

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
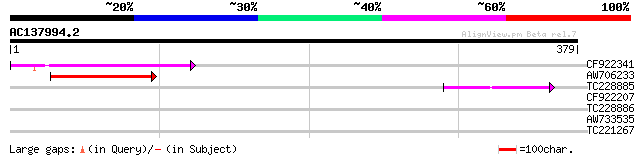
Query= AC137994.2 + phase: 0 /pseudo
(379 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CF922341 87 9e-18
AW706233 60 1e-09
TC228885 46 3e-05
CF922207 40 0.002
TC228886 39 0.005
AW733535 weakly similar to GP|4156245|dbj ATHP3 {Arabidopsis tha... 32 0.62
TC221267 30 2.3
>CF922341
Length = 675
Score = 87.4 bits (215), Expect = 9e-18
Identities = 51/133 (38%), Positives = 73/133 (54%), Gaps = 9/133 (6%)
Frame = +1
Query: 1 IPTTQQNHGPIFHAE--------SVKSYVRVDNLQEKYNEIQREMKALHRKELFGI-NAY 51
IP GP +H + S +V + K + ++ ++A+ E + N
Sbjct: 202 IPLRNTLEGPQYHPQLHLLHSTTSKNPHVMAE--MGKLDHLEEGLRAIEGGEDYAFANLE 375
Query: 52 DLCLVSNVVIPPKFKVPDFEKYKGNTFPKIHLVMYVRKMSYQADNDELFIHYFQDSLTGT 111
+L LV N++ PPKFKV DF+KYKG T PK HL MY +KM A ++EL IH FQ+SLTG
Sbjct: 376 ELFLVPNIITPPKFKVLDFDKYKGTTCPKNHLKMYCQKMGAYAKDEELLIHSFQESLTGV 555
Query: 112 ALLGYMGLDKSDI 124
A+ Y L+ S +
Sbjct: 556 AVTWYTNLEPSRV 594
>AW706233
Length = 376
Score = 60.5 bits (145), Expect = 1e-09
Identities = 30/72 (41%), Positives = 45/72 (61%), Gaps = 1/72 (1%)
Frame = -3
Query: 28 EKYNEIQREMKALHRKELFGI-NAYDLCLVSNVVIPPKFKVPDFEKYKGNTFPKIHLVMY 86
EK + ++ KA+ + + N +L LV N++ PPKFKV +F+KYKG T PK HL MY
Sbjct: 347 EKLDHLKERFKAIEGGQDYAFANLEELFLVXNIISPPKFKVLNFDKYKGTTCPKNHLKMY 168
Query: 87 VRKMSYQADNDE 98
+KM A +++
Sbjct: 167 CQKMGAYAKDEK 132
>TC228885
Length = 901
Score = 45.8 bits (107), Expect = 3e-05
Identities = 31/75 (41%), Positives = 39/75 (51%), Gaps = 1/75 (1%)
Frame = -3
Query: 291 IPMKYADLFPILLERNLVH-TKAPLPVPARLSARYRPNLFCAFHQGASCHDIEHCFSLKK 349
IP+ YADL P LL+ ++V T A + P L Y N CA H A IEH +LK+
Sbjct: 608 IPVSYADLLPYLLDNSMVAITLAKVHQPPFLR-EYDSNAMCACHGEAPGRSIEHYRALKR 432
Query: 350 IVQNLIPENLLPLED 364
VQ LI L E+
Sbjct: 431 KVQGLIDAGWLKFEE 387
>CF922207
Length = 616
Score = 39.7 bits (91), Expect = 0.002
Identities = 27/71 (38%), Positives = 35/71 (49%), Gaps = 3/71 (4%)
Frame = -1
Query: 297 DLFPILLERNLVH---TKAPLPVPARLSARYRPNLFCAFHQGASCHDIEHCFSLKKIVQN 353
+L P L+ ++V TK P P R Y N CA+H GA H IEHC + K V +
Sbjct: 535 NLIPYPLDNSMVAITPTKVPQPPFFR---EYDSNATCAYHGGAPGHSIEHCMTPKHKV*S 365
Query: 354 LIPENLLPLED 364
LI L E+
Sbjct: 364 LIDTG*LKFEE 332
>TC228886
Length = 748
Score = 38.5 bits (88), Expect = 0.005
Identities = 19/41 (46%), Positives = 23/41 (55%)
Frame = +3
Query: 324 YRPNLFCAFHQGASCHDIEHCFSLKKIVQNLIPENLLPLED 364
Y N CA+H GAS H IEHC + K V +LI L E+
Sbjct: 153 YDSNATCAYHGGASGHSIEHCMTPKHKV*SLIDTGWLKFEE 275
>AW733535 weakly similar to GP|4156245|dbj ATHP3 {Arabidopsis thaliana},
partial (88%)
Length = 454
Score = 31.6 bits (70), Expect = 0.62
Identities = 15/30 (50%), Positives = 18/30 (60%)
Frame = -2
Query: 241 SEYAEKSTSTTVSGLSPCCRCYA*RQCCSK 270
S Y ++TSTT SGL C C +* C SK
Sbjct: 153 SSYKRETTSTTKSGLFSSCSCCS*SNCPSK 64
>TC221267
Length = 539
Score = 29.6 bits (65), Expect = 2.3
Identities = 22/65 (33%), Positives = 29/65 (43%), Gaps = 8/65 (12%)
Frame = -3
Query: 243 YAEKSTSTTVSGLSPCCRCYA*RQCCSKSRLSTPISTISTTG*--------TQIDLIPMK 294
YA S STT+S PCC C KSR P T+ + * +++ P+K
Sbjct: 492 YANLSGSTTISVYKPCC*I---NSLC*KSRKHYPTPTLPGSP*P*KKSKNTSKVQQEPLK 322
Query: 295 YADLF 299
YA F
Sbjct: 321 YAPKF 307
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.340 0.147 0.501
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,876,080
Number of Sequences: 63676
Number of extensions: 318156
Number of successful extensions: 2479
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 2453
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2479
length of query: 379
length of database: 12,639,632
effective HSP length: 99
effective length of query: 280
effective length of database: 6,335,708
effective search space: 1773998240
effective search space used: 1773998240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC137994.2


