

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137822.15 + phase: 0
(279 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
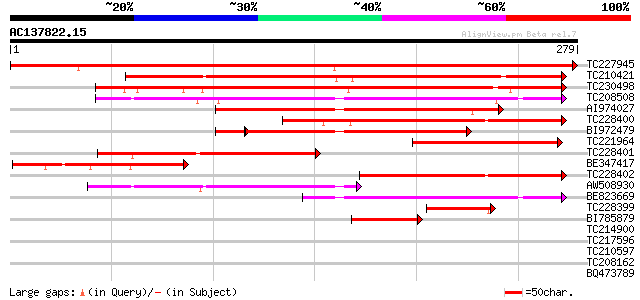
Score E
Sequences producing significant alignments: (bits) Value
TC227945 similar to GB|AAO24559.1|27808558|BT003127 At5g39890 {A... 446 e-126
TC210421 similar to GB|AAO24559.1|27808558|BT003127 At5g39890 {A... 213 6e-56
TC230498 similar to UP|Q9LPR3 (Q9LPR3) F15H18.4, partial (9%) 213 8e-56
TC208508 similar to UP|Q9SJI9 (Q9SJI9) Expressed protein, partia... 174 5e-44
AI974027 similar to GP|20197959|gb expressed protein {Arabidopsi... 154 6e-38
TC228400 similar to GB|AAO24559.1|27808558|BT003127 At5g39890 {A... 137 7e-33
BI972479 similar to GP|20197959|gb expressed protein {Arabidopsi... 136 9e-33
TC221964 weakly similar to GB|AAO24559.1|27808558|BT003127 At5g3... 133 1e-31
TC228401 weakly similar to GB|AAO24559.1|27808558|BT003127 At5g3... 119 1e-27
BE347417 119 2e-27
TC228402 similar to GB|AAO24559.1|27808558|BT003127 At5g39890 {A... 103 9e-23
AW508930 100 1e-21
BE823669 98 5e-21
TC228399 similar to GB|AAO24559.1|27808558|BT003127 At5g39890 {A... 49 2e-06
BI785879 42 3e-04
TC214900 32 0.42
TC217596 weakly similar to UP|Q8K357 (Q8K357) Procr protein, par... 30 1.2
TC210597 weakly similar to UP|Q9M3Y4 (Q9M3Y4) Germin-like protei... 29 2.1
TC208162 similar to GB|AAR23711.1|38564264|BT010741 At1g55090 {A... 28 6.0
BQ473789 similar to GP|26106026|dbj mannose-binding lectin-assoc... 28 6.0
>TC227945 similar to GB|AAO24559.1|27808558|BT003127 At5g39890 {Arabidopsis
thaliana;} , partial (47%)
Length = 1772
Score = 446 bits (1147), Expect = e-126
Identities = 215/281 (76%), Positives = 241/281 (85%), Gaps = 2/281 (0%)
Frame = +3
Query: 1 MVEQGRERVGHVNKVCYVKRVIAKKRNKPYNH-RRVKKHVPKALQELFDSCKQTFKGPGT 59
+VE+GRER GHVNKV YV+RVIAKK+ + Y RR + V K L +LFDSC++ FKGPGT
Sbjct: 357 LVERGRERAGHVNKVGYVRRVIAKKKKQLYRRVRRPELSVSKTLHQLFDSCREAFKGPGT 536
Query: 60 VPSPRDVHKLCHILDNMKPEDVGLSRDLQFFKPGNIIKENQRVTYTTVYKCDNFSLCIFF 119
VPSP+DV +L HILDNMKPEDVGLSRDLQFFKPGNI+KENQRVTYTTVYKCDNFSLCIFF
Sbjct: 537 VPSPQDVKRLTHILDNMKPEDVGLSRDLQFFKPGNIVKENQRVTYTTVYKCDNFSLCIFF 716
Query: 120 LPERGVIPLHNHPGMTVFSKLLLGQMHIKSYDWVDHEAS-HNLLQPSSKLRLAKLKANKT 178
+PE GVIPLHNHP MTVFSKLLLG MHIKSYDWVD EAS N+LQP S+LRLA LK +K
Sbjct: 717 IPEGGVIPLHNHPDMTVFSKLLLGLMHIKSYDWVDPEASDDNMLQPQSQLRLAMLKVDKV 896
Query: 179 FTAPCDTSVLYPTTGGNIHEFTAITPCAVLDVIGPPYSKEDGRDCSYYKDYPYNAFPNEE 238
FT+ CDTSVLYPTTGGNIHEFTAITPCAVLDVIGPPYSKEDGRDCSYY+D+PY FPNE
Sbjct: 897 FTSSCDTSVLYPTTGGNIHEFTAITPCAVLDVIGPPYSKEDGRDCSYYRDHPYTCFPNER 1076
Query: 239 KIGEVKDKDDSYGLLEEIDMPENCQMDGIEYLGPPIDDTMF 279
IGE K+++DSY LEEI+MPEN +M+G+EYLGP MF
Sbjct: 1077IIGEAKEENDSYTWLEEIEMPENSEMNGVEYLGPTPS*CMF 1199
>TC210421 similar to GB|AAO24559.1|27808558|BT003127 At5g39890 {Arabidopsis
thaliana;} , partial (47%)
Length = 816
Score = 213 bits (543), Expect = 6e-56
Identities = 108/224 (48%), Positives = 148/224 (65%), Gaps = 7/224 (3%)
Frame = +2
Query: 58 GTVPSPRDVHKLCHILDNMKPEDVGLSRDLQFFKPGNIIKENQRVTYTTVYKCDNFSLCI 117
G SP+++ L +L +K EDVGL ++ FF N + ++TY +Y+C FS+ I
Sbjct: 8 GLFLSPQNIEMLLSVLGEIKQEDVGLKPEMAFFSSNNP-RRTPKITYLHIYECQQFSMGI 184
Query: 118 FFLPERGVIPLHNHPGMTVFSKLLLGQMHIKSYDWVDHEASH--NLLQPSSK-----LRL 170
F LP GVIPLHNHPGMTVFSKLL G MHIKSYDWV H +++PSS+ +RL
Sbjct: 185 FCLPPSGVIPLHNHPGMTVFSKLLFGTMHIKSYDWVVDLPPHMPTIVKPSSETQTPDMRL 364
Query: 171 AKLKANKTFTAPCDTSVLYPTTGGNIHEFTAITPCAVLDVIGPPYSKEDGRDCSYYKDYP 230
AK+K + F APCD S+LYP GGN+H FTA+T CAVLDV+GPPYS DGR C+YY+++P
Sbjct: 365 AKVKVDADFNAPCDPSILYPADGGNMHWFTAVTACAVLDVLGPPYSDPDGRHCTYYQNFP 544
Query: 231 YNAFPNEEKIGEVKDKDDSYGLLEEIDMPENCQMDGIEYLGPPI 274
++++ + I E ++ +Y L+E + PEN ++ Y GP I
Sbjct: 545 FSSYSDGLSIPE--EERTAYEWLQEKEKPENLKVVVKMYSGPKI 670
>TC230498 similar to UP|Q9LPR3 (Q9LPR3) F15H18.4, partial (9%)
Length = 1137
Score = 213 bits (542), Expect = 8e-56
Identities = 112/247 (45%), Positives = 154/247 (62%), Gaps = 15/247 (6%)
Frame = +2
Query: 43 LQELFDSCKQTFK--GPGTVP-SPRDVHKLCHILDNMKPEDVGLS-------RDLQFFKP 92
+Q L+D CK G GT P S + + KL ILD ++P DVGL R L FF
Sbjct: 68 VQALYDHCKTILSPSGSGTAPPSSQALMKLSSILDTIQPADVGLKEETADDDRGLGFFGA 247
Query: 93 G---NIIKENQRVTYTTVYKCDNFSLCIFFLPERGVIPLHNHPGMTVFSKLLLGQMHIKS 149
+ + Q +TY +++CD+F++CIF P VIPLH+HPGMTVFSKLL G +H+K+
Sbjct: 248 NALSRVTRWAQPITYVDIHECDSFTMCIFCFPTSSVIPLHDHPGMTVFSKLLYGSLHVKA 427
Query: 150 YDWVDHEASHNLLQPS-SKLRLAKLKANKTFTAPCDTSVLYPTTGGNIHEFTAITPCAVL 208
YDWV+ +P +++RLAKL +K APCDTSVLYP GGN+H FTA+TPCA+L
Sbjct: 428 YDWVEPPCIIESKEPGYAQVRLAKLAVDKVLNAPCDTSVLYPKHGGNLHCFTAVTPCAML 607
Query: 209 DVIGPPYSKEDGRDCSYYKDYPYNAFPNEEKIGEVKD-KDDSYGLLEEIDMPENCQMDGI 267
D++ PPY +E+GR C+YY DYPY+AF ++D +++ Y L E++ P + M
Sbjct: 608 DILTPPYREEEGRRCTYYHDYPYSAFSAAN--APIRDGEEEEYAWLTELESPSDLYMRQG 781
Query: 268 EYLGPPI 274
Y GP I
Sbjct: 782 VYAGPAI 802
>TC208508 similar to UP|Q9SJI9 (Q9SJI9) Expressed protein, partial (57%)
Length = 1135
Score = 174 bits (440), Expect = 5e-44
Identities = 99/248 (39%), Positives = 144/248 (57%), Gaps = 16/248 (6%)
Frame = +1
Query: 43 LQELFDSCKQTFKGPGTVPSPRDVHKLCHILDNMKPEDVGLSRDLQFFK--PGNIIKENQ 100
+Q+L+D+CK + G + S + K+ +LD +KP +VGL ++ Q + G++ N
Sbjct: 271 VQKLYDTCKASLSPEGPI-SEEALEKVRTLLDELKPSNVGLEQEAQLVRGWKGSLNGTNG 447
Query: 101 R------------VTYTTVYKCDNFSLCIFFLPERGVIPLHNHPGMTVFSKLLLGQMHIK 148
+ + Y +++CD FS+ IF + VIPLHNHPGMTV SKLL G + ++
Sbjct: 448 KKGRNGSYQYPPSIKYIHLHECDKFSMGIFCMSPGSVIPLHNHPGMTVLSKLLYGSLLVR 627
Query: 149 SYDWVDHEASHNLLQPSSKLRLAKLKANKTFTAPCDTSVLYPTTGGNIHEFTAITPCAVL 208
SYDW+D + S+ R AKL + +APC+T+VLYP+ GGNIH F A+TPCA+
Sbjct: 628 SYDWLDLPGPDD----PSQARPAKLVKDCQMSAPCNTTVLYPSKGGNIHCFKALTPCALF 795
Query: 209 DVIGPPYSKEDGRDCSYYKDYPYNAFPNEE--KIGEVKDKDDSYGLLEEIDMPENCQMDG 266
DV+ PPYS EDGR CSY++ P E ++ VK + ++ LEEI PEN +
Sbjct: 796 DVLSPPYSSEDGRHCSYFRKSTRKDLPGVELDQLSGVKPSEITW--LEEIQAPENLVVRR 969
Query: 267 IEYLGPPI 274
Y GP I
Sbjct: 970 GVYKGPTI 993
>AI974027 similar to GP|20197959|gb expressed protein {Arabidopsis thaliana},
partial (51%)
Length = 431
Score = 154 bits (388), Expect = 6e-38
Identities = 74/147 (50%), Positives = 98/147 (66%), Gaps = 5/147 (3%)
Frame = +3
Query: 102 VTYTTVYKCDNFSLCIFFLPERGVIPLHNHPGMTVFSKLLLGQMHIKSYDWVDHEASHNL 161
+ Y +++CD+FS+ IF +P +IPLHNHPGMTV SKLL G M++KSYDW+D S++
Sbjct: 3 IKYLHLHECDSFSIGIFCMPPSSIIPLHNHPGMTVLSKLLYGSMYVKSYDWIDAPGSND- 179
Query: 162 LQPSSKLRLAKLKANKTFTAPCDTSVLYPTTGGNIHEFTAITPCAVLDVIGPPYSKEDGR 221
S+ R AKL + TAP T+VLYPT+GGNIH F AITPCA+ D++ PPYS + GR
Sbjct: 180 ---PSEARPAKLVKDTEMTAPSPTTVLYPTSGGNIHCFRAITPCAIFDILSPPYSSDHGR 350
Query: 222 DCSYY-----KDYPYNAFPNEEKIGEV 243
C+Y+ KD P N N + EV
Sbjct: 351 HCTYFRRSQRKDLPVNVQLNGVTVSEV 431
>TC228400 similar to GB|AAO24559.1|27808558|BT003127 At5g39890 {Arabidopsis
thaliana;} , partial (41%)
Length = 797
Score = 137 bits (344), Expect = 7e-33
Identities = 69/146 (47%), Positives = 100/146 (68%), Gaps = 6/146 (4%)
Frame = +3
Query: 135 TVFSKLLLGQMHIKSYDWV--DHEASHNLLQPSS----KLRLAKLKANKTFTAPCDTSVL 188
TVFSKLL G MHIKSYDWV S ++PS ++RLAK+K + FTAPC+ S+L
Sbjct: 18 TVFSKLLFGTMHIKSYDWVVDSPPESPTTIKPSENQGPEMRLAKVKVDADFTAPCNPSIL 197
Query: 189 YPTTGGNIHEFTAITPCAVLDVIGPPYSKEDGRDCSYYKDYPYNAFPNEEKIGEVKDKDD 248
YP GGN+H FTA+T CAVLDV+GPPYS +GR C+YY ++P++ F + + + +++ +
Sbjct: 198 YPEDGGNLHCFTAVTACAVLDVLGPPYSDAEGRHCTYYHNFPFSNF-SADGLSIPEEEKN 374
Query: 249 SYGLLEEIDMPENCQMDGIEYLGPPI 274
+Y L+E + E+ +++G Y GP I
Sbjct: 375 AYEWLQEREELEDLEVNGKMYNGPKI 452
>BI972479 similar to GP|20197959|gb expressed protein {Arabidopsis thaliana},
partial (51%)
Length = 422
Score = 136 bits (342), Expect(2) = 9e-33
Identities = 62/111 (55%), Positives = 80/111 (71%)
Frame = +2
Query: 117 IFFLPERGVIPLHNHPGMTVFSKLLLGQMHIKSYDWVDHEASHNLLQPSSKLRLAKLKAN 176
IF +P +IPLHNHPGMTV SKLL G M++KSYDW+D S++ S+ R AKL +
Sbjct: 95 IFCMPPSSIIPLHNHPGMTVLSKLLYGSMYVKSYDWIDAPGSND----PSEARPAKLVKD 262
Query: 177 KTFTAPCDTSVLYPTTGGNIHEFTAITPCAVLDVIGPPYSKEDGRDCSYYK 227
TAP T+VLYPT+GGNIH F AITPCA+ D++ PPYS + GR C+Y++
Sbjct: 263 TEMTAPSPTTVLYPTSGGNIHCFRAITPCAIFDILSPPYSSDHGRHCTYFR 415
Score = 21.6 bits (44), Expect(2) = 9e-33
Identities = 7/17 (41%), Positives = 12/17 (70%)
Frame = +3
Query: 102 VTYTTVYKCDNFSLCIF 118
+ Y +++CD+FSL F
Sbjct: 27 IKYLHLHECDSFSLICF 77
>TC221964 weakly similar to GB|AAO24559.1|27808558|BT003127 At5g39890
{Arabidopsis thaliana;} , partial (13%)
Length = 824
Score = 133 bits (334), Expect = 1e-31
Identities = 59/74 (79%), Positives = 67/74 (89%)
Frame = +3
Query: 199 FTAITPCAVLDVIGPPYSKEDGRDCSYYKDYPYNAFPNEEKIGEVKDKDDSYGLLEEIDM 258
FTAITPCAVLDVIGPPYSKEDGRDCSYY+D+PY +FPNE IGE K+++DSY LEEI+M
Sbjct: 3 FTAITPCAVLDVIGPPYSKEDGRDCSYYRDHPYASFPNERIIGEAKEENDSYAWLEEIEM 182
Query: 259 PENCQMDGIEYLGP 272
PEN +MDGIEYLGP
Sbjct: 183 PENSEMDGIEYLGP 224
>TC228401 weakly similar to GB|AAO24559.1|27808558|BT003127 At5g39890
{Arabidopsis thaliana;} , partial (41%)
Length = 408
Score = 119 bits (299), Expect = 1e-27
Identities = 58/112 (51%), Positives = 76/112 (67%), Gaps = 2/112 (1%)
Frame = +1
Query: 44 QELFDSCKQTFKGPGT--VPSPRDVHKLCHILDNMKPEDVGLSRDLQFFKPGNIIKENQR 101
++L ++CK F GT VP D+ +L +LD +KPEDVGL D+ +F+ + + R
Sbjct: 7 EKLLETCKVVFASAGTGFVPPHEDIDELQSVLDGIKPEDVGLRPDMPYFRT-SATQRVPR 183
Query: 102 VTYTTVYKCDNFSLCIFFLPERGVIPLHNHPGMTVFSKLLLGQMHIKSYDWV 153
+TY +Y+C+ FS+ IF LP GVIPLHNHPGMTVFSK G MHIKSYDWV
Sbjct: 184 ITYLHIYECEKFSMGIFCLPPSGVIPLHNHPGMTVFSKPTFGTMHIKSYDWV 339
>BE347417
Length = 480
Score = 119 bits (298), Expect = 2e-27
Identities = 66/97 (68%), Positives = 74/97 (76%), Gaps = 10/97 (10%)
Frame = +1
Query: 2 VEQGRERVGHVNKVC-YVKRVIAKKRNKPYNHRRVKKH--------VPKALQELFDSCKQ 52
+EQG +RVGHVNKV Y KRVI KKR KPY+H + H VPKALQELF SC++
Sbjct: 193 MEQGGDRVGHVNKVIGYGKRVIVKKR-KPYHHHHRRIHNKKTVPNKVPKALQELFVSCRE 369
Query: 53 TFKGPG-TVPSPRDVHKLCHILDNMKPEDVGLSRDLQ 88
TFKGPG TVPSP+DV KLCHILD+MKPEDVGL DLQ
Sbjct: 370 TFKGPGGTVPSPQDVXKLCHILDSMKPEDVGLRSDLQ 480
>TC228402 similar to GB|AAO24559.1|27808558|BT003127 At5g39890 {Arabidopsis
thaliana;} , partial (31%)
Length = 546
Score = 103 bits (257), Expect = 9e-23
Identities = 47/102 (46%), Positives = 72/102 (70%)
Frame = +1
Query: 173 LKANKTFTAPCDTSVLYPTTGGNIHEFTAITPCAVLDVIGPPYSKEDGRDCSYYKDYPYN 232
+K + FTAPC+ S+LYP GGN+H FTA+T CAVLDV+GPPYS +GR C+YY D+P++
Sbjct: 1 VKVDADFTAPCNPSILYPEDGGNLHCFTAVTACAVLDVLGPPYSDAEGRHCTYYHDFPFS 180
Query: 233 AFPNEEKIGEVKDKDDSYGLLEEIDMPENCQMDGIEYLGPPI 274
F + + + +++ ++Y L+E D E+ +++G Y GP I
Sbjct: 181 NF-SVDGLSIPEEEKNAYEWLQERDELEDLEVNGKMYNGPKI 303
>AW508930
Length = 444
Score = 99.8 bits (247), Expect = 1e-21
Identities = 58/144 (40%), Positives = 83/144 (57%), Gaps = 9/144 (6%)
Frame = +1
Query: 39 VPKALQELFDSCKQTFKGPGTVPSPRDVHKLCHILDNMKPEDVGLSRDLQFFKP------ 92
+P +Q L+ CK +F G V S + K+C L+ +KP DVGL ++ Q +
Sbjct: 25 MPYYIQRLYRLCKASFSPNGPV-SEEAIAKVCEKLEKIKPSDVGLEQEAQVVRNWSSQMP 201
Query: 93 ---GNIIKENQRVTYTTVYKCDNFSLCIFFLPERGVIPLHNHPGMTVFSKLLLGQMHIKS 149
GN ++ + Y +Y+ D+FS+ IF +P VIPLHNHPGMTV SKLL G +++KS
Sbjct: 202 ECNGNH-QQPPPIKYLHLYEDDSFSIGIFCMPPSSVIPLHNHPGMTVLSKLLYGSVYVKS 378
Query: 150 YDWVDHEASHNLLQPSSKLRLAKL 173
YDW+D + SS+ R AKL
Sbjct: 379 YDWIDFPGPTD----SSEARAAKL 438
>BE823669
Length = 641
Score = 97.8 bits (242), Expect = 5e-21
Identities = 53/130 (40%), Positives = 73/130 (55%)
Frame = -3
Query: 145 MHIKSYDWVDHEASHNLLQPSSKLRLAKLKANKTFTAPCDTSVLYPTTGGNIHEFTAITP 204
+++KSYD +D + S + R AKL + TAP T+VLYPT GGNIH F AITP
Sbjct: 636 VYVKSYDXIDFXGXXD----SXEARAAKLVKDTEMTAPTATTVLYPTXGGNIHTFRAITP 469
Query: 205 CAVLDVIGPPYSKEDGRDCSYYKDYPYNAFPNEEKIGEVKDKDDSYGLLEEIDMPENCQM 264
CA+ DV+ PPYS E GR C+Y++ P ++ V D ++ LEE P++ +
Sbjct: 468 CAIFDVLSPPYSSEHGRHCTYFRKSQRKDLPGNLQLNGVTVSDVTW--LEEFQPPDDFVI 295
Query: 265 DGIEYLGPPI 274
Y GP I
Sbjct: 294 RRGIYKGPVI 265
>TC228399 similar to GB|AAO24559.1|27808558|BT003127 At5g39890 {Arabidopsis
thaliana;} , partial (17%)
Length = 458
Score = 49.3 bits (116), Expect = 2e-06
Identities = 22/41 (53%), Positives = 28/41 (67%), Gaps = 7/41 (17%)
Frame = +2
Query: 206 AVLDVIGPPYSKEDGRDCSYYKDYPYNAF-------PNEEK 239
AVLDV+GPPYS +GR C+YY D+P++ F P EEK
Sbjct: 17 AVLDVLGPPYSDAEGRHCTYYHDFPFSNFSVDGLSIPEEEK 139
>BI785879
Length = 421
Score = 42.0 bits (97), Expect = 3e-04
Identities = 19/35 (54%), Positives = 26/35 (74%)
Frame = +3
Query: 169 RLAKLKANKTFTAPCDTSVLYPTTGGNIHEFTAIT 203
R AKL + +APC+T+VL+P+ GGNIH F A+T
Sbjct: 315 RPAKLMKDCLMSAPCNTTVLHPSKGGNIHCFKALT 419
>TC214900
Length = 802
Score = 31.6 bits (70), Expect = 0.42
Identities = 14/40 (35%), Positives = 25/40 (62%), Gaps = 1/40 (2%)
Frame = -3
Query: 14 KVCYVKRVIAKKRNKP-YNHRRVKKHVPKALQELFDSCKQ 52
++C ++++ +K+N P Y ++ KKH K L L +S KQ
Sbjct: 527 QICIQRKILIRKKNDP*YQKKKKKKHYLKVLVNLRNSYKQ 408
>TC217596 weakly similar to UP|Q8K357 (Q8K357) Procr protein, partial (9%)
Length = 988
Score = 30.0 bits (66), Expect = 1.2
Identities = 18/65 (27%), Positives = 28/65 (42%), Gaps = 4/65 (6%)
Frame = -2
Query: 75 NMKPEDVGLSRDLQFFKPGNI----IKENQRVTYTTVYKCDNFSLCIFFLPERGVIPLHN 130
N K +D+ LS D QF K K R +T+Y N+ + + G++ H+
Sbjct: 777 NSKAKDLTLSLDKQFIKLKTSQI**SKHTGRYNASTLYLVGNYGMILIHTASTGILITHH 598
Query: 131 HPGMT 135
H T
Sbjct: 597 HNSCT 583
>TC210597 weakly similar to UP|Q9M3Y4 (Q9M3Y4) Germin-like protein, partial
(87%)
Length = 830
Score = 29.3 bits (64), Expect = 2.1
Identities = 27/106 (25%), Positives = 40/106 (37%), Gaps = 7/106 (6%)
Frame = +2
Query: 48 DSCKQTFKGPGTVPSPRDVHKLCHILDNMKPEDVGLSRDLQF-FKPGN---IIKENQRVT 103
D C KGP SP H KP S D F PGN + K
Sbjct: 83 DFCVADLKGPD---SPTGYH--------CKPPKTVTSHDFVFHLGPGNTSNVFKSAITSA 229
Query: 104 YTTVYKCDN---FSLCIFFLPERGVIPLHNHPGMTVFSKLLLGQMH 146
+ + N S+ + + GV+P+H HPG ++ G+++
Sbjct: 230 FVKDFPAVNGLSLSVARIDIAQGGVVPMHTHPGANEIVMMVPGEIN 367
>TC208162 similar to GB|AAR23711.1|38564264|BT010741 At1g55090 {Arabidopsis
thaliana;} , partial (22%)
Length = 779
Score = 27.7 bits (60), Expect = 6.0
Identities = 18/57 (31%), Positives = 26/57 (45%), Gaps = 2/57 (3%)
Frame = +2
Query: 195 NIHEFTAITPCAVLDVIGPPYSKEDGRDCSYYKDYPYNAFPNEEKIGE--VKDKDDS 249
N H+ T +TP + P ++ D R Y +PY +E + E VKD DS
Sbjct: 353 NRHKMTVLTPSYHAESYSPEDNRFDLRQFLYNARWPYQFRKIDELVSELDVKDVKDS 523
>BQ473789 similar to GP|26106026|dbj mannose-binding lectin-associated serine
protease {Lethenteron japonicum}, partial (1%)
Length = 421
Score = 27.7 bits (60), Expect = 6.0
Identities = 27/113 (23%), Positives = 46/113 (39%), Gaps = 17/113 (15%)
Frame = +2
Query: 20 RVIAKKRNKPYNHRRVKKHVPKALQELFDSCKQTFKGPGTVPSPRDVHKLCHILDNMKPE 79
R+I K P R K +Q L+D+ F G +P+ + +H L +LD ++
Sbjct: 104 RLIITKLYYPKTMERSK------IQVLYDASHAVFSQEG-LPTFQQIHYLKTLLDKIEAI 262
Query: 80 DVGLSRDLQFFKP-------GNIIKENQR----------VTYTTVYKCDNFSL 115
DVG+ P + N + +TY +++CD FS+
Sbjct: 263 DVGVDESGLCDSPTSDATVDSSSSSSNSKGLLCGHGFSEITYIHIHECDYFSM 421
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.140 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,994,102
Number of Sequences: 63676
Number of extensions: 222010
Number of successful extensions: 1053
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 1035
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1036
length of query: 279
length of database: 12,639,632
effective HSP length: 96
effective length of query: 183
effective length of database: 6,526,736
effective search space: 1194392688
effective search space used: 1194392688
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC137822.15


