

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138012.7 + phase: 0 /pseudo
(990 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
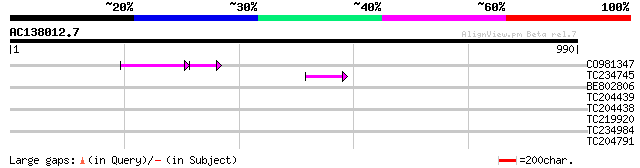
Score E
Sequences producing significant alignments: (bits) Value
CO981347 66 4e-15
TC234745 weakly similar to UP|Q9FFM0 (Q9FFM0) Copia-like retrotr... 57 5e-08
BE802806 38 0.019
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 36 0.074
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 34 0.37
TC219920 32 1.8
TC234984 similar to UP|Q84LM4 (Q84LM4) Acylamino acid-releasing ... 31 3.1
TC204791 weakly similar to UP|O82196 (O82196) Copia-like retroel... 30 5.3
>CO981347
Length = 624
Score = 65.9 bits (159), Expect(2) = 4e-15
Identities = 51/120 (42%), Positives = 63/120 (52%)
Frame = +3
Query: 194 KTKVTPLKSSKNGTLS*KIKWELN*KV*ELTMAWSLFQSSLMSFAG*KESRGIEPYQEHC 253
K K K+S+N L +I N*K * LTMAWSLF SS MSFAG S+G + H
Sbjct: 90 KNKSESFKNSENDILLLEINLVQN*KF*GLTMAWSLF*SSSMSFAGK*ASKGTK*SLTHH 269
Query: 254 NKMALRNA*IGHFWSV*GV*F*ELGYLRVSGVKP*QRLLI*SIDVHQRE*TSRHLWRYGV 313
++M + *I FW *G +* R G K + I* ID + * SRH W+ GV
Sbjct: 270 SRMV*QKG*I*PFWKE*GACY*VQDCQRPFGEKLQTQHHI*LIDALHQP*VSRHQWKLGV 449
Score = 34.7 bits (78), Expect(2) = 4e-15
Identities = 23/56 (41%), Positives = 29/56 (51%)
Frame = +1
Query: 315 NRRITPL*RFSELWHMRISSKTSLSLEL*NVSSLVIRKV*RDTSCGNWNLVEDQES 370
N I *R + W + + +K S L*+V SL I K + TS GNWNLV S
Sbjct: 454 NHLIIQD*RCLDHWPLIMLNKESWMQGL*SVCSLAILKELKGTSYGNWNLVRQDAS 621
>TC234745 weakly similar to UP|Q9FFM0 (Q9FFM0) Copia-like retrotransposable
element, partial (5%)
Length = 494
Score = 56.6 bits (135), Expect = 5e-08
Identities = 34/74 (45%), Positives = 45/74 (59%)
Frame = -1
Query: 517 *LDASGSSRKRMAFRVLKVQGIKQDLWQRVSLKCRGSTTTRSFHRW*SIVP*GYLWL**I 576
*LDAS SSRKR F+V + +G +QDLW RV K + H W*+I+ *G W *I
Sbjct: 266 *LDASRSSRKRKTFQV*RSRGTRQDLWLRVLHKWTELIIMKFSHLW*NIIL*GS*WQL*I 87
Query: 577 NSILSWNKWM*RPL 590
+ I S++ W+ R L
Sbjct: 86 SMIWSYHNWLPRQL 45
>BE802806
Length = 285
Score = 38.1 bits (87), Expect = 0.019
Identities = 28/85 (32%), Positives = 36/85 (41%)
Frame = -3
Query: 706 FLGLISRGIVTKANFSYLNLVI*RRWWSVLECQTLKL*ALPWVTTQSCP*NNVLNPRMRS 765
+ GLI GI +AN+S + R+WW L C L A QS +
Sbjct: 256 YSGLIFIGIEQRANYSCPKAITSRKWWKGLGCIKANLLAHHLAIIQSYLSLKHQKQLKKG 77
Query: 766 I*WKVLPMQVGLEASCMEWFVVGRT 790
+ PM + LEA CMEWF T
Sbjct: 76 LK*IKHPMPMVLEA*CMEWFAADLT 2
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 36.2 bits (82), Expect = 0.074
Identities = 42/143 (29%), Positives = 68/143 (47%), Gaps = 2/143 (1%)
Frame = +2
Query: 495 KKWIHWKGTKLGNL*NYLITKG*LDASGSSRKRMAFRVLKVQGIKQDLWQRVSLKCRGST 554
K W + KG K G+ L *L SGSSR + +V + + D + +L+ + T
Sbjct: 3257 KNWSNSKGMKSGS*FLGLRELM*LAPSGSSRTKPMKKVS*PE-TRPDWLLKATLRLKV*T 3433
Query: 555 TTRSFHRW*SIVP*GYLWL**INSILSWNKWM*RPLSYMVTLKRQSTWSNRKVLWRTSLR 614
R + + P Y +* ++S S +WM*R M T ++S WS+++ L ++
Sbjct: 3434 LMRLLPQLLDLSPSDYYLV*LVSSNSSCTRWM*RAHF*MDT*MKKSMWSSQRDLQTRLIQ 3613
Query: 615 CVF--*RNLCMG*SKALGNGIVG 635
++ R L M *SK G+ G
Sbjct: 3614 IMYTGSRRLSMD*SKLQELGMKG 3682
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 33.9 bits (76), Expect = 0.37
Identities = 43/155 (27%), Positives = 67/155 (42%)
Frame = +2
Query: 165 LLRLLLMGEVPIFFLSLMIILGECGSLF*KTKVTPLKSSKNGTLS*KIKWELN*KV*ELT 224
L +L + E + L MI GS + TPLK S++ K K ++ + +T
Sbjct: 2288 LCKLKALEEKGMPMLLWMISPDLPGSTLSERNQTPLKYSRS*V*DFKEKKTVSSRESGVT 2467
Query: 225 MAWSLFQSSLMSFAG*KESRGIEPYQEHCNKMALRNA*IGHFWSV*GV*F*ELGYLRVSG 284
MA SL +SL++ A K S H NKMA G + G F + +SG
Sbjct: 2468 MAESLKTASLLNSAHLKASLMSSLQPLHHNKMA*LKGKTGLCKKLLGSCFMPKNFPIISG 2647
Query: 285 VKP*QRLLI*SIDVHQRE*TSRHLWRYGVGNRRIT 319
+KP* + + + H E H + G G +++
Sbjct: 2648 LKP*TQHATSTTESHLEEGLQPHCMKSGKGGSQLS 2752
>TC219920
Length = 941
Score = 31.6 bits (70), Expect = 1.8
Identities = 22/58 (37%), Positives = 29/58 (49%), Gaps = 4/58 (6%)
Frame = -3
Query: 748 VTTQSCP*NNVLNPRMRSI*WKVLPMQVGLEA----SCMEWFVVGRTWLMRLAL*VGL 801
V QSC * V +M+ K+ P++ LE CME + RTWLM+ L V L
Sbjct: 366 VVYQSCS*RLVAESKMKGQLMKIHPLEGQLELLLSPPCMELCHLIRTWLMKKTLRVSL 193
>TC234984 similar to UP|Q84LM4 (Q84LM4) Acylamino acid-releasing enzyme ,
partial (14%)
Length = 682
Score = 30.8 bits (68), Expect = 3.1
Identities = 17/39 (43%), Positives = 23/39 (58%)
Frame = +3
Query: 781 CMEWFVVGRTWLMRLAL*VGLWQIRELCIGKL*SGF*GI 819
CM+W + LM L*V +Q E +G L*+G+*GI
Sbjct: 555 CMQWSIYD*ALLML*EL*VDSYQTLERSVGML*NGY*GI 671
>TC204791 weakly similar to UP|O82196 (O82196) Copia-like retroelement pol
polyprotein, partial (4%)
Length = 880
Score = 30.0 bits (66), Expect = 5.3
Identities = 21/55 (38%), Positives = 30/55 (54%)
Frame = +2
Query: 931 IKCIMKGQSTLTFACTLLET*LNQKRLWLRKWHRRKIRRMCSPSHCLDQDSSIAW 985
IK IM+G STLT T + + +K RK R +I+++CS S + S I W
Sbjct: 305 IKSIMRGLSTLTLDTT---SSILRKGSKSRKLIRGRIQQICSRSRF*ETSSCIIW 460
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.370 0.164 0.642
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 47,408,851
Number of Sequences: 63676
Number of extensions: 704531
Number of successful extensions: 9486
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 3818
Number of HSP's successfully gapped in prelim test: 298
Number of HSP's that attempted gapping in prelim test: 5455
Number of HSP's gapped (non-prelim): 4521
length of query: 990
length of database: 12,639,632
effective HSP length: 106
effective length of query: 884
effective length of database: 5,889,976
effective search space: 5206738784
effective search space used: 5206738784
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.8 bits)
S2: 64 (29.3 bits)
Medicago: description of AC138012.7


