

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137823.16 + phase: 0
(621 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
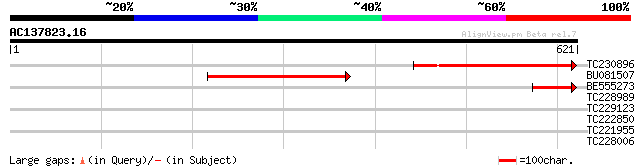
Score E
Sequences producing significant alignments: (bits) Value
TC230896 similar to UP|Q7XYR0 (Q7XYR0) At5g38880, partial (23%) 283 2e-76
BU081507 280 1e-75
BE555273 86 5e-17
TC228989 similar to UP|Q9LSS5 (Q9LSS5) Myosin heavy chain-like, ... 32 0.64
TC229123 32 1.1
TC222850 similar to UP|Q8LBL6 (Q8LBL6) Cell division protein Fts... 30 4.2
TC221955 weakly similar to UP|Q6UAL1 (Q6UAL1) Myosin heavy chain... 29 5.4
TC228006 homologue to UP|IM30_PEA (Q03943) Membrane-associated 3... 28 9.3
>TC230896 similar to UP|Q7XYR0 (Q7XYR0) At5g38880, partial (23%)
Length = 697
Score = 283 bits (723), Expect = 2e-76
Identities = 143/178 (80%), Positives = 155/178 (86%)
Frame = +3
Query: 443 ARAGARDPSAIPSICRVSAALQYAAGGLEGSDAGLASILESLEFCLKRRGSEASVLEDLL 502
ARAGARDPSAIPSICRVSAAL Y AG LEGSDAGLAS+LESLEFCLK RGSEASVLEDLL
Sbjct: 12 ARAGARDPSAIPSICRVSAALHYPAG-LEGSDAGLASVLESLEFCLKLRGSEASVLEDLL 188
Query: 503 KAINLVHIRRDLVQSGHALLNHAYFVQQDYERTTNFSLNLAAEQERAVMEKWLPELKTGV 562
+AINLV+IRRDLVQSG ALLNHA VQQ+YE+TT F L+ A EQE+ +ME+WLPELK V
Sbjct: 189 RAINLVYIRRDLVQSGEALLNHANLVQQEYEKTTKFCLSKADEQEKTIMEEWLPELKNAV 368
Query: 563 LNAQQSLEACKYVWGLLDEWWEQPASTVVDWATVDGSNVAFWHNHVKKLLTCYDQELL 620
L+AQQSLE CKYV GLLDEWWEQPASTVVDW TVDG NV WHNHVK+LL D+ELL
Sbjct: 369 LSAQQSLEDCKYVRGLLDEWWEQPASTVVDWVTVDGQNVTAWHNHVKQLLAFCDKELL 542
>BU081507
Length = 474
Score = 280 bits (717), Expect = 1e-75
Identities = 142/157 (90%), Positives = 149/157 (94%)
Frame = +3
Query: 217 QNQLLERQKAHVQQFLATEDALNNAAEARDLCEKLMKRLHGGTDVTSRSIGIGATSQNVG 276
QNQLL+RQKAHVQQFLATEDALN AAEARD+CEKLMKRLHGGTDV+SRS+GIG+ SQNVG
Sbjct: 3 QNQLLDRQKAHVQQFLATEDALNKAAEARDMCEKLMKRLHGGTDVSSRSLGIGSNSQNVG 182
Query: 277 SLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKWKKIEEFDGR 336
SLRQ LDVWAKEREV+GLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKWKKIEEFD R
Sbjct: 183 SLRQF*LDVWAKEREVAGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKWKKIEEFDAR 362
Query: 337 RSELESIYTALLKANTDAASFWSQQPSTAREYALSTI 373
RSELE+IY ALLKAN DAASFWSQQP TAREYALSTI
Sbjct: 363 RSELETIYMALLKANMDAASFWSQQPLTAREYALSTI 473
>BE555273
Length = 356
Score = 85.9 bits (211), Expect = 5e-17
Identities = 38/48 (79%), Positives = 40/48 (83%)
Frame = +1
Query: 573 KYVWGLLDEWWEQPASTVVDWATVDGSNVAFWHNHVKKLLTCYDQELL 620
KYV GLLDEWWEQPASTVVDW TVDG NV WHNHVK+LL D+ELL
Sbjct: 7 KYVRGLLDEWWEQPASTVVDWVTVDGQNVTAWHNHVKQLLAFCDKELL 150
>TC228989 similar to UP|Q9LSS5 (Q9LSS5) Myosin heavy chain-like, partial
(23%)
Length = 884
Score = 32.3 bits (72), Expect = 0.64
Identities = 39/175 (22%), Positives = 71/175 (40%), Gaps = 19/175 (10%)
Frame = +1
Query: 280 QLQLDVWAKEREVSGLKASLNTL-----------MSEIQRLNKLCAERK--------EAE 320
QL+ +V + + EV L++++ T M +I+ +L + K E E
Sbjct: 34 QLEAEVVSLKSEVEQLRSAIETAETKYQEEQIRSMVQIRNAYELVEQIKSESGKRECELE 213
Query: 321 DSLKKKWKKIEEFDGRRSELESIYTALLKANTDAASFWSQQPSTAREYALSTIIPACSAV 380
LKKK IEE + E+ +++ N + + S+ E+ L I +
Sbjct: 214 AELKKKKADIEELKANLMDKETELQGIVEENDNLNLKLEESMSSKNEHKLKREIKRLAEC 393
Query: 381 VETSNSAKDLIEKEVSAFYRSPDNSLYMLPSSPQALLEAIGSSGSSGQEAVANAE 435
V + D+++KE + S +N + L I +SG + +E A E
Sbjct: 394 V--AELKADMMDKETTLQSISEENEM---------LKSEINNSGKAREEVAAEVE 525
>TC229123
Length = 1034
Score = 31.6 bits (70), Expect = 1.1
Identities = 16/51 (31%), Positives = 31/51 (60%)
Frame = +3
Query: 275 VGSLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKK 325
V S+ QLQ + + K+ ++S L++ + + +E +RLNK + +SL+K
Sbjct: 402 VSSMEQLQSEKYEKDSKISELQSKMAEMEAETKRLNKEISGLSVELESLRK 554
>TC222850 similar to UP|Q8LBL6 (Q8LBL6) Cell division protein FtsH-like
protein, partial (14%)
Length = 458
Score = 29.6 bits (65), Expect = 4.2
Identities = 18/50 (36%), Positives = 28/50 (56%)
Frame = +1
Query: 456 ICRVSAALQYAAGGLEGSDAGLASILESLEFCLKRRGSEASVLEDLLKAI 505
IC + A+L GL G+D LA+++ RRGSE ED+++A+
Sbjct: 76 ICHLIASL---TTGLVGAD--LANVVNEAALLAARRGSETVAREDIMEAV 210
>TC221955 weakly similar to UP|Q6UAL1 (Q6UAL1) Myosin heavy chain class XI E3
protein, partial (6%)
Length = 728
Score = 29.3 bits (64), Expect = 5.4
Identities = 20/64 (31%), Positives = 32/64 (49%)
Frame = +1
Query: 308 RLNKLCAERKEAEDSLKKKWKKIEEFDGRRSELESIYTALLKANTDAASFWSQQPSTARE 367
+L L + +E E +++ KKI+EF+ SE+E+ A LK +A +Q T
Sbjct: 157 KLELLTNKNEELETEVEELKKKIKEFEESYSEIENENQARLKEAEEAQLKATQLQETIER 336
Query: 368 YALS 371
LS
Sbjct: 337 LELS 348
>TC228006 homologue to UP|IM30_PEA (Q03943) Membrane-associated 30 kDa
protein, chloroplast precursor (M30), partial (34%)
Length = 939
Score = 28.5 bits (62), Expect = 9.3
Identities = 13/25 (52%), Positives = 17/25 (68%)
Frame = -3
Query: 286 WAKEREVSGLKASLNTLMSEIQRLN 310
WA R +S L S TL+SEI++LN
Sbjct: 793 WASSRTIS*LAFSCTTLISEIKKLN 719
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.314 0.129 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,416,673
Number of Sequences: 63676
Number of extensions: 280332
Number of successful extensions: 1062
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1057
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1061
length of query: 621
length of database: 12,639,632
effective HSP length: 103
effective length of query: 518
effective length of database: 6,081,004
effective search space: 3149960072
effective search space used: 3149960072
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 62 (28.5 bits)
Medicago: description of AC137823.16


