

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137822.5 - phase: 0
(203 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
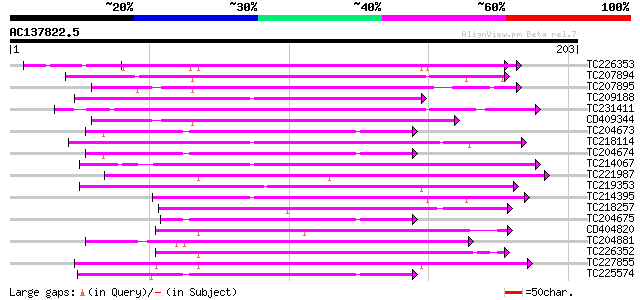
Score E
Sequences producing significant alignments: (bits) Value
TC226353 weakly similar to UP|Q9M510 (Q9M510) Dicyanin, partial ... 103 7e-23
TC207894 similar to UP|O81500 (O81500) F9D12.16 protein (At5g263... 97 5e-21
TC207895 similar to UP|O81500 (O81500) F9D12.16 protein (At5g263... 94 3e-20
TC209188 similar to UP|BCP_PEA (Q41001) Blue copper protein prec... 93 7e-20
TC231411 similar to GB|AAP21370.1|30102904|BT006562 At3g27200 {A... 91 4e-19
CD409344 90 8e-19
TC204673 similar to UP|Q9ZRV5 (Q9ZRV5) Basic blue copper protein... 88 3e-18
TC218114 similar to UP|BCP_PEA (Q41001) Blue copper protein prec... 85 3e-17
TC204674 similar to UP|Q9ZRV5 (Q9ZRV5) Basic blue copper protein... 85 3e-17
TC214067 weakly similar to UP|Q949E8 (Q949E8) Uclacyanin 3-like ... 82 2e-16
TC221987 nodulin 81 4e-16
TC219353 weakly similar to UP|Q8L555 (Q8L555) Uclacyanin 3-like ... 78 2e-15
TC214395 weakly similar to UP|Q949E8 (Q949E8) Uclacyanin 3-like ... 77 4e-15
TC218257 76 1e-14
TC204675 homologue to UP|Q8GV76 (Q8GV76) Auxin efflux carrier pr... 75 2e-14
CD404820 weakly similar to SP|Q41001|BCP_P Blue copper protein p... 75 2e-14
TC204881 weakly similar to UP|ENL1_ARATH (Q9SK27) Early nodulin-... 75 3e-14
TC226352 weakly similar to UP|Q9M510 (Q9M510) Dicyanin, partial ... 74 3e-14
TC227855 weakly similar to GB|AAP88350.1|32815931|BT009716 At3g2... 74 5e-14
TC225574 similar to UP|Q9ZRV5 (Q9ZRV5) Basic blue copper protein... 73 8e-14
>TC226353 weakly similar to UP|Q9M510 (Q9M510) Dicyanin, partial (26%)
Length = 1341
Score = 103 bits (256), Expect = 7e-23
Identities = 68/195 (34%), Positives = 96/195 (48%), Gaps = 17/195 (8%)
Frame = +3
Query: 6 RIQSLILSNKIKQRKSNTSTSTMALSRALFLFAF-IATIFSTMAVAKDFVVGDEKGWTT- 63
R + L+L+ +IKQ +S+ +A++R L L F +AT+ A +VGD GW
Sbjct: 12 RHKDLVLA-RIKQGRSHLQEH-IAMARNLLLVLFAVATLLHGSAAQTRHMVGDATGWIIP 185
Query: 64 ---LFDYQTWTANKVFRLGDTLTFNYVGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKI 120
Y W +NK F + DTL FN+ G+ NV +V S F +C+ LTSG +
Sbjct: 186 AGGAATYTAWASNKTFTVNDTLVFNFATGQHNVAKVTKSAFDACNGGSAVFTLTSGPATV 365
Query: 121 IITTYGRRWYISSVTDHCENGQKLFITV------------QPKQDGWSPVPSPSPSPSLD 168
+ G ++YI SV HC GQKL I V QP+ G P SP P+ +
Sbjct: 366 TLNETGEQYYICSVGSHCSAGQKLAINVNRASSTGPSPAPQPRGSGSPPRASPVPTQAPQ 545
Query: 169 LVTPEAPPSNAPWPA 183
+P PP +AP PA
Sbjct: 546 ASSPTPPPRSAPAPA 590
Score = 75.9 bits (185), Expect = 1e-14
Identities = 47/148 (31%), Positives = 66/148 (43%), Gaps = 9/148 (6%)
Frame = +3
Query: 41 ATIFSTMAVAKDFVVGDEKGWTTLFD---YQTWTANKVFRLGDTLTFNYVGGKDNVVRVN 97
A F + F+VG+ GW + Y W + K FR+GD L FNY NV V
Sbjct: 582 APAFGPSSEPATFIVGETAGWIVPGNASFYTAWASGKNFRVGDVLVFNYASNTHNVEEVT 761
Query: 98 GSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFI------TVQPK 151
++F +CS T+ ++ + G+ ++I + HC GQKL I T P
Sbjct: 762 KANFDACSSASPIATFTTPPARVTLNKSGQHFFICGIPGHCLGGQKLAINVTGSSTATPP 941
Query: 152 QDGWSPVPSPSPSPSLDLVTPEAPPSNA 179
P SPSP+ VTP PP N+
Sbjct: 942 SAAAPPTTPSSPSPA-GAVTP--PPQNS 1016
>TC207894 similar to UP|O81500 (O81500) F9D12.16 protein (At5g26330), partial
(56%)
Length = 755
Score = 97.1 bits (240), Expect = 5e-21
Identities = 60/164 (36%), Positives = 87/164 (52%), Gaps = 5/164 (3%)
Frame = +2
Query: 21 SNTSTSTMALSRALFLFAFIATIFSTMAVAKDFVVGDEKGWTTL--FDYQTWTANKVFRL 78
++ TM L + + + TI ++ A + VGD GWTTL DY+ W A K F++
Sbjct: 26 TSAGLETMTLVERVVVLFIVMTIVK-VSYAAVYKVGDSAGWTTLGTIDYRKWAATKNFQI 202
Query: 79 GDTLTFNYVGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHC 138
GDT+ F Y NV+RV + +K+C+ T+G+D I IT +G ++ V HC
Sbjct: 203 GDTIIFEYNAKFHNVMRVTHAMYKTCNASSPIATFTTGKDSINITNHGHHFFFCGVPGHC 382
Query: 139 ENGQKLFITVQPKQDGWSPVPSPS--PSPSLDLVTPEAP-PSNA 179
+ GQK+ I V K +P PS S SP++ T AP PSNA
Sbjct: 383 QAGQKVDINVL-KVSAEAPTPSGSALASPTVQASTVPAPSPSNA 511
>TC207895 similar to UP|O81500 (O81500) F9D12.16 protein (At5g26330), partial
(53%)
Length = 858
Score = 94.4 bits (233), Expect = 3e-20
Identities = 55/158 (34%), Positives = 84/158 (52%), Gaps = 4/158 (2%)
Frame = +3
Query: 30 LSRALFLFAFIATIF--STMAVAKDFVVGDEKGWTTL--FDYQTWTANKVFRLGDTLTFN 85
+ +A+F + T F S AV K VGD GWT + DY+ W A K F++GDT+ F
Sbjct: 138 IEKAVFFLMMMMTAFQVSHAAVHK---VGDSAGWTIIGNIDYKKWAATKNFQVGDTIIFE 308
Query: 86 YVGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLF 145
Y NV+RV + +KSC+ +++G D I IT YG +++ + HC+ GQK+
Sbjct: 309 YNAKFHNVMRVTHAMYKSCNASSPLTTMSTGNDTIKITNYGHHFFLCGIPGHCQAGQKVD 488
Query: 146 ITVQPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPA 183
I V + +PSP+ + +P PP+N P P+
Sbjct: 489 INVVKVS------AAAAPSPTSAMASP-VPPANVPAPS 581
>TC209188 similar to UP|BCP_PEA (Q41001) Blue copper protein precursor,
partial (57%)
Length = 597
Score = 93.2 bits (230), Expect = 7e-20
Identities = 49/126 (38%), Positives = 69/126 (53%)
Frame = +1
Query: 24 STSTMALSRALFLFAFIATIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLT 83
S MA S AL L++ +A + +A + VGD GW DY TWT +K+F +GD+L
Sbjct: 46 SHKIMAFSSALILWSLLAINMALPTLATVYTVGDTSGWAIGTDYSTWTGDKIFSVGDSLA 225
Query: 84 FNYVGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQK 143
FNY G V V SD+KSC+ + +SG I + + G ++I SV HC G K
Sbjct: 226 FNY-GAGHTVDEVKESDYKSCTAGNSISTDSSGATTIALKSAGTHYFICSVPGHCSGGMK 402
Query: 144 LFITVQ 149
L +TV+
Sbjct: 403 LAVTVK 420
>TC231411 similar to GB|AAP21370.1|30102904|BT006562 At3g27200 {Arabidopsis
thaliana;} , partial (57%)
Length = 855
Score = 90.9 bits (224), Expect = 4e-19
Identities = 58/174 (33%), Positives = 87/174 (49%)
Frame = +2
Query: 17 KQRKSNTSTSTMALSRALFLFAFIATIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVF 76
+QRK+ T M +FL A IAT+ + A A VVG +GW D+++WT+ + F
Sbjct: 38 EQRKNQT----MGHKNTIFL-ALIATLIAKEAFAAQHVVGGSQGWDQSTDFKSWTSGQTF 202
Query: 77 RLGDTLTFNYVGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTD 136
++GD L F Y V N S +K+C + L++G D + + G R++
Sbjct: 203 KVGDKLVFKYSSFHSVVELGNESAYKNCDISSPVQSLSTGNDVVKLDKPGTRYFTCGTLG 382
Query: 137 HCENGQKLFITVQPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSVPRRS 190
HC G K+ IT++ K + SP SP+ SPSL +P P + SS P S
Sbjct: 383 HCSQGMKVKITIR-KGNAPSPALSPATSPSL---SPSLSPLLSSTSLSSPPSSS 532
>CD409344
Length = 438
Score = 89.7 bits (221), Expect = 8e-19
Identities = 50/134 (37%), Positives = 69/134 (51%), Gaps = 2/134 (1%)
Frame = -2
Query: 30 LSRALFLFAFIATIFSTMAVAKDFVVGDEKGWTTL--FDYQTWTANKVFRLGDTLTFNYV 87
+ +A+ A S AV K VGD GWT + DY+ W A K F++GDT+ F Y
Sbjct: 419 IEKAVVFLMMTAFQVSNSAVHK---VGDSAGWTIIGNIDYKKWAATKNFQVGDTIIFEYN 249
Query: 88 GGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFIT 147
NV+RV +KSC+ +++G D I IT YG ++ V HC+ GQK+ I
Sbjct: 248 AKFHNVMRVTHGMYKSCNASSPLTRMSTGNDTIKITNYGHHLFLCGVPGHCQAGQKVDIN 69
Query: 148 VQPKQDGWSPVPSP 161
V K +P PSP
Sbjct: 68 VVKKVSAEAPTPSP 27
>TC204673 similar to UP|Q9ZRV5 (Q9ZRV5) Basic blue copper protein, partial
(98%)
Length = 582
Score = 87.8 bits (216), Expect = 3e-18
Identities = 51/121 (42%), Positives = 66/121 (54%), Gaps = 2/121 (1%)
Frame = +3
Query: 28 MALSR--ALFLFAFIATIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFN 85
MAL R A+ L + S MA A + VGD GWT F+ W K+FR GDTL FN
Sbjct: 45 MALGRGSAVVLLLCFLLLHSQMARAATYTVGDSGGWT--FNTVAWPKGKLFRAGDTLAFN 218
Query: 86 YVGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLF 145
Y G NVV VN + + SC P A V SG D+I + G+ ++I + HCE+G K+
Sbjct: 219 YSPGTHNVVAVNKAGYDSCKTPRGAKVYKSGTDQIRLAK-GQNYFICNYVGHCESGMKIA 395
Query: 146 I 146
I
Sbjct: 396 I 398
>TC218114 similar to UP|BCP_PEA (Q41001) Blue copper protein precursor,
partial (57%)
Length = 901
Score = 84.7 bits (208), Expect = 3e-17
Identities = 57/168 (33%), Positives = 81/168 (47%), Gaps = 4/168 (2%)
Frame = +3
Query: 22 NTSTSTMALSRALFLFAFIATIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDT 81
+T + MA S AL L +A A VGD GW DY TW + ++GD+
Sbjct: 33 HTHSH*MASSVALVLGLCLALNMVLPTRAATHTVGDTSGWALGADYSTWASGLKLKVGDS 212
Query: 82 LTFNYVGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENG 141
L FNY G V V SD+KSC+ + +SG I + T G ++I + HC+ G
Sbjct: 213 LVFNY-GAGHTVDEVKESDYKSCTTGNSLSTDSSGTTTITLKTAGTHYFICASPGHCDGG 389
Query: 142 QKLFITVQPKQDGWSPVPSPSP----SPSLDLVTPEAPPSNAPWPASS 185
KL + V+ K+ +P +PSP SPS T + P S + P +S
Sbjct: 390 MKLAVKVKAKKAS-APATAPSPAAKDSPSDSDDTKDTPTSTSTNPKTS 530
>TC204674 similar to UP|Q9ZRV5 (Q9ZRV5) Basic blue copper protein, partial
(98%)
Length = 635
Score = 84.7 bits (208), Expect = 3e-17
Identities = 50/121 (41%), Positives = 67/121 (55%), Gaps = 2/121 (1%)
Frame = +2
Query: 28 MALSR--ALFLFAFIATIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFN 85
MAL R A+ L + S MA A + VGD +GWT F+ TW K F GDTL FN
Sbjct: 53 MALGRGSAVVLLLCFLVLQSEMARAATYRVGDSRGWT--FNTVTWPQGKRFTAGDTLAFN 226
Query: 86 YVGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLF 145
Y G NVV V+ + + SC P A V SG+D+I + G+ ++I + HCE+G K+
Sbjct: 227 YSPGAHNVVAVSKAGYDSCKTPRGAKVYRSGKDQIRLAR-GQNYFICNYVGHCESGMKIA 403
Query: 146 I 146
I
Sbjct: 404 I 406
>TC214067 weakly similar to UP|Q949E8 (Q949E8) Uclacyanin 3-like protein,
partial (50%)
Length = 948
Score = 81.6 bits (200), Expect = 2e-16
Identities = 52/165 (31%), Positives = 75/165 (44%)
Frame = +1
Query: 26 STMALSRALFLFAFIATIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFN 85
S +A S + L AF T+F D VGD GW +Y TW + K F +GDTL F
Sbjct: 94 SAIAASFLVLLLAF-PTVFGA-----DHEVGDTSGWALGVNYNTWASGKTFTVGDTLVFK 255
Query: 86 YVGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLF 145
Y V V+ S + SCS + G KI +T+ G+R+++ ++ HC G KL
Sbjct: 256 Y-DSTHQVDEVDESGYNSCSSSNSIKNYQDGNSKIELTSPGKRYFLCPISGHCAGGMKLQ 432
Query: 146 ITVQPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSVPRRS 190
I V PS +P + +P P + +S P+ S
Sbjct: 433 INVAAASGTPPTTPSGTPPTTPSNPSPSPPSESGSTNTTSPPKPS 567
>TC221987 nodulin
Length = 844
Score = 80.9 bits (198), Expect = 4e-16
Identities = 51/164 (31%), Positives = 77/164 (46%), Gaps = 5/164 (3%)
Frame = +1
Query: 35 FLFAFIATIFSTMAVAKDFVVGDEKGWTTLFD----YQTWTANKVFRLGDTLTFNYVGGK 90
FLF + +++ A +VVG + W W ++ F++GDTL F Y
Sbjct: 46 FLFMLSMWLLISISEAAKYVVGGSETWKFPLSKPDSLSHWASSHRFKIGDTLIFKYDERT 225
Query: 91 DNVVRVNGSDFKSCSVPLTAPVL-TSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQ 149
++V VN +D++ C+ VL G K+++T G R +IS HC+ G KL + V
Sbjct: 226 ESVHEVNETDYEQCNTVGKEHVLFNDGNTKVMLTKSGFRHFISGNQSHCQMGLKLMVVVM 405
Query: 150 PKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSVPRRSLLP 193
+ SPSPS +P PS +P P+ S P S LP
Sbjct: 406 SNNTKKKLIHSPSPSSPSPSPSPSPSPSPSPSPSLSSPSPSPLP 537
>TC219353 weakly similar to UP|Q8L555 (Q8L555) Uclacyanin 3-like protein-like
protein, partial (44%)
Length = 713
Score = 78.2 bits (191), Expect = 2e-15
Identities = 45/158 (28%), Positives = 74/158 (46%), Gaps = 1/158 (0%)
Frame = +2
Query: 26 STMALSRALFLFAFIATIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFN 85
+ M + +F +F+A + + V G GW T + Q+W ++++F +GD+L F
Sbjct: 47 ANMGVPELMFRVSFMAVLIKLASATNYIVGGPSGGWDTNSNLQSWASSQIFSVGDSLVFQ 226
Query: 86 YVGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLF 145
Y D VV V +D+ SC G I +T G+R++I HC G K+
Sbjct: 227 YPPNHD-VVEVTKADYDSCQPTNPIQSYNDGATTIPLTLPGKRYFICGTIGHCSQGMKVE 403
Query: 146 I-TVQPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWP 182
I T+ + +P SP S + +PE S++P P
Sbjct: 404 IDTLASATNSVTPAASPEDSTTSPAESPEVIISSSPSP 517
>TC214395 weakly similar to UP|Q949E8 (Q949E8) Uclacyanin 3-like protein,
partial (44%)
Length = 652
Score = 77.4 bits (189), Expect = 4e-15
Identities = 47/140 (33%), Positives = 67/140 (47%), Gaps = 5/140 (3%)
Frame = +2
Query: 52 DFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVGGKDNVVRVNGSDFKSCSVPLTAP 111
D VGD GW +Y TW + K FR+GD L F Y V V+ S + SCS
Sbjct: 11 DHEVGDTGGWALGVNYNTWASGKTFRIGDNLVFKY-DSTHQVDEVDESGYNSCSSSNIIK 187
Query: 112 VLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITV----QPKQDGWSPVPSPS-PSPS 166
G KI +T+ G+R+++ ++ HC G KL I V P +P +PS PSPS
Sbjct: 188 NYKDGNTKIELTSTGKRYFLCPISGHCAGGMKLQINVVAGTPPTTPSGTPPTTPSNPSPS 367
Query: 167 LDLVTPEAPPSNAPWPASSV 186
+ ++ P P+ +V
Sbjct: 368 PPSDSGSTNTTSPPKPSGAV 427
>TC218257
Length = 971
Score = 75.9 bits (185), Expect = 1e-14
Identities = 41/128 (32%), Positives = 67/128 (52%), Gaps = 1/128 (0%)
Frame = +3
Query: 54 VVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVGGKDNVVRVNG-SDFKSCSVPLTAPV 112
VVG ++GW D +W+A++VFR+GD + Y + V + ++++C+V V
Sbjct: 297 VVGADRGWDQTSDLVSWSASRVFRVGDQIWLTYSVAQGLVAELKSREEYEACNVSNPINV 476
Query: 113 LTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQPKQDGWSPVPSPSPSPSLDLVTP 172
T G I + + G R+++SS ++C+NG KL + V PK D + PS S D
Sbjct: 477 YTEGLHTIPLESEGMRYFVSSEPENCKNGLKLHVEVLPKAD--ERITEPSTSTLTDEAVA 650
Query: 173 EAPPSNAP 180
PS +P
Sbjct: 651 PTTPSGSP 674
>TC204675 homologue to UP|Q8GV76 (Q8GV76) Auxin efflux carrier protein, partial
(40%)
Length = 2164
Score = 75.1 bits (183), Expect = 2e-14
Identities = 40/92 (43%), Positives = 52/92 (56%)
Frame = +3
Query: 55 VGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVGGKDNVVRVNGSDFKSCSVPLTAPVLT 114
VGD GWT F+ W K+FR GDTL FNY G NVV VN + + SC P A V
Sbjct: 1737 VGDSGGWT--FNTVAWPKGKLFRAGDTLAFNYSPGTHNVVAVNKAGYDSCKTPRGAKVYK 1910
Query: 115 SGQDKIIITTYGRRWYISSVTDHCENGQKLFI 146
SG D+I + G+ ++I + CE+G K+ I
Sbjct: 1911 SGTDQIRLAK-GQNYFICNYVGPCESGMKIAI 2003
>CD404820 weakly similar to SP|Q41001|BCP_P Blue copper protein precursor.
[Garden pea] {Pisum sativum}, partial (22%)
Length = 632
Score = 75.1 bits (183), Expect = 2e-14
Identities = 43/132 (32%), Positives = 63/132 (47%), Gaps = 4/132 (3%)
Frame = -3
Query: 53 FVVGDEKGWTTLFD---YQTWTANKVFRLGDTLTFNYVGGKDNVVRVNGSDFKSC-SVPL 108
+VVGD GW D Y W ++K F +GDTL+F + G NV+ V+ + SC S
Sbjct: 615 YVVGDGTGWRVPQDASTYHNWASDKNFTVGDTLSFIFQTGLHNVIEVSEESYNSCSSANP 436
Query: 109 TAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQPKQDGWSPVPSPSPSPSLD 168
+G + + G +YI S +HC NGQ+L I+V + P + +P P
Sbjct: 435 IGTTYNTGPANVTLNRGGEHYYICSFGNHCNNGQRLAISVSGSSTAFPPATTTAPPP--- 265
Query: 169 LVTPEAPPSNAP 180
P S+AP
Sbjct: 264 ------PSSSAP 247
>TC204881 weakly similar to UP|ENL1_ARATH (Q9SK27) Early nodulin-like protein
1 precursor (Phytocyanin-like protein), partial (65%)
Length = 968
Score = 74.7 bits (182), Expect = 3e-14
Identities = 47/144 (32%), Positives = 69/144 (47%), Gaps = 5/144 (3%)
Frame = +3
Query: 28 MALSRALFLFAFIATIFSTMAVAKDFVVGDE-KGW----TTLFDYQTWTANKVFRLGDTL 82
MA S A LF F+ FS AK+ +VG + W + W FR+GD L
Sbjct: 93 MAGSSASLLFLFLLFGFSA---AKELLVGGKIDAWKIPSSESDTLNQWAERSRFRVGDHL 263
Query: 83 TFNYVGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQ 142
+ Y GKD+V+ V D+ +CS G K+ + G ++IS HCE GQ
Sbjct: 264 VWKYESGKDSVLEVTREDYANCSTSNPIKEYNDGNTKVKLEHPGPFYFISGSKGHCEKGQ 443
Query: 143 KLFITVQPKQDGWSPVPSPSPSPS 166
KL + V + ++ + SP+P+PS
Sbjct: 444 KLIVVVMSPRHTFTAIISPAPTPS 515
>TC226352 weakly similar to UP|Q9M510 (Q9M510) Dicyanin, partial (27%)
Length = 795
Score = 74.3 bits (181), Expect = 3e-14
Identities = 43/130 (33%), Positives = 62/130 (47%), Gaps = 3/130 (2%)
Frame = +1
Query: 53 FVVGDEKGWTTLFD---YQTWTANKVFRLGDTLTFNYVGGKDNVVRVNGSDFKSCSVPLT 109
+ VG+ GW + Y W + K F++GD L FNY NV V +++ SCS
Sbjct: 163 YTVGETAGWIVPGNASFYPAWASAKNFKVGDILVFNYPSNAHNVEEVTKANYDSCSSASP 342
Query: 110 APVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQPKQDGWSPVPSPSPSPSLDL 169
T+ ++ ++ G +YI + HC GQKL I V +P SPSPS
Sbjct: 343 IATFTTPPARVPLSKSGEHYYICGIPGHCLGGQKLSINVTGGSTATAPTTPSSPSPS-GA 519
Query: 170 VTPEAPPSNA 179
V+P PP N+
Sbjct: 520 VSP--PPQNS 543
Score = 35.0 bits (79), Expect = 0.023
Identities = 20/52 (38%), Positives = 26/52 (49%), Gaps = 3/52 (5%)
Frame = +1
Query: 137 HCENGQKLFITVQPKQDGWSPVPSPSP---SPSLDLVTPEAPPSNAPWPASS 185
HC GQKL I V S PSP+P +P+ +P +PP + P P S
Sbjct: 1 HCSGGQKLAINVNRAS---SSTPSPAPEAQAPAPKATSPISPPRSTPAPGPS 147
>TC227855 weakly similar to GB|AAP88350.1|32815931|BT009716 At3g20570
{Arabidopsis thaliana;} , partial (44%)
Length = 988
Score = 73.9 bits (180), Expect = 5e-14
Identities = 47/184 (25%), Positives = 79/184 (42%), Gaps = 20/184 (10%)
Frame = +1
Query: 24 STSTMALSRALFLFAFIATIFSTMAVAK-DFVVGDEKGWTTLFD-----YQTWTANKVFR 77
+T + ++A+ F ++ + A DFVVG +KGW+ D + W F+
Sbjct: 67 ATFILRSNKAVHAFGWLCLLLMVHKGASYDFVVGGQKGWSVPNDPSFNPFNQWAEKSRFQ 246
Query: 78 LGDTLTFNYVGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDH 137
+GD+L FNY G+D+V+ V D+ SC++ + G + G ++IS D+
Sbjct: 247 IGDSLVFNYQSGQDSVLYVKSEDYASCNIDSPYAKYSDGHTVYKLNQSGPHFFISGNKDN 426
Query: 138 CENGQKLFI--------------TVQPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPA 183
C +KL + T P S PS +P+ +P + + P P
Sbjct: 427 CNKNEKLTVIVLADRNKNTNQTTTASPPSPQTSSSPSAAPTGQDQGQSPTSDTNQTPSPV 606
Query: 184 SSVP 187
S P
Sbjct: 607 SEPP 618
>TC225574 similar to UP|Q9ZRV5 (Q9ZRV5) Basic blue copper protein, partial
(83%)
Length = 868
Score = 73.2 bits (178), Expect = 8e-14
Identities = 44/130 (33%), Positives = 64/130 (48%), Gaps = 8/130 (6%)
Frame = +1
Query: 25 TSTMALSRALFLFAFIATIFSTMAV--------AKDFVVGDEKGWTTLFDYQTWTANKVF 76
T+ M+ R + T+ S + + A + VG GWT F+ W K F
Sbjct: 82 TTRMSQGRGSASLPIVVTVVSLLCLLVLLERADAATYTVGGPGGWT--FNTNAWPKGKRF 255
Query: 77 RLGDTLTFNYVGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTD 136
R GD L FNY NVV V+ S + SC P A V +SG+D+I + G+ ++I +
Sbjct: 256 RAGDILIFNYDSTTHNVVAVDRSGYNSCKTPGGAKVFSSGKDQIKLAR-GQNYFICNYPG 432
Query: 137 HCENGQKLFI 146
HCE+G K+ I
Sbjct: 433 HCESGMKVAI 462
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.134 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,597,292
Number of Sequences: 63676
Number of extensions: 156870
Number of successful extensions: 4023
Number of sequences better than 10.0: 389
Number of HSP's better than 10.0 without gapping: 2640
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3673
length of query: 203
length of database: 12,639,632
effective HSP length: 93
effective length of query: 110
effective length of database: 6,717,764
effective search space: 738954040
effective search space used: 738954040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC137822.5


