

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
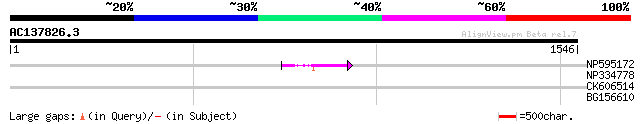
Query= AC137826.3 - phase: 0 /pseudo
(1546 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
NP595172 polyprotein [Glycine max] 57 8e-08
NP334778 reverse transcriptase [Glycine max] 32 2.8
CK606514 30 8.2
BG156610 weakly similar to GP|9828614|gb|A F1N21.3 {Arabidopsis ... 30 8.2
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 56.6 bits (135), Expect = 8e-08
Identities = 56/217 (25%), Positives = 95/217 (42%), Gaps = 22/217 (10%)
Frame = +1
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D S C DYR LN +T ++ + +P +D+L ++L A + +SK+ L +
Sbjct: 1930 DGSWRFCTDYRALNAITVKDSFPMPTVDELLDELHGAQY---FSKLDL-----------R 2067
Query: 800 KGIQDPLWAL*VSSDIVWVDKCSFGCHE------------------FD*LCD---QMTLG 838
G L V + +K +F H F L + Q L
Sbjct: 2068 SGYHQIL----VQPEDR--EKTAFRTHHGHYEWLVMPFGLTNAPATFQCLMNKIFQFALR 2229
Query: 839 FILIVCVDDILLFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNE 897
++V DDIL++ S D + HL V + +L+A SF ++ +G+ V
Sbjct: 2230 KFVLVFFDDILIYSASWKDHLKHLESVLQTLKQHQLFARLSKCSFGDTEVDYLGHKVSGL 2409
Query: 898 RIMVDLQNIEPVMNWV*PSFVIESRGFMGLASYYRGF 934
+ ++ ++ V++W P+ V + RGF+GL YYR F
Sbjct: 2410 GVSMENTKVQAVLDWPTPNNVKQLRGFLGLTGYYRRF 2520
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 31.6 bits (70), Expect = 2.8
Identities = 16/34 (47%), Positives = 21/34 (61%)
Frame = +3
Query: 745 MCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASF 778
MC DYR LNR + ++ + LP ID L + ASF
Sbjct: 6 MCVDYRDLNRASPKDNFPLPHIDILMANM--ASF 101
>CK606514
Length = 280
Score = 30.0 bits (66), Expect = 8.2
Identities = 15/35 (42%), Positives = 23/35 (64%), Gaps = 2/35 (5%)
Frame = +3
Query: 27 RSLFKGRERGRVEQA--RGGAPFEKSPREKAPYAP 59
++L KG++RG++ A RGG +K PRE P +P
Sbjct: 60 QNLKKGKKRGKIFWAKKRGGGGGKKKPRETPPISP 164
>BG156610 weakly similar to GP|9828614|gb|A F1N21.3 {Arabidopsis thaliana},
partial (23%)
Length = 293
Score = 30.0 bits (66), Expect = 8.2
Identities = 13/42 (30%), Positives = 19/42 (44%)
Frame = +1
Query: 405 YNCGDFRHISRYCPKPRILDLMQK*TRAVVPASNDKNSRGHP 446
+NCG + H R CP+PR + + N +S HP
Sbjct: 55 FNCGSYNHSLRECPRPRDNIAVNNARDKLKSRRNQNSSSRHP 180
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.359 0.161 0.583
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 74,009,100
Number of Sequences: 63676
Number of extensions: 1125316
Number of successful extensions: 10553
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 5581
Number of HSP's successfully gapped in prelim test: 378
Number of HSP's that attempted gapping in prelim test: 4770
Number of HSP's gapped (non-prelim): 6384
length of query: 1546
length of database: 12,639,632
effective HSP length: 110
effective length of query: 1436
effective length of database: 5,635,272
effective search space: 8092250592
effective search space used: 8092250592
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 65 (29.6 bits)
Medicago: description of AC137826.3


