

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137703.1 + phase: 0
(233 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
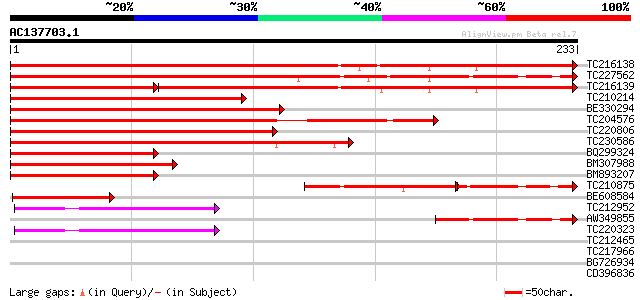
TC216138 similar to UP|LB37_ARATH (Q9FN11) LOB domain protein 37... 338 2e-93
TC227562 similar to UP|LB37_ARATH (Q9FN11) LOB domain protein 37... 319 6e-88
TC216139 similar to UP|LB37_ARATH (Q9FN11) LOB domain protein 37... 212 6e-88
TC210214 homologue to UP|LB39_ARATH (Q9SZE8) LOB domain protein ... 201 2e-52
BE330294 181 3e-46
TC204576 similar to UP|Q7XYW2 (Q7XYW2) Seed specific protein Bn1... 175 1e-44
TC220806 similar to UP|LB40_ARATH (Q9ZW96) LOB domain protein 40... 171 3e-43
TC230586 similar to UP|LB42_ARATH (Q9CA30) LOB domain protein 42... 142 2e-34
BQ299324 127 6e-30
BM307988 102 2e-22
BM893207 100 4e-22
TC210875 similar to UP|Q79FV6 (Q79FV6) PE-PGRS FAMILY PROTEIN, p... 61 2e-18
BE608584 73 1e-13
TC212952 similar to UP|LBD1_ARATH (Q9LQR0) LOB domain protein 1,... 55 4e-08
AW349855 53 1e-07
TC220323 similar to UP|LBD1_ARATH (Q9LQR0) LOB domain protein 1,... 52 2e-07
TC212465 similar to UP|LBD1_ARATH (Q9LQR0) LOB domain protein 1,... 33 0.14
TC217966 homologue to UP|Q9LX13 (Q9LX13) (3R)-hydroxymyristoyl-[... 31 0.42
BG726934 28 2.7
CD396836 28 3.6
>TC216138 similar to UP|LB37_ARATH (Q9FN11) LOB domain protein 37, partial
(50%)
Length = 1169
Score = 338 bits (866), Expect = 2e-93
Identities = 179/242 (73%), Positives = 194/242 (79%), Gaps = 9/242 (3%)
Frame = +1
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSESCILRPCLQWI+TPEAQGHATVFVAKFFGRA LMSFISNVPEPQR
Sbjct: 121 MSCNGCRVLRKGCSESCILRPCLQWIDTPEAQGHATVFVAKFFGRADLMSFISNVPEPQR 300
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPLPELTLLDAPL 120
PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGT+RP+PEL LDAP
Sbjct: 301 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTVRPMPELFGLDAPT 480
Query: 121 TATDEASEAEDGGADAMWKIRE--PNFGLASSRSSNKVCSGSKRRRSEEFVKV-PAAINL 177
TD+ASE E D W++R+ PNF SR S+K SG KR+R EE K+ A +L
Sbjct: 481 PTTDDASEGEVTCTD-KWRVRDPNPNFRFPGSR-SDKGSSGGKRKRFEELAKLQTATTDL 654
Query: 178 DLRLTPIFQQKAV------EERRHGSPSMTSEESVTTTACLETGIGDRWSHGGDRKVLNL 231
+LRLTP F Q A E R GSPSM SEES TTTACLE+GIGD ++ GDRKVLNL
Sbjct: 655 NLRLTPSFLQNAANFGCRQEIRWPGSPSMNSEESGTTTACLESGIGDHYAPDGDRKVLNL 834
Query: 232 FI 233
FI
Sbjct: 835 FI 840
>TC227562 similar to UP|LB37_ARATH (Q9FN11) LOB domain protein 37, partial
(45%)
Length = 1148
Score = 319 bits (818), Expect = 6e-88
Identities = 179/239 (74%), Positives = 186/239 (76%), Gaps = 6/239 (2%)
Frame = +1
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSESCILRPCLQWIET EAQGHATVFVAKFFGRAGLMSFISNVPE QR
Sbjct: 121 MSCNGCRVLRKGCSESCILRPCLQWIETAEAQGHATVFVAKFFGRAGLMSFISNVPENQR 300
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPLPELTLLD--- 117
PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRP+PEL LD
Sbjct: 301 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPMPELMGLDAAA 480
Query: 118 APLTATDEASEAEDGGADAMWKIREPNFG--LASSRSSNKVCSGSKRRRSEEFVKV-PAA 174
A A DE SEAE G D W+IR PN SSRSS+ G KR+RSEE K+
Sbjct: 481 AAAAAADETSEAEVGCTDT-WRIRNPNTNCRFTSSRSSSG-GGGGKRKRSEELAKLQEQQ 654
Query: 175 INLDLRLTPIFQQKAVEERRHGSPSMTSEESVTTTACLETGIGDRWSHGGDRKVLNLFI 233
NLDLRLTP+F QK E RR SPSMTSEES TTT E GD+WSH KVLNLFI
Sbjct: 655 PNLDLRLTPVFLQKE-ESRRPESPSMTSEESGTTT---ENRFGDQWSHW---KVLNLFI 810
>TC216139 similar to UP|LB37_ARATH (Q9FN11) LOB domain protein 37, partial
(50%)
Length = 996
Score = 212 bits (539), Expect(2) = 6e-88
Identities = 117/183 (63%), Positives = 133/183 (71%), Gaps = 11/183 (6%)
Frame = +3
Query: 62 ALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPLPELTLLDAPLT 121
ALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGT+RP+PEL LDAP
Sbjct: 417 ALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTVRPMPELLGLDAPTP 596
Query: 122 ATDEASEAEDGGADAMWKIREPNFGLASSR----SSNKVCSGSKRRRSEEFVKV-PAAIN 176
D+ASE E D W++R+PN +R S+K SG KR+R EE K+ A +
Sbjct: 597 TADDASEGEVTCTD-KWRVRDPNPNFPGTRFPGSRSDKAGSGGKRKRFEELAKLQTATTD 773
Query: 177 LDLRLTPIFQQKAV------EERRHGSPSMTSEESVTTTACLETGIGDRWSHGGDRKVLN 230
L+LRLTP F Q A E R GSPSM SEES TTTACLE+GIG+ ++H GDRKVLN
Sbjct: 774 LNLRLTPSFLQNAASYGCRQEIRWPGSPSMNSEESGTTTACLESGIGEYFAHDGDRKVLN 953
Query: 231 LFI 233
LFI
Sbjct: 954 LFI 962
Score = 129 bits (325), Expect(2) = 6e-88
Identities = 59/61 (96%), Positives = 60/61 (97%)
Frame = +1
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSESCILRPCLQWI+TPEAQGHATVFVAKFFGRA LMSFISNVPEPQR
Sbjct: 133 MSCNGCRVLRKGCSESCILRPCLQWIDTPEAQGHATVFVAKFFGRADLMSFISNVPEPQR 312
Query: 61 P 61
P
Sbjct: 313 P 315
>TC210214 homologue to UP|LB39_ARATH (Q9SZE8) LOB domain protein 39, partial
(40%)
Length = 424
Score = 201 bits (511), Expect = 2e-52
Identities = 93/97 (95%), Positives = 95/97 (97%)
Frame = +1
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSESC+LRPCLQWIET +AQGHATVFVAKFFGRAGLMSFISNVPE QR
Sbjct: 133 MSCNGCRVLRKGCSESCMLRPCLQWIETAQAQGHATVFVAKFFGRAGLMSFISNVPENQR 312
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAA 97
PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAA
Sbjct: 313 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAA 423
>BE330294
Length = 475
Score = 181 bits (459), Expect = 3e-46
Identities = 87/113 (76%), Positives = 94/113 (82%)
Frame = +1
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRK CSE C LR L WI+ P AQ HAT+F+AKFFGR+ LM FIS+VP R
Sbjct: 31 MSCNGCRVLRKRCSERCSLRSSLNWIDCPHAQAHATLFLAKFFGRSDLMFFISSVPLNNR 210
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPLPEL 113
PALFQSLLFEACGRTVNPVNGAVGLL +GNWHVCQAAV TVL GG LRPL E+
Sbjct: 211 PALFQSLLFEACGRTVNPVNGAVGLLLSGNWHVCQAAVHTVLAGGVLRPLSEV 369
>TC204576 similar to UP|Q7XYW2 (Q7XYW2) Seed specific protein Bn15D17A,
partial (66%)
Length = 1320
Score = 175 bits (444), Expect = 1e-44
Identities = 89/176 (50%), Positives = 122/176 (68%)
Frame = +3
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSE+C +RPCLQWI++PE+Q +ATVF+AKF+GRAGLM+ ++ PE R
Sbjct: 114 MSCNGCRVLRKGCSENCSIRPCLQWIKSPESQANATVFLAKFYGRAGLMNLVNAGPEHLR 293
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPLPELTLLDAPL 120
PA+F+SLL+EACGR VNP+ G+VGLLW+G+W +CQAAVE VL+G + P+
Sbjct: 294 PAIFRSLLYEACGRIVNPIYGSVGLLWSGSWQLCQAAVENVLKGAPITPI---------- 443
Query: 121 TATDEASEAEDGGADAMWKIREPNFGLASSRSSNKVCSGSKRRRSEEFVKVPAAIN 176
T EA+ + G + IR + S+ +SN+ + +R ++ VK P A N
Sbjct: 444 --TSEAAASGRGPPLKAYDIRHVSKDENSAAASNE--TQHQRVKTRSRVKRPLANN 599
>TC220806 similar to UP|LB40_ARATH (Q9ZW96) LOB domain protein 40, partial
(53%)
Length = 689
Score = 171 bits (433), Expect = 3e-43
Identities = 75/110 (68%), Positives = 94/110 (85%)
Frame = +2
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSE C +RPCLQWI+ PE+Q +ATVF+AKF+GRAGLM+ I+ PE R
Sbjct: 125 MSCNGCRVLRKGCSEDCSIRPCLQWIKNPESQANATVFLAKFYGRAGLMNLINAGPENLR 304
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPL 110
PA+F+SLL+EACGR VNP+ G+VGLLW+G+W +CQAAVE VL+G + P+
Sbjct: 305 PAIFRSLLYEACGRIVNPIYGSVGLLWSGSWQLCQAAVEAVLKGEPITPI 454
>TC230586 similar to UP|LB42_ARATH (Q9CA30) LOB domain protein 42, partial
(43%)
Length = 805
Score = 142 bits (357), Expect = 2e-34
Identities = 67/144 (46%), Positives = 98/144 (67%), Gaps = 3/144 (2%)
Frame = +3
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
+SCNGCRVLR+GC+ C +R LQWI +P +Q +AT+F+AKF+GR GL++ ++N P +
Sbjct: 54 LSCNGCRVLRRGCTTECPIRISLQWINSPNSQSNATLFLAKFYGRQGLLTLLTNAPSQLQ 233
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLR--PLPELTLLDA 118
A+ +SLL+EACGR VNPV G+ GLL TGNWH+C+AAV+ VL G + PL T L
Sbjct: 234 QAVLKSLLYEACGRIVNPVLGSTGLLSTGNWHLCEAAVKAVLNGAPIEQAPLDGATSLQL 413
Query: 119 PLTATDEASEAEDG-GADAMWKIR 141
++++ G+D + K++
Sbjct: 414 KACNIRHVAKSDKSCGSDTLRKVK 485
>BQ299324
Length = 421
Score = 127 bits (318), Expect = 6e-30
Identities = 59/61 (96%), Positives = 59/61 (96%)
Frame = +2
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSESCILRPCLQWIET EAQGHATVFVAKFFGRAGLMSFISNVPE QR
Sbjct: 98 MSCNGCRVLRKGCSESCILRPCLQWIETAEAQGHATVFVAKFFGRAGLMSFISNVPENQR 277
Query: 61 P 61
P
Sbjct: 278 P 280
>BM307988
Length = 455
Score = 102 bits (253), Expect = 2e-22
Identities = 46/69 (66%), Positives = 55/69 (79%)
Frame = +2
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSE C +RPCLQWI+ PE+Q +ATVF+AKF+GRAGLM+ I+ PE R
Sbjct: 122 MSCNGCRVLRKGCSEDCSIRPCLQWIKNPESQANATVFLAKFYGRAGLMNLINAGPENLR 301
Query: 61 PALFQSLLF 69
P +LF
Sbjct: 302 PGSCYFVLF 328
>BM893207
Length = 423
Score = 100 bits (250), Expect = 4e-22
Identities = 44/61 (72%), Positives = 52/61 (85%)
Frame = +2
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSE C +RPCLQWI+ PE+Q +ATVF+AKF+GRAGLM+ I+ PE R
Sbjct: 122 MSCNGCRVLRKGCSEDCSIRPCLQWIKNPESQANATVFLAKFYGRAGLMNLINAGPESLR 301
Query: 61 P 61
P
Sbjct: 302 P 304
>TC210875 similar to UP|Q79FV6 (Q79FV6) PE-PGRS FAMILY PROTEIN, partial (3%)
Length = 649
Score = 61.2 bits (147), Expect(2) = 2e-18
Identities = 35/65 (53%), Positives = 41/65 (62%), Gaps = 1/65 (1%)
Frame = +3
Query: 122 ATDEASEAEDGGADAMWKIREPNFGLASSRSSNKVCSGS-KRRRSEEFVKVPAAINLDLR 180
ATDEASEAE D W+IR PN + S + C G KR+RSEE K+ A NLDLR
Sbjct: 9 ATDEASEAEVACTDT-WRIRNPNPNCRFTSSRSSGCGGGGKRKRSEELAKLQAQPNLDLR 185
Query: 181 LTPIF 185
LTP+F
Sbjct: 186 LTPVF 200
Score = 47.8 bits (112), Expect(2) = 2e-18
Identities = 31/49 (63%), Positives = 32/49 (65%)
Frame = +1
Query: 185 FQQKAVEERRHGSPSMTSEESVTTTACLETGIGDRWSHGGDRKVLNLFI 233
F QK VE RR SPSMTS ES TTT E GD+W H RKVLNLFI
Sbjct: 199 FSQK-VESRRPESPSMTSVESGTTT---ENRFGDQWGH---RKVLNLFI 324
>BE608584
Length = 127
Score = 73.2 bits (178), Expect = 1e-13
Identities = 31/42 (73%), Positives = 37/42 (87%)
Frame = +2
Query: 2 SCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFF 43
SCNGCRVLRKGC ++C LR CLQWIE+ E+Q HAT+F+AKFF
Sbjct: 2 SCNGCRVLRKGCIDTCPLRSCLQWIESLESQRHATLFLAKFF 127
>TC212952 similar to UP|LBD1_ARATH (Q9LQR0) LOB domain protein 1, partial
(63%)
Length = 436
Score = 54.7 bits (130), Expect = 4e-08
Identities = 25/84 (29%), Positives = 42/84 (49%)
Frame = +2
Query: 3 CNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQRPA 62
C C++LR+ C+E C+L P P + + FG + ++ F+ +PE QR
Sbjct: 17 CAACKILRRKCAEKCVLAPYF-----PPTEPAKFTIAHRVFGASNIIKFLQELPESQRAD 181
Query: 63 LFQSLLFEACGRTVNPVNGAVGLL 86
S+++EA R +PV G G +
Sbjct: 182AVTSMVYEASARIRDPVYGCAGAI 253
>AW349855
Length = 501
Score = 52.8 bits (125), Expect = 1e-07
Identities = 34/58 (58%), Positives = 36/58 (61%)
Frame = -3
Query: 176 NLDLRLTPIFQQKAVEERRHGSPSMTSEESVTTTACLETGIGDRWSHGGDRKVLNLFI 233
NLDL TP+F K E RR SPSMTSEES T E GD+WSH KVLNLFI
Sbjct: 442 NLDLXXTPVFLHKE-ESRRPESPSMTSEESGMPT---ENRFGDQWSHW---KVLNLFI 290
>TC220323 similar to UP|LBD1_ARATH (Q9LQR0) LOB domain protein 1, partial
(69%)
Length = 854
Score = 52.0 bits (123), Expect = 2e-07
Identities = 25/84 (29%), Positives = 40/84 (46%)
Frame = +1
Query: 3 CNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQRPA 62
C C++LR+ C E C+L P P + FG + ++ F+ +PE QR
Sbjct: 280 CAACKILRRRCVEKCVLAPYF-----PPTDPLKFTIAHRVFGASNIIKFLQELPESQRAD 444
Query: 63 LFQSLLFEACGRTVNPVNGAVGLL 86
S+++EA R +PV G G +
Sbjct: 445 AVSSMVYEANARIRDPVYGCAGAI 516
>TC212465 similar to UP|LBD1_ARATH (Q9LQR0) LOB domain protein 1, partial
(34%)
Length = 411
Score = 32.7 bits (73), Expect = 0.14
Identities = 14/53 (26%), Positives = 25/53 (46%)
Frame = +3
Query: 3 CNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNV 55
C C++LR+ C+E C+L P P + + FG + ++ F+ V
Sbjct: 216 CAACKILRRKCAEKCVLAPYF-----PPTEPAKFTIAHRVFGASNIIKFLQVV 359
>TC217966 homologue to UP|Q9LX13 (Q9LX13) (3R)-hydroxymyristoyl-[acyl carrier
protein] dehydratase-like protein (At5g10160), partial
(71%)
Length = 910
Score = 31.2 bits (69), Expect = 0.42
Identities = 23/65 (35%), Positives = 31/65 (47%)
Frame = +3
Query: 143 PNFGLASSRSSNKVCSGSKRRRSEEFVKVPAAINLDLRLTPIFQQKAVEERRHGSPSMTS 202
PNF L++S + + + KRRRS F VP + + L +TP H PS TS
Sbjct: 180 PNFHLSNSPTPDPIPCRGKRRRSPRF--VPPMLPMLLNMTPQLN*GI----PHSQPSWTS 341
Query: 203 EESVT 207
VT
Sbjct: 342 TRFVT 356
>BG726934
Length = 387
Score = 28.5 bits (62), Expect = 2.7
Identities = 16/52 (30%), Positives = 27/52 (51%)
Frame = -1
Query: 130 EDGGADAMWKIREPNFGLASSRSSNKVCSGSKRRRSEEFVKVPAAINLDLRL 181
+ GG+ ++ ++R +F A +R + + G RRR E F V +N L L
Sbjct: 219 DPGGSASVCRLRNWSFAAAKNRGAWRGFGGEARRREEGFGPV*LGLNWGLGL 64
>CD396836
Length = 549
Score = 28.1 bits (61), Expect = 3.6
Identities = 24/73 (32%), Positives = 29/73 (38%)
Frame = +3
Query: 142 EPNFGLASSRSSNKVCSGSKRRRSEEFVKVPAAINLDLRLTPIFQQKAVEERRHGSPSMT 201
E G+A ++CS RRRS + + P TP QQ RH S T
Sbjct: 231 EGRLGVARCSHKIRMCSHRTRRRSRAWKREPRTRTPSRLSTPTRQQ-----WRHSSSPTT 395
Query: 202 SEESVTTTACLET 214
SV TT L T
Sbjct: 396 LATSVGTTEDLCT 434
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.134 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,760,334
Number of Sequences: 63676
Number of extensions: 160139
Number of successful extensions: 868
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 852
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 855
length of query: 233
length of database: 12,639,632
effective HSP length: 94
effective length of query: 139
effective length of database: 6,654,088
effective search space: 924918232
effective search space used: 924918232
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC137703.1


