

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137825.4 - phase: 0
(161 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
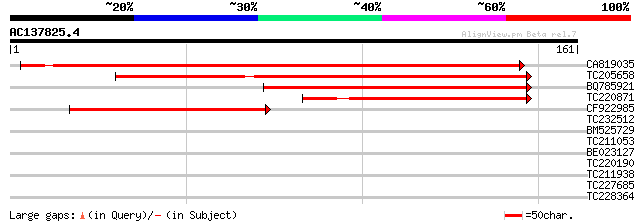
Score E
Sequences producing significant alignments: (bits) Value
CA819035 234 1e-62
TC205658 weakly similar to UP|Q9LWK9 (Q9LWK9) Oryza sativa (japo... 136 4e-33
BQ785921 129 6e-31
TC220871 weakly similar to UP|Q9LWK9 (Q9LWK9) Oryza sativa (japo... 100 4e-22
CF922985 98 2e-21
TC232512 28 2.5
BM525729 homologue to GP|11994408|dbj CTP synthase {Arabidopsis ... 27 3.2
TC211053 27 5.5
BE023127 26 7.2
TC220190 weakly similar to UP|Q84630 (Q84630) A316R protein, par... 26 9.4
TC211938 similar to GB|AAD17339.1|4325339|F15P23 F15P23.1 {Arabi... 26 9.4
TC227685 similar to UP|O23509 (O23509) HSP like protein (Fragmen... 26 9.4
TC228364 26 9.4
>CA819035
Length = 447
Score = 234 bits (598), Expect = 1e-62
Identities = 112/143 (78%), Positives = 129/143 (89%)
Frame = +3
Query: 4 LLGVLVMIVLLGNSIYAYGAINQLSWREISNINNEGPSIGIVVPNAYELNPLLHSSSFVP 63
LLG+LV+ LLG+SI A GA++++SWREISNIN+EGP +GIVVPN++EL PLL SSSFVP
Sbjct: 24 LLGLLVL--LLGSSIMANGAVSEVSWREISNINSEGPYLGIVVPNSFELGPLLRSSSFVP 197
Query: 64 HNKFPYFDFAGRHFRIGELEKKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGN 123
HNKFPYFDFAG+HFRIG LEKK+VIVVM G GMLNAGL+TQLLLTLFN+ GVLHYGIAGN
Sbjct: 198 HNKFPYFDFAGKHFRIGVLEKKRVIVVMCGLGMLNAGLSTQLLLTLFNVNGVLHYGIAGN 377
Query: 124 LNSRFQIGDVTIPKYWAHTGLWH 146
N + QIGDVTIP+YWAHTGLWH
Sbjct: 378 ANPKLQIGDVTIPQYWAHTGLWH 446
>TC205658 weakly similar to UP|Q9LWK9 (Q9LWK9) Oryza sativa (japonica
cultivar-group) genomic DNA, chromosome 1, PAC
clone:P0485D09, partial (72%)
Length = 1153
Score = 136 bits (343), Expect = 4e-33
Identities = 61/118 (51%), Positives = 83/118 (69%)
Frame = +1
Query: 31 EISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKFPYFDFAGRHFRIGELEKKKVIVV 90
+I+ N EGP +G+++PN++EL+PLL + + + DFAGR FR G + K VI+V
Sbjct: 115 KIAKANQEGPYLGLIIPNSFELDPLLQNPGYTASDTI--IDFAGRRFRFGAIGDKPVILV 288
Query: 91 MSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTIPKYWAHTGLWHWQ 148
M+G ++NA + TQLLL+LF +EGV+HYGIAGN N IGDV IP+YWAH LW WQ
Sbjct: 289 MTGLSVINAAITTQLLLSLFTVEGVVHYGIAGNANPSLHIGDVAIPQYWAHLALWSWQ 462
>BQ785921
Length = 481
Score = 129 bits (324), Expect = 6e-31
Identities = 59/76 (77%), Positives = 69/76 (90%)
Frame = +1
Query: 73 AGRHFRIGELEKKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGD 132
AG+HFRIG LEKK+VIVVM+ MLNAG++TQLLLTLF+IEGV+HYGIAGN N + QIGD
Sbjct: 130 AGKHFRIGVLEKKRVIVVMTFLSMLNAGISTQLLLTLFDIEGVVHYGIAGNANPKLQIGD 309
Query: 133 VTIPKYWAHTGLWHWQ 148
VTIP+YWAHTGLW+WQ
Sbjct: 310 VTIPQYWAHTGLWNWQ 357
>TC220871 weakly similar to UP|Q9LWK9 (Q9LWK9) Oryza sativa (japonica
cultivar-group) genomic DNA, chromosome 1, PAC
clone:P0485D09, partial (66%)
Length = 902
Score = 100 bits (248), Expect = 4e-22
Identities = 47/65 (72%), Positives = 56/65 (85%)
Frame = +2
Query: 84 KKKVIVVMSGEGMLNAGLATQLLLTLFNIEGVLHYGIAGNLNSRFQIGDVTIPKYWAHTG 143
KKKV+V + LNAG++TQLLLTLF+IEGV+HYGIAGN N + QIGDVTIP+YWAHTG
Sbjct: 2 KKKVLVSLL*---LNAGISTQLLLTLFDIEGVVHYGIAGNANPKLQIGDVTIPQYWAHTG 172
Query: 144 LWHWQ 148
LW+WQ
Sbjct: 173 LWNWQ 187
>CF922985
Length = 497
Score = 97.8 bits (242), Expect = 2e-21
Identities = 44/57 (77%), Positives = 50/57 (87%)
Frame = +2
Query: 18 IYAYGAINQLSWREISNINNEGPSIGIVVPNAYELNPLLHSSSFVPHNKFPYFDFAG 74
I A GA++++SWREIS IN EGP +GIVVPNA+ELNPLL S SFVPHNKFPYFDFAG
Sbjct: 2 ISANGAVSEVSWREISKINREGPYVGIVVPNAFELNPLLRSPSFVPHNKFPYFDFAG 172
>TC232512
Length = 804
Score = 27.7 bits (60), Expect = 2.5
Identities = 8/16 (50%), Positives = 12/16 (75%)
Frame = -2
Query: 145 WHWQVHTSCFNYLDTF 160
W+W+ H SCFN+ + F
Sbjct: 416 WNWKPHLSCFNFSNFF 369
>BM525729 homologue to GP|11994408|dbj CTP synthase {Arabidopsis thaliana},
partial (6%)
Length = 429
Score = 27.3 bits (59), Expect = 3.2
Identities = 15/32 (46%), Positives = 18/32 (55%), Gaps = 3/32 (9%)
Frame = -1
Query: 28 SWREISNINNEGPSIGIVV---PNAYELNPLL 56
S RE + EG S+ + V PNAY LNP L
Sbjct: 114 SARECERVEREGRSLRLCVGLMPNAYWLNPFL 19
>TC211053
Length = 617
Score = 26.6 bits (57), Expect = 5.5
Identities = 9/38 (23%), Positives = 21/38 (54%)
Frame = -3
Query: 118 YGIAGNLNSRFQIGDVTIPKYWAHTGLWHWQVHTSCFN 155
+GI N++ + ++ + I +YW + + +H C+N
Sbjct: 195 FGIRTNISLKIELDLLLIRRYWQNLVGLRFSIHRECWN 82
>BE023127
Length = 277
Score = 26.2 bits (56), Expect = 7.2
Identities = 13/32 (40%), Positives = 16/32 (49%), Gaps = 2/32 (6%)
Frame = +2
Query: 126 SRFQIGDVTIPK--YWAHTGLWHWQVHTSCFN 155
S I D IP YW +G W+ HT CF+
Sbjct: 50 STITIQDQVIPSALYWWESGGRAWKRHTHCFD 145
>TC220190 weakly similar to UP|Q84630 (Q84630) A316R protein, partial (11%)
Length = 602
Score = 25.8 bits (55), Expect = 9.4
Identities = 6/12 (50%), Positives = 9/12 (75%)
Frame = +1
Query: 136 PKYWAHTGLWHW 147
P++W+ GLW W
Sbjct: 289 PRFWSRNGLWSW 324
>TC211938 similar to GB|AAD17339.1|4325339|F15P23 F15P23.1 {Arabidopsis
thaliana;} , partial (20%)
Length = 518
Score = 25.8 bits (55), Expect = 9.4
Identities = 8/20 (40%), Positives = 11/20 (55%)
Frame = +2
Query: 131 GDVTIPKYWAHTGLWHWQVH 150
G V +P+ W H W +VH
Sbjct: 293 GPVQVPRRWDHVSTWRRRVH 352
>TC227685 similar to UP|O23509 (O23509) HSP like protein (Fragment), partial
(28%)
Length = 463
Score = 25.8 bits (55), Expect = 9.4
Identities = 7/14 (50%), Positives = 10/14 (71%)
Frame = -3
Query: 144 LWHWQVHTSCFNYL 157
LWHW +H S +N +
Sbjct: 431 LWHWHLHPSVYNLI 390
>TC228364
Length = 1129
Score = 25.8 bits (55), Expect = 9.4
Identities = 15/41 (36%), Positives = 21/41 (50%), Gaps = 5/41 (12%)
Frame = -2
Query: 33 SNINNEGPSIGIVVPNAYELNPLLHSSS-----FVPHNKFP 68
S N+ G + G +VP + P+L SSS PH +FP
Sbjct: 267 SERNDFGRTEGQMVPTVFPF*PILDSSSQDLPFSYPHRRFP 145
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.142 0.448
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,313,661
Number of Sequences: 63676
Number of extensions: 115176
Number of successful extensions: 930
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 926
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 928
length of query: 161
length of database: 12,639,632
effective HSP length: 90
effective length of query: 71
effective length of database: 6,908,792
effective search space: 490524232
effective search space used: 490524232
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 55 (25.8 bits)
Medicago: description of AC137825.4


