2e-20
TC220146 87 2e-17
TC228510 similar to UP|Q9FK34 (Q9FK34) Gb|AAD25781.1, partial (18%) 77 2e-14
BI322056 76 5e-14
TC210040 70 2e-12
AW394508 70 2e-12
BG789722 67 3e-11
AW734116 60 2e-09
TC207607 similar to UP|Q9LJH9 (Q9LJH9) Gb|AAD25781.1, partial (20%) 58 2e-08
>TC218187 similar to UP|Q9FGH9 (Q9FGH9) Leucine zipper protein
(AT5g58430/mqj2_20), partial (33%)
Length = 1098
Score = 181 bits (460), Expect = 7e-46
Identities = 104/237 (43%), Positives = 147/237 (61%), Gaps = 11/237 (4%)
Frame = +2
Query: 439 VVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILE-----EYPEVHNEVEASSFFLKQMEQ 493
+V + G H +T VM+Y+ +ACR R+ LEQ+ E EYP++ + V +SS QM+
Sbjct: 5 LVPNGGGLHPITRYVMNYLRAACRSRQSLEQVFEDYGLKEYPKLDDRVPSSSSLSVQMDW 184
Query: 494 IMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSRLETIFGNDWFQNNKAKIQQ 553
IM +L+ L KS+ KD AL +IF++NN +I K S L T+ G DW + + AK++Q
Sbjct: 185 IMELLESNLEAKSKIYKDPALCYIFLMNNGRYIVQKTKDSELGTLLGEDWIRKHAAKVRQ 364
Query: 554 NLDLYKRSAWDEVMDFLKLDNNESI----TKELLKEKIHLFNNRFEAICRVQSAWFIYGS 609
Y+RS+W++++ LKLD+N S+ + +KEK+ FN FE IC+ QS+WF++
Sbjct: 365 FHVHYQRSSWNKLLGILKLDSNGSMPHINLAKSMKEKLKSFNTVFEEICKEQSSWFVFDE 544
Query: 610 QLRGEIISSVGNILLPAYGIFVGRLHGI--LGNQAYKYIKYGMIEIQDLLNHLFLGN 664
QLR EI S+ ILLPAYG FV R + LG A KYIKYG EIQ LN LF G+
Sbjct: 545 QLREEIRISLEKILLPAYGNFVARFQSVPELGKHADKYIKYGTEEIQARLNGLFQGS 715
>TC206841 similar to UP|Q9FHC6 (Q9FHC6) Emb|CAB83315.1, partial (50%)
Length = 1568
Score = 171 bits (434), Expect = 7e-43
Identities = 118/442 (26%), Positives = 217/442 (48%), Gaps = 23/442 (5%)
Frame = +1
Query: 248 LPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKINTERERE 307
+P L L+ + + M+ AG++++ Y R +L++ L K+ V +K + ++ +
Sbjct: 22 IPPRILPLLNNLTQQMVQAGHQQQLLKTYRDTRSKVLEESL-QKLGVEKLSKDDVQKLQW 198
Query: 308 RYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQLLNFAD 367
L+ W+ IA +LF E+K CD +F GF S + CF E+ + LL+F +
Sbjct: 199 EVLEAKIGNWIHFMRIAVKLLFAAERKVCDQIFEGFDSLSDQCFAEVTTNSISMLLSFGE 378
Query: 368 VIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPN---SLVNEAI-AVRNRLGDASRVLF 423
IA S +LF ++D++ L + + E LF + + EA+ + +L ++ F
Sbjct: 379 AIAKSKRSPEKLFVLLDMYEILQEIHAEIEILFKGRACTKIREAVMGLTKQLAQTAQETF 558
Query: 424 MKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEEYPEVHNEVEA 483
+ + A K V+ G H +T V++YV R L Q+ + + E ++
Sbjct: 559 GDFEEAVEK-DATKTAVTD-GTVHPLTSYVINYVKFLFDYRSTLHQL---FQGIEGEGDS 723
Query: 484 SSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSRLETIFGNDW 543
S M +I++ LQ L KS++ +D AL H+F++NN +I + S + + G+DW
Sbjct: 724 SQLASVTM-RILQALQTNLDGKSKHYRDPALTHLFLMNNIHYIVRSVRRSEAKDLLGDDW 900
Query: 544 FQNNKAKIQQNLDLYKRSAWDEVMDFLKLD-----------------NNESITKELLKEK 586
Q ++ +QQ+ + YKR+AW +++ L + + ++ ++K++
Sbjct: 901 IQRHRKIVQQHANQYKRNAWAKILQSLSIQGLISSSGGGSSNAGGDAGSSGASRTMVKDR 1080
Query: 587 IHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGILGN--QAYK 644
FN FE + + QS W + ++LR +I +V +LLPAY FV R ++ N +
Sbjct: 1081FKTFNTMFEELHQKQSQWTVPDAELRESLILAVAEVLLPAYRSFVKRFGPLVENVKSTQR 1260
Query: 645 YIKYGMIEIQDLLNHLFLGNKM 666
YIKY +++ +L F G M
Sbjct: 1261YIKYTAEDLERILGEFFEGKSM 1326
>TC209612
Length = 916
Score = 166 bits (419), Expect = 4e-41
Identities = 89/207 (42%), Positives = 132/207 (62%), Gaps = 4/207 (1%)
Frame = +3
Query: 462 RKRRKLEQILEEYPEVHNEVEASSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLN 521
R ++ LEQILE+YP+ NEV S+ Q++Q+++ L+ +L+ S+N ALR+ F++N
Sbjct: 12 RAQKILEQILEQYPKFANEVAKSNSVSDQIDQVIKRLETELVTVSKNYDKPALRYFFLMN 191
Query: 522 NRSHIEAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDNNESI--- 578
N +E RL F + K+QQNL+LY+ S+W+ V++FLKL+NNE +
Sbjct: 192 NWRCVELEAIKLRLNL----GCFHKDTTKVQQNLELYQSSSWNMVLNFLKLENNELVEPN 359
Query: 579 -TKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGI 637
E LK ++LFN F+ IC QS W + QL +II S+ +ILLPAYG F+ +L +
Sbjct: 360 ANAESLKGSLNLFNMHFKDICSTQSRWLAFDKQLSEKIIMSLQHILLPAYGNFIEKLQDV 539
Query: 638 LGNQAYKYIKYGMIEIQDLLNHLFLGN 664
LG A +YIKYG+ +I+D LNHLFLG+
Sbjct: 540 LGIHASEYIKYGLFDIKDQLNHLFLGS 620
>TC218186 weakly similar to UP|Q9FGH9 (Q9FGH9) Leucine zipper protein
(AT5g58430/mqj2_20), partial (28%)
Length = 738
Score = 159 bits (403), Expect = 3e-39
Identities = 93/215 (43%), Positives = 127/215 (58%), Gaps = 12/215 (5%)
Frame = +1
Query: 462 RKRRKLEQILE-----EYPEVHNEVEASSFFLKQMEQIMRMLQRKLIVKSENCKDRALRH 516
R R+ LEQ+ E EY ++ + V +SS QM+ IM +L+ L KS KD ALR+
Sbjct: 4 RSRQSLEQVFEDYGLKEYTKLEDRVPSSSSLSVQMDWIMELLESNLEAKSRIYKDPALRY 183
Query: 517 IFMLNNRSHIEAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDNNE 576
+F++NN +I K S L T+ G+DW + + AK++Q Y+R +W +V+ LKLD+N
Sbjct: 184 VFLMNNGRYIVQKTKDSELGTLLGDDWIRKHAAKVRQFHVHYQRCSWTKVLGILKLDSNG 363
Query: 577 SI-----TKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFV 631
S + +KE + LFN FE CR S+WF++ QLR EI S+ ILLPAYG FV
Sbjct: 364 SSLPPNGLAKSMKETLKLFNTVFEETCREHSSWFVFDEQLREEIRISLEKILLPAYGNFV 543
Query: 632 GRLHGI--LGNQAYKYIKYGMIEIQDLLNHLFLGN 664
R + LG A KYIKYG EIQ LN LF G+
Sbjct: 544 ARFESVAELGKNADKYIKYGTEEIQATLNGLFQGS 648
>TC221038 similar to UP|Q40156 (Q40156) L.esculentum protein with leucine
zipper, partial (5%)
Length = 519
Score = 156 bits (394), Expect = 3e-38
Identities = 75/156 (48%), Positives = 108/156 (69%)
Frame = +3
Query: 507 ENCKDRALRHIFMLNNRSHIEAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEV 566
+N D AL H+FM+NN ++ K+ IFG DW+ K+KI QN++LY+RS+ D++
Sbjct: 3 KNYTDPALGHVFMINNLMLLQ-YEKYIYRVVIFGEDWY---KSKINQNIELYQRSSLDKI 170
Query: 567 MDFLKLDNNESITKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPA 626
+DFL LD+NE + E +K+K+ LFN F IC+ QS W I+ QL+ ++I S+ N LLPA
Sbjct: 171 LDFLNLDSNELLLAESMKKKLKLFNQHFNEICKAQSEWLIFDEQLKEQMIKSIENKLLPA 350
Query: 627 YGIFVGRLHGILGNQAYKYIKYGMIEIQDLLNHLFL 662
YG F+GR+H +LG AY +I+YG+ IQDLL+ LFL
Sbjct: 351 YGTFLGRIHDVLGKDAYDFIRYGIQNIQDLLSGLFL 458
>TC224288
Length = 736
Score = 135 bits (339), Expect = 8e-32
Identities = 75/171 (43%), Positives = 102/171 (58%), Gaps = 10/171 (5%)
Frame = +3
Query: 8 IRMCLLQTKVWRFVGFASAAVGLLCYALSTSFNHLFGNWNLLKIFLYTLFS-----FIMI 62
I L+ +WR GF S+ VG CYALS SF+++FG+WN LKIF+YT+ S F+
Sbjct: 27 ISKLLIHPLLWRLTGFVSSIVGFSCYALSPSFHNMFGHWNALKIFVYTVVSSLLSIFMFF 206
Query: 63 LYANIWKNSRSLRFKAHTAFLVLTITSVYSFFFD-----KVVNGKPDAYSLISCAAFAIM 117
+ W + RS KA F+VLT+TS++S F D KV N +L S AFA+M
Sbjct: 207 IKRCSWGHGRSFLLKAQVGFVVLTLTSLWSVFEDRSEEGKVENAHGKMMNLTSSGAFALM 386
Query: 118 SMSLSRQSQCGIEVDLLYFFLGCLIMQLMKIKLQLVIVGAGFSYSLIIIRS 168
+MSLSRQ Q G EV + F +GC ++ +MK+ +L V A F Y L+ IRS
Sbjct: 387 AMSLSRQLQLGFEVGVFNFLVGCFLVTVMKMSFKLAPVAALFCYLLVNIRS 539
>AW508895 weakly similar to GP|10177020|dbj leucine zipper protein
{Arabidopsis thaliana}, partial (12%)
Length = 376
Score = 132 bits (333), Expect = 4e-31
Identities = 60/123 (48%), Positives = 88/123 (70%)
Frame = +3
Query: 322 DIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQLLNFADVIAYGSLSKWRLFK 381
++A +LFP E++ CD +FS FSS + CF E+C+ A QLLNFA+ +A GS S+WRL K
Sbjct: 6 NVAHRILFPSERRLCDCIFSRFSSVAALCFNEVCRGALIQLLNFAEAVASGSPSEWRLSK 185
Query: 382 MVDIFVKLNNLVPKFESLFPNSLVNEAIAVRNRLGDASRVLFMKMHNFIFRVPAAKQVVS 441
++D+F L +L+P+F+SLFP S+V E + V ++LG+ASRV+FM M N IF +P K +
Sbjct: 186 ILDMFETLRDLIPEFQSLFPESMVKEVMKVHDKLGEASRVIFMNMENVIFHIPETKVIAP 365
Query: 442 SYG 444
+ G
Sbjct: 366 ADG 374
>TC206842 similar to UP|LORI_MOUSE (P18165) Loricrin, partial (4%)
Length = 1122
Score = 110 bits (276), Expect = 2e-24
Identities = 72/253 (28%), Positives = 126/253 (49%), Gaps = 21/253 (8%)
Frame = +2
Query: 435 AAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEEYPEVHNEVEASSFFLKQMEQI 494
A K V+ G H +T V++YV + L+Q+ +E+ + + +S ++ I
Sbjct: 38 ATKTAVTD-GTVHPLTSYVINYVKFLFDYQSTLKQLFQEFEGGDDSSQLASVTVR----I 202
Query: 495 MRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSRLETIFGNDWFQNNKAKIQQN 554
M+ LQ L KS+ KD AL H+F++NN +I + S + + G+DW Q ++ +QQ+
Sbjct: 203 MQALQTNLDGKSKQYKDLALTHLFLMNNIHYIVRSVRRSEAKDLLGDDWVQRHRRIVQQH 382
Query: 555 LDLYKRSAWDEVMDFLKLD-------------------NNESITKELLKEKIHLFNNRFE 595
+ YKR+AW +++ L + ++ ++ ++K++ FN FE
Sbjct: 383 ANQYKRNAWAKILQCLSIQGLTSSGGGSGTAGGDSGTGSSSGASRAIVKDRFKAFNIMFE 562
Query: 596 AICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGIL--GNQAYKYIKYGMIEI 653
+ + QS W + S+LR + +V +LLPAY FV R ++ G KYIKY ++
Sbjct: 563 ELHQKQSQWTVPDSELRESLRLAVAEVLLPAYRSFVKRFGPLVESGKNPQKYIKYSAEDL 742
Query: 654 QDLLNHLFLGNKM 666
+L F G M
Sbjct: 743 DRMLGEFFEG*NM 781
>TC210824 weakly similar to UP|Q9FNR3 (Q9FNR3) Leucine zipper protein-like
(AT5g61010/maf19_10), partial (13%)
Length = 1027
Score = 107 bits (268), Expect = 1e-23
Identities = 86/321 (26%), Positives = 150/321 (45%), Gaps = 12/321 (3%)
Frame = +3
Query: 356 QEATFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPNSLVN----EAIAV 411
Q LLNF + +A G + ++F+++D++ L L + LF + + E +
Sbjct: 6 QSFMLHLLNFGEAVAMGMHTPEKMFRLLDMYEVLEKLDVDVDVLFFEEVGSFVRGELHKL 185
Query: 412 RNRLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQIL 471
GD + + N I +K G HH +T VM+Y+ + L +L
Sbjct: 186 LRSFGDTIKSTLLAFRNAIAS-NHSKTPFPQGGVHH-VTKYVMNYIMALVEYGDTLNLLL 359
Query: 472 EEYPEVH---NEVEASSFFLK----QMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRS 524
+ + N+ + L Q I L+ L KS+ KD AL+HIFM+NN
Sbjct: 360 VDDTSIDPAGNKDDTPCLSLCPVACQFRSIXATLESNLSNKSKLYKDEALQHIFMMNNIH 539
Query: 525 HIEAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDN-NESITKELL 583
++ K S L FG+ W + + A Q++ Y+R +W V+ LK + + +++ L
Sbjct: 540 YMVQKVKCSDLSHFFGDCWLRQHIAMYQRDARCYERISWGSVLSMLKEGSVSNCVSQRTL 719
Query: 584 KEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGILGNQAY 643
+++ F+ F + R+Q+ WFI +LR ++ SV L+ AY ++GR + A
Sbjct: 720 EKRCKEFSTAFGEVYRIQTGWFILDPRLREDLQISVSQKLVLAYRTYIGRNSSSI---AE 890
Query: 644 KYIKYGMIEIQDLLNHLFLGN 664
KY+KY ++Q + LF G+
Sbjct: 891 KYVKYTEDDLQSYILDLFQGS 953
>BE804620
Length = 400
Score = 102 bits (255), Expect = 4e-22
Identities = 49/100 (49%), Positives = 68/100 (68%), Gaps = 3/100 (3%)
Frame = +1
Query: 566 VMDFLKLDNNES---ITKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNI 622
+++ LKLD NES + +L+K K+ FN + IC Q+ W + +LR +II S+ NI
Sbjct: 1 ILNILKLDINESEPNVAAKLMKNKLCSFNEHLDDICNTQATWSVLNEELREQIIKSIENI 180
Query: 623 LLPAYGIFVGRLHGILGNQAYKYIKYGMIEIQDLLNHLFL 662
LLPAYG F+ RL LGN A++YI+YGM +IQD LN+LFL
Sbjct: 181 LLPAYGNFIARLQDFLGNHAFEYIEYGMFDIQDRLNNLFL 300
>BF324127 similar to GP|10177020|db leucine zipper protein {Arabidopsis
thaliana}, partial (17%)
Length = 334
Score = 98.2 bits (243), Expect = 1e-20
Identities = 45/104 (43%), Positives = 72/104 (68%)
Frame = +1
Query: 310 LDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQLLNFADVI 369
L+ ++W+ AS++A +LFP E++ CD VF GF+SA F+E+C+ + QLLNFAD +
Sbjct: 28 LEDEIEKWIKASNVALKILFPSERRLCDRVFFGFASAADFSFMEVCRGSAIQLLNFADAV 207
Query: 370 AYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPNSLVNEAIAVRN 413
A GS S RLF+++D+F L +L P+FE+LF + ++++RN
Sbjct: 208 AIGSRSPERLFRILDVFETLRDLFPEFEALFSDQF---SVSLRN 330
>TC230660 weakly similar to UP|Q9FNR3 (Q9FNR3) Leucine zipper protein-like
(AT5g61010/maf19_10), partial (12%)
Length = 903
Score = 97.1 bits (240), Expect = 2e-20
Identities = 70/262 (26%), Positives = 127/262 (47%), Gaps = 27/262 (10%)
Frame = +3
Query: 350 CFIEICQEATFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPN----SLV 405
CF++ + + QLLNF + ++ G +LF+++DI+ L +L+P ++L+ + S+
Sbjct: 123 CFVDASKASMLQLLNFGEAMSIGPHQPEKLFRVLDIYEVLQDLMPDIDALYSDEVGSSVK 302
Query: 406 NEAIAVRNRLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRR 465
E V RLGD RV F++ N I ++ V G H +T VM+Y+ +
Sbjct: 303 IECHEVLKRLGDCVRVTFLEFENAIATNVSSTPFVG--GGIHPLTKYVMNYLRTLTDYSD 476
Query: 466 KLEQILEEY--------PEVHNEVEASS----------FFLKQMEQIMRMLQRKLIVKSE 507
L +L++ P++ E S I +L+ L KS+
Sbjct: 477 ILNLLLKDQDEDAISLSPDMSPGTEEDSRSQGSPGRVSSMALHFRSIASILESNLEEKSK 656
Query: 508 NCKDRALRHIFMLNNRSHIEAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVM 567
K+ +L+H+F++NN ++ K S L I G++W + K QQ+ Y+R++W+ ++
Sbjct: 657 LYKEVSLQHLFLMNNLHYMAEKVKGSELRLIHGDEWIRKCNWKFQQHAMKYERASWNPIL 836
Query: 568 DFLK-----LDNNESITKELLK 584
+ LK + S++K LLK
Sbjct: 837 NLLKDEGIHVPGTNSVSKSLLK 902
>TC220146
Length = 1013
Score = 87.0 bits (214), Expect = 2e-17
Identities = 72/294 (24%), Positives = 141/294 (47%), Gaps = 24/294 (8%)
Frame = +3
Query: 392 LVPKFESLFP---NSLVNEAIAVR-NRLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHH 447
L+P+ ES+F NS V V RL +++++L + + I + +K V+ G H
Sbjct: 6 LLPEIESIFSSDYNSGVRSQFLVSLQRLTESAQILLAEFESTIQK-GTSKPAVNGGGVH- 179
Query: 448 QMTIQVMSYVSSACRKRRKLEQILEEY---PEVHNEVEASSFFLKQMEQ----------- 493
+TIQ M+Y+S L I P+ + + S + + +
Sbjct: 180 SLTIQTMNYLSVLADYLNVLSDIFPRDWLPPQKSSSLPESYLYSPESDYSASTPALTARF 359
Query: 494 --IMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSRLETIFGNDWFQNNKAKI 551
++ +L KL K+++CKD +L ++F+ NN ++ A + S L+ + G+DW ++AK
Sbjct: 360 AWLILVLLCKLDGKAKHCKDVSLSYLFLANNLWYVVARVRSSNLQYVLGDDWILKHEAKA 539
Query: 552 QQNLDLYKRSAWDEVMDFLKLDNNESITKELLKEKIHLFNNRFEAICRVQSAWFIYGSQL 611
++ + Y++ AW EV+ L E+ + FN +FE R Q+++ + +L
Sbjct: 540 KRFVSNYEKVAWGEVVSSL----TENPAAAEARAVFENFNRKFEEAYRKQNSFVVADREL 707
Query: 612 RGEIISSVGNILLPA----YGIFVGRLHGILGNQAYKYIKYGMIEIQDLLNHLF 661
R EI S+ ++P Y + + ++ + A +Y+ + +I++ L +LF
Sbjct: 708 RDEIKGSIARSIVPRYREWYNVLLAKVGSVRDLTATEYVTFTPEDIENYLVNLF 869
>TC228510 similar to UP|Q9FK34 (Q9FK34) Gb|AAD25781.1, partial (18%)
Length = 795
Score = 77.0 bits (188), Expect = 2e-14
Identities = 47/137 (34%), Positives = 74/137 (53%), Gaps = 5/137 (3%)
Frame = +1
Query: 533 SRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLD----NNESITKELLKEKIH 588
S L +G +W + + +I+Q Y R++W + + LK + ++ + +K LKE+
Sbjct: 4 SDLXGXWGXNWIRKRRGQIRQYATGYLRASWSKALSCLKDEGIGGSSNNASKMALKERFK 183
Query: 589 LFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLH-GILGNQAYKYIK 647
FN FE I RVQ+AW + QLR E+ S+ ++PAY FVGR + G KYIK
Sbjct: 184 SFNACFEEIYRVQTAWKVPDDQLREELRISISEKVIPAYRSFVGRFRIQLEGRHVGKYIK 363
Query: 648 YGMIEIQDLLNHLFLGN 664
Y +++ L LF G+
Sbjct: 364 YTPEDLETYLLDLFEGS 414
>BI322056
Length = 437
Score = 75.9 bits (185), Expect = 5e-14
Identities = 43/117 (36%), Positives = 68/117 (57%), Gaps = 11/117 (9%)
Frame = +1
Query: 17 VWRFVGFASAAVGLLCYALSTSFNHLFGNWNLLKIFLYTLFSFIM---ILYANIW---KN 70
+W+ GF S+ VG CYALS SF L W+ KI Y++ S ++ +L+ W ++
Sbjct: 85 LWKLTGFGSSIVGFSCYALSPSFLCLVKEWSPKKIVAYSVVSSLLSISMLFVKKWSLGRH 264
Query: 71 SRSLRFKAHTAFLVLTITSVYSFFFDKVVNGKPD-----AYSLISCAAFAIMSMSLS 122
+SL K H F+VL +T+++SF+ D+ GK + +L S AFA+M++SLS
Sbjct: 265 GKSLLLKGHVVFVVLALTNLWSFWEDRCQQGKVENRFRKIMNLASIGAFALMALSLS 435
>TC210040
Length = 684
Score = 70.5 bits (171), Expect = 2e-12
Identities = 56/222 (25%), Positives = 105/222 (47%), Gaps = 13/222 (5%)
Frame = +3
Query: 327 VLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQLLNFADVIAYGSLSKWRLFKMVDIF 386
VL E++ CD +F CF E + QLLNF + IA S +LF+++D++
Sbjct: 12 VLLSGEKRLCDGLFGDLDDLKEICFNETAKGCVMQLLNFGEAIAICKRSPEKLFRILDMY 191
Query: 387 VKLNNLVPKFESLFPNS-LVNEAIAVRNRLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQ 445
L + +P +++ + ++ EA V + LG+A++ F + N I + K V++ G
Sbjct: 192 EALRDAMPDLQAMVSDEFVIGEANGVLSGLGEAAKGTFAEFENCIRNETSKKPVIT--GD 365
Query: 446 HHQMTIQVMSYVSSACRKRRKLEQILEEYPE----VHNEVEASSF--------FLKQMEQ 493
H + VM+Y+ ++ +LE E N++ F +++
Sbjct: 366 VHPLPRYVMNYLKLLVDYGDPMDSLLELSEEDLYRFKNDLGGEWFRS*RPMSPLGQRILL 545
Query: 494 IMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSRL 535
+M L+ L KS+ +D A++ +F++NN ++ K S L
Sbjct: 546 LMSELEYNLEEKSKLYEDSAMQQVFLMNNLYYLVRKVKDSDL 671
>AW394508
Length = 396
Score = 70.5 bits (171), Expect = 2e-12
Identities = 42/102 (41%), Positives = 59/102 (57%), Gaps = 5/102 (4%)
Frame = -1
Query: 72 RSLRFKAHTAFLVLTITSVYSFFFDKVVNGKPD-----AYSLISCAAFAIMSMSLSRQSQ 126
RS KA +VLT+TS+ S + GK + +L AFA+M+MSLSRQ Q
Sbjct: 396 RSFLLKAQVCSVVLTLTSL*SVLEHRSEEGKVENAHGKMMNLTYSGAFALMAMSLSRQLQ 217
Query: 127 CGIEVDLLYFFLGCLIMQLMKIKLQLVIVGAGFSYSLIIIRS 168
G EV + YF +GC ++ +MK+ L+L + A F Y L+ IRS
Sbjct: 216 LGFEVGVFYFLVGCFLVTVMKMSLKLASLAALFCYLLVNIRS 91
>BG789722
Length = 392
Score = 67.0 bits (162), Expect = 3e-11
Identities = 35/126 (27%), Positives = 60/126 (46%)
Frame = +3
Query: 505 KSENCKDRALRHIFMLNNRSHIEAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWD 564
K+E KD A ++F+ NN ++ + S L + G +W +K K+++ Y+R W
Sbjct: 12 KAELYKDVAHSYLFLANNMQYVVVKVRKSNLGFLLGEEWLDKHKLKVREYASKYERVGWS 191
Query: 565 EVMDFLKLDNNESITKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILL 624
V L + +T E + F+ F CR Q++WF+ + R EI S+ + L+
Sbjct: 192 AVFSALPENPAAELTAEQARACFVRFDAAFHEACRKQASWFVSDPKFRDEIKGSIASKLV 371
Query: 625 PAYGIF 630
Y F
Sbjct: 372 QKYSEF 389
>AW734116
Length = 434
Score = 60.5 bits (145), Expect = 2e-09
Identities = 26/43 (60%), Positives = 33/43 (76%)
Frame = +3
Query: 12 LLQTKVWRFVGFASAAVGLLCYALSTSFNHLFGNWNLLKIFLY 54
L Q VW+FV F S VGL+CYALS+SF+ LFG W +LKIF++
Sbjct: 69 LKQVAVWKFVVFVSTVVGLVCYALSSSFSCLFGEWTMLKIFIF 197
Score = 32.7 bits (73), Expect = 0.53
Identities = 33/113 (29%), Positives = 56/113 (49%), Gaps = 3/113 (2%)
Frame = +2
Query: 125 SQCGIEVDLLYFFLGCLIMQLM--KIKLQLVIVGAGFSYSLIIIRSSLSSTNSVPENEYF 182
++ G +++ F C L+ ++ LQL I L + S+S + EN
Sbjct: 68 AEAGSSMEICCFCFNCCWTCLLCSQLLLQLSIWRMDHVEDLHLPLESVSKDTTSYEN--- 238
Query: 183 GVGLR-GENSVVIEMDSLLRPSNDIYSGMMQQLTTCINALQQDSLNIIDRLME 234
LR ++SVVI++++ + MMQQL TCI+ALQ+ + N+ + L E
Sbjct: 239 ---LRVQDDSVVIDVNN---------ARMMQQLMTCISALQRGNSNLTNILFE 361
>TC207607 similar to UP|Q9LJH9 (Q9LJH9) Gb|AAD25781.1, partial (20%)
Length = 1260
Score = 57.8 bits (138), Expect = 2e-08
Identities = 35/116 (30%), Positives = 58/116 (49%), Gaps = 10/116 (8%)
Frame = +1
Query: 558 YKRSAWDEVMDFLKLDN-------NESITKELLKEK-IHLFNNRFEAICRVQSAWFIYGS 609
Y+R W +V +L+ + + ++K + + FN+ FE + R Q+ W I S
Sbjct: 25 YQRETWVKVSYYLRDEGLHASGGFSSGVSKSXSQGXGLRTFNSMFEEVHRTQAVWLIPDS 204
Query: 610 QLRGEIISSVGNILLPAYGIFVGRLHGIL--GNQAYKYIKYGMIEIQDLLNHLFLG 663
QLR E+ S+ L+PAY F+GR + G YIKY + +++D + F G
Sbjct: 205 QLREELRISISEKLIPAYRSFLGRFRSYIESGRHPENYIKYSVEDLEDAVLDFFEG 372
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.327 0.139 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,017,747
Number of Sequences: 63676
Number of extensions: 430531
Number of successful extensions: 3038
Number of sequences better than 10.0: 58
Number of HSP's better than 10.0 without gapping: 2997
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3018
length of query: 672
length of database: 12,639,632
effective HSP length: 104
effective length of query: 568
effective length of database: 6,017,328
effective search space: 3417842304
effective search space used: 3417842304
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC138012.9


