

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
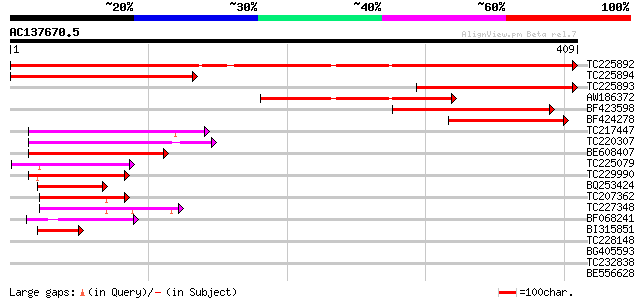
Query= AC137670.5 + phase: 0
(409 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC225892 similar to UP|ARG3_HUMAN (Q9NP61) ADP-ribosylation fact... 627 e-180
TC225894 similar to UP|Q9FIQ0 (Q9FIQ0) Zinc finger protein Glo3-... 244 5e-65
TC225893 similar to UP|Q9FIQ0 (Q9FIQ0) Zinc finger protein Glo3-... 191 7e-49
AW186372 180 9e-46
BF423598 weakly similar to GP|9758511|dbj| zinc finger protein G... 138 4e-33
BF424278 138 5e-33
TC217447 similar to GB|AAM65362.1|21655301|AY103310 At2g37550/F1... 118 5e-27
TC220307 similar to GB|AAM65362.1|21655301|AY103310 At2g37550/F1... 114 6e-26
BE608407 similar to GP|4519792|dbj Asp1 {Arabidopsis thaliana}, ... 110 9e-25
TC225079 similar to UP|Q8L8M0 (Q8L8M0) ARF GAP-like zinc finger-... 94 1e-19
TC229990 similar to GB|AAP68261.1|31711810|BT008822 At4g21160 {A... 81 7e-16
BQ253424 81 1e-15
TC207362 similar to UP|Q9FIT8 (Q9FIT8) GCN4-complementing protei... 71 1e-12
TC227348 similar to UP|Q9SMX5 (Q9SMX5) GCN4-complementing protei... 69 4e-12
BF068241 58 9e-09
BI315851 51 8e-07
TC228148 similar to UP|Q7YXC8 (Q7YXC8) C. elegans GRD-1 protein ... 39 0.006
BG405593 homologue to GP|10441354|gb| ARF GAP-like zinc finger-c... 37 0.016
TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {A... 37 0.016
BE556628 36 0.027
>TC225892 similar to UP|ARG3_HUMAN (Q9NP61) ADP-ribosylation factor
GTPase-activating protein 3 (ARF GAP 3), partial (12%)
Length = 1727
Score = 627 bits (1617), Expect = e-180
Identities = 328/412 (79%), Positives = 362/412 (87%), Gaps = 3/412 (0%)
Frame = +2
Query: 1 MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
MASD FTDKN VFRKLK KSENK CFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS
Sbjct: 209 MASDGFTDKNTVFRKLKAKSENKMCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 388
Query: 61 FVRSTNLDSWTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSKE 120
FVRSTNLDSW+PEQLK MSFGGN+RA FF+QHGW GK+EAKYTSRAA+LY+Q+LSKE
Sbjct: 389 FVRSTNLDSWSPEQLKTMSFGGNNRAHGFFKQHGWTDGGKIEAKYTSRAADLYRQILSKE 568
Query: 121 VAKSMSEEAALSAPPAASSQSAQGTNGLPDVKTNEVPIEKTVEKTVEKPEKTESSSSPRA 180
VAKSM+E+ L + P A SQSAQG NGLP+VKTNEVP E T+EKPEK ES+SSPRA
Sbjct: 569 VAKSMAEDGGLPSSPVA-SQSAQGVNGLPEVKTNEVP----KENTLEKPEKPESTSSPRA 733
Query: 181 -YTAVSNNLKKPIGAKKTGKSGGLGARKLTRKPSESLYEQKPEELPAPVSSSTITKNNLP 239
++ +S +KKPIGAKK KSGGLGARKLT+KPSESLYEQKPEE PAPV SST NN+P
Sbjct: 734 SHSVISGTVKKPIGAKKAVKSGGLGARKLTKKPSESLYEQKPEEPPAPVPSST---NNMP 904
Query: 240 SGPPLTSRFEYTEDVQSSELNSGGSNVTGHVSVPKSSSSFFSDFGMDSGFQKKSGPSSSK 299
+GP TSRFEY E+VQSS+LN+GGS+V HVS PK SSSFF+DFGMD GF KKSGPSSSK
Sbjct: 905 AGPSPTSRFEYVENVQSSDLNTGGSHVLSHVSPPK-SSSFFADFGMDGGFPKKSGPSSSK 1081
Query: 300 VQIQESDEARKKFSNAKSISSSQFFGDQNK-ANADAQATLSKFSGSSAISSADLFGDSSD 358
VQIQE+DEAR+KFSNAKSISSSQFFGDQNK A+ D+QATLSKFSGSSAISSADLFGDS D
Sbjct: 1082VQIQETDEARRKFSNAKSISSSQFFGDQNKAADVDSQATLSKFSGSSAISSADLFGDSRD 1261
Query: 359 -NVDLAASDLINRISFQAQQDISSLKNIAGETGKKLTSLASSLMTDLQDRIL 409
N+DL A DLINR+SFQAQQD+SSLKNIAGETGKKL+SLAS+LMTDLQDRIL
Sbjct: 1262NNIDLTAGDLINRLSFQAQQDLSSLKNIAGETGKKLSSLASTLMTDLQDRIL 1417
>TC225894 similar to UP|Q9FIQ0 (Q9FIQ0) Zinc finger protein Glo3-like,
partial (32%)
Length = 612
Score = 244 bits (623), Expect = 5e-65
Identities = 114/135 (84%), Positives = 124/135 (91%)
Frame = +1
Query: 1 MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
MASD FTDKN VFRKLK KSENK CFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS
Sbjct: 208 MASDGFTDKNTVFRKLKAKSENKMCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 387
Query: 61 FVRSTNLDSWTPEQLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSKE 120
FVRSTNLDSW+PEQLK MSFGGN+RAQVFF+QHGWN GK+EAKYTSRAA+LY+Q+LSKE
Sbjct: 388 FVRSTNLDSWSPEQLKTMSFGGNNRAQVFFKQHGWNDGGKIEAKYTSRAADLYRQILSKE 567
Query: 121 VAKSMSEEAALSAPP 135
VAKSM+E+ L + P
Sbjct: 568 VAKSMAEDGGLPSSP 612
>TC225893 similar to UP|Q9FIQ0 (Q9FIQ0) Zinc finger protein Glo3-like,
partial (28%)
Length = 674
Score = 191 bits (484), Expect = 7e-49
Identities = 102/118 (86%), Positives = 112/118 (94%), Gaps = 2/118 (1%)
Frame = +1
Query: 294 GPSSSKVQIQESDEARKKFSNAKSISSSQFFGDQNKA-NADAQATLSKFSGSSAISSADL 352
GPSSSKVQIQE+DEAR+KFSNAKSISSSQ+FGDQNKA + D+QATLSKFSGSSAISSADL
Sbjct: 1 GPSSSKVQIQETDEARRKFSNAKSISSSQYFGDQNKAADVDSQATLSKFSGSSAISSADL 180
Query: 353 FGDSSDN-VDLAASDLINRISFQAQQDISSLKNIAGETGKKLTSLASSLMTDLQDRIL 409
FGDS DN +DL A DLINR+SFQAQQD+SSLKNIAGETGKKL+SLAS+LMTDLQDRIL
Sbjct: 181 FGDSRDNNIDLTAGDLINRLSFQAQQDLSSLKNIAGETGKKLSSLASTLMTDLQDRIL 354
>AW186372
Length = 418
Score = 180 bits (457), Expect = 9e-46
Identities = 97/141 (68%), Positives = 111/141 (77%)
Frame = +2
Query: 182 TAVSNNLKKPIGAKKTGKSGGLGARKLTRKPSESLYEQKPEELPAPVSSSTITKNNLPSG 241
+ +S KPIGAKK KSGGLGARKLT+KPSESLYEQKPEE PAPV SST N++P+G
Sbjct: 8 SVISGTGNKPIGAKKAVKSGGLGARKLTKKPSESLYEQKPEEPPAPVPSST---NSMPAG 178
Query: 242 PPLTSRFEYTEDVQSSELNSGGSNVTGHVSVPKSSSSFFSDFGMDSGFQKKSGPSSSKVQ 301
P TSRFEY E+VQS +LN+GGS+V HV PK SSSFF+DFGMD G KKSGPSS KVQ
Sbjct: 179 PSPTSRFEYVENVQSCDLNTGGSHVLSHVYPPK-SSSFFADFGMDGGCPKKSGPSSCKVQ 355
Query: 302 IQESDEARKKFSNAKSISSSQ 322
I E+DEAR K NAK+ SSS+
Sbjct: 356 IHETDEARMKCVNAKADSSSE 418
>BF423598 weakly similar to GP|9758511|dbj| zinc finger protein Glo3-like
{Arabidopsis thaliana}, partial (20%)
Length = 364
Score = 138 bits (348), Expect = 4e-33
Identities = 78/119 (65%), Positives = 92/119 (76%), Gaps = 2/119 (1%)
Frame = +2
Query: 277 SSFFSDFGMDSGFQKKSGPSSSKVQIQESDEARKKFSNAKSISSSQFFGDQNK-ANADAQ 335
SSFF+D GMD GF KKSGPSSSKVQIQE+DE+R+KFS+AKSISSSQ GD N+ A+ D +
Sbjct: 8 SSFFADLGMDGGFAKKSGPSSSKVQIQETDESRRKFSHAKSISSSQILGDHNRAADVDDE 187
Query: 336 ATLSKFSGSSAISSADLFGDSSDN-VDLAASDLINRISFQAQQDISSLKNIAGETGKKL 393
ATL+K++ SAISSADL G DN VDL A DL +R S QA QD+S LKNI+ ET L
Sbjct: 188 ATLTKWTAPSAISSADLVGC*RDNTVDLTAGDLSHRSSMQAPQDLSVLKNISRETRSTL 364
>BF424278
Length = 417
Score = 138 bits (347), Expect = 5e-33
Identities = 74/89 (83%), Positives = 83/89 (93%), Gaps = 2/89 (2%)
Frame = +3
Query: 317 SISSSQFFGDQNKA-NADAQATLSKFSGSSAISSADLFGDSSDN-VDLAASDLINRISFQ 374
+ISSSQ+FGDQNKA + D+QATLSKFSGSSAISSADLFGDS DN +DL A DLINR+SFQ
Sbjct: 150 AISSSQYFGDQNKAADVDSQATLSKFSGSSAISSADLFGDSRDNNIDLTAGDLINRLSFQ 329
Query: 375 AQQDISSLKNIAGETGKKLTSLASSLMTD 403
AQQD+SSLKNIAGETGKKL+SLAS+LMTD
Sbjct: 330 AQQDLSSLKNIAGETGKKLSSLASTLMTD 416
>TC217447 similar to GB|AAM65362.1|21655301|AY103310 At2g37550/F13M22.5
{Arabidopsis thaliana;} , partial (35%)
Length = 563
Score = 118 bits (295), Expect = 5e-27
Identities = 56/142 (39%), Positives = 82/142 (57%), Gaps = 11/142 (7%)
Frame = +3
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWTPE 73
R L+++ NK C DC+ KNP WASV+YG+F+C++CS HR LGVHISFVRS +DSW+
Sbjct: 60 RDLQSQPANKICVDCSQKNPQWASVSYGVFMCLECSGKHRGLGVHISFVRSVTMDSWSEI 239
Query: 74 QLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLS-----------KEVA 122
Q+K M GGN + F Q+G + + AKY S AA +Y+ + V
Sbjct: 240 QIKKMEAGGNDKLNAFLTQYGIPKETDIVAKYNSNAAAVYRDRIQALADGRPWRDPPVVK 419
Query: 123 KSMSEEAALSAPPAASSQSAQG 144
+++ A+ PP ++ + G
Sbjct: 420 EAVRSSASKGKPPLSAGNNNNG 485
>TC220307 similar to GB|AAM65362.1|21655301|AY103310 At2g37550/F13M22.5
{Arabidopsis thaliana;} , partial (28%)
Length = 602
Score = 114 bits (286), Expect = 6e-26
Identities = 53/136 (38%), Positives = 79/136 (57%)
Frame = +2
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWTPE 73
R L++++ NK C DC+ KNP WASV+YG+F+C++CS HR LGVHISFVRS +DSW+
Sbjct: 170 RDLQSEAGNKICVDCSQKNPQWASVSYGVFMCLECSGKHRGLGVHISFVRSVTMDSWSDI 349
Query: 74 QLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYKQLLSKEVAKSMSEEAALSA 133
Q+K M GGN + F Q+ + + KY + AA +Y+ + ++++E
Sbjct: 350 QIKKMEAGGNDKLNAFLAQYSIPKETDIVTKYNTNAASVYRNRI-----QAIAEGRPWRD 514
Query: 134 PPAASSQSAQGTNGLP 149
PP + G P
Sbjct: 515 PPVLKENLSAAGKGKP 562
>BE608407 similar to GP|4519792|dbj Asp1 {Arabidopsis thaliana}, partial
(26%)
Length = 421
Score = 110 bits (276), Expect = 9e-25
Identities = 48/101 (47%), Positives = 67/101 (65%)
Frame = +2
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWTPE 73
R L+++ NK C DC+ KNP WASV+YG+F+C++CS HR L VHI FVRS +DSW+
Sbjct: 20 RDLQSQPANKICVDCSQKNPQWASVSYGVFMCLECSGKHRGLCVHICFVRSVTMDSWSEI 199
Query: 74 QLKMMSFGGNSRAQVFFRQHGWNGDGKVEAKYTSRAAELYK 114
Q+K M GGN + F Q+G + + KY+S AA +Y+
Sbjct: 200 QIKKMEAGGNDKLNAFLLQYGIPKETDIVVKYSSNAASVYR 322
>TC225079 similar to UP|Q8L8M0 (Q8L8M0) ARF GAP-like zinc finger-containing
protein ZIGA3, partial (48%)
Length = 1845
Score = 94.0 bits (232), Expect = 1e-19
Identities = 46/100 (46%), Positives = 58/100 (58%), Gaps = 11/100 (11%)
Frame = +3
Query: 2 ASDSFTDKNAVFRKLKTKS-----------ENKSCFDCNAKNPTWASVTYGIFLCIDCSA 50
AS + K V ++L K EN+ C DC AK P WASV GIF+C+ CS
Sbjct: 123 ASSTMNSKANVSKELNAKHKKILEGLLKLPENRECADCKAKGPRWASVNLGIFICMQCSG 302
Query: 51 VHRSLGVHISFVRSTNLDSWTPEQLKMMSFGGNSRAQVFF 90
+HRSLGVHIS VRS LD+W PEQ+ + GN +A F+
Sbjct: 303 IHRSLGVHISKVRSATLDTWLPEQVAFIQSMGNEKANCFW 422
>TC229990 similar to GB|AAP68261.1|31711810|BT008822 At4g21160 {Arabidopsis
thaliana;} , partial (22%)
Length = 444
Score = 81.3 bits (199), Expect = 7e-16
Identities = 38/77 (49%), Positives = 51/77 (65%), Gaps = 4/77 (5%)
Frame = +3
Query: 14 RKLKT---KSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSW 70
RKLK +S+N+ C DCNA +P WAS G+F+C+ C VHRSLG HIS V S LD W
Sbjct: 207 RKLKDLLLQSDNRLCADCNAPDPKWASANIGVFICLKCCGVHRSLGTHISKVLSVTLDDW 386
Query: 71 TPEQL-KMMSFGGNSRA 86
+ +++ M+ GGN+ A
Sbjct: 387 SEDEIDAMIEVGGNASA 437
>BQ253424
Length = 277
Score = 80.9 bits (198), Expect = 1e-15
Identities = 34/50 (68%), Positives = 38/50 (76%)
Frame = +2
Query: 21 ENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSW 70
ENK C DC AK P WASV GIF+C+ CS +HRSLGVHIS VRS LD+W
Sbjct: 128 ENKECADCKAKGPRWASVNLGIFICMQCSGIHRSLGVHISKVRSATLDTW 277
>TC207362 similar to UP|Q9FIT8 (Q9FIT8) GCN4-complementing protein homolog,
partial (51%)
Length = 1959
Score = 70.9 bits (172), Expect = 1e-12
Identities = 33/67 (49%), Positives = 41/67 (60%), Gaps = 2/67 (2%)
Frame = +1
Query: 22 NKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLD--SWTPEQLKMMS 79
N C DC A P WAS+ G+ +CI+CS VHR+LGVHIS VRS LD W P + +
Sbjct: 793 NDKCADCGAPEPDWASLNLGVLVCIECSGVHRNLGVHISKVRSLTLDVKVWEPSVISLFQ 972
Query: 80 FGGNSRA 86
GN+ A
Sbjct: 973 SLGNTFA 993
>TC227348 similar to UP|Q9SMX5 (Q9SMX5) GCN4-complementing protein (GCP1),
partial (29%)
Length = 1197
Score = 68.9 bits (167), Expect = 4e-12
Identities = 43/113 (38%), Positives = 59/113 (52%), Gaps = 9/113 (7%)
Frame = +3
Query: 22 NKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLD--SWTPEQLKMMS 79
N C +C+A P WAS+ GI LCI+CS VHR+LGVH+S VRS LD W L++
Sbjct: 594 NDKCAECSAPEPDWASLNLGILLCIECSGVHRNLGVHVSKVRSITLDVRVWENTVLELFD 773
Query: 80 FGGNSRAQ-----VFFRQHGWNGDGKVEAKYTSRAAELYKQ--LLSKEVAKSM 125
GN+ + H G+ V K S A +K+ + +K V KS+
Sbjct: 774 NLGNAYCNSIWEGLLLLDHERVGEPNVPMKPCSADAFQHKEKYIQAKYVEKSL 932
>BF068241
Length = 387
Score = 57.8 bits (138), Expect = 9e-09
Identities = 30/82 (36%), Positives = 43/82 (51%), Gaps = 1/82 (1%)
Frame = +2
Query: 13 FRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVRSTNLDSWTP 72
F+ +K N +C C S G+F+C+ C VHRSLG HIS V S LD W+
Sbjct: 134 FKLVKE*LPNGTCKTC------LKSANIGVFICLKCCGVHRSLGTHISKVLSVTLDDWSE 295
Query: 73 EQL-KMMSFGGNSRAQVFFRQH 93
+++ MM GGN+ A + +
Sbjct: 296 DEIDAMMEVGGNASANSIYEAY 361
>BI315851
Length = 385
Score = 51.2 bits (121), Expect = 8e-07
Identities = 19/33 (57%), Positives = 23/33 (69%)
Frame = +1
Query: 21 ENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHR 53
EN+ C DC K P WASV GIF+C+ CS +HR
Sbjct: 286 ENRECADCRNKAPRWASVNLGIFICMQCSGIHR 384
>TC228148 similar to UP|Q7YXC8 (Q7YXC8) C. elegans GRD-1 protein
(Corresponding sequence R08B4.1b), partial (3%)
Length = 1180
Score = 38.5 bits (88), Expect = 0.006
Identities = 47/184 (25%), Positives = 77/184 (41%)
Frame = +3
Query: 159 EKTVEKTVEKPEKTESSSSPRAYTAVSNNLKKPIGAKKTGKSGGLGARKLTRKPSESLYE 218
E+ + K EKP T + S R+Y ++NN K + T K+G G KPSE++
Sbjct: 228 EEELPKAKEKPV-TGARSLRRSYKNLNNNNNKKPSSNSTPKTGSSG-----NKPSETV-- 383
Query: 219 QKPEELPAPVSSSTITKNNLPSGPPLTSRFEYTEDVQSSELNSGGSNVTGHVSVPKSSSS 278
KPE+ A K N + R + ++S+ SGGS G SSSS
Sbjct: 384 -KPEKSKA--EGGPDKKRN--AAADFLKRIKRNTSAEASKGGSGGSGGGGSGGGGGSSSS 548
Query: 279 FFSDFGMDSGFQKKSGPSSSKVQIQESDEARKKFSNAKSISSSQFFGDQNKANADAQATL 338
G + ++K ++ K D+ +++ S ++ GD+ + +
Sbjct: 549 SKGGGGGNGVKEQKKMVNNGK-----GDKGKERASRHNNVGGGSGSGDKRNSKNVENNSQ 713
Query: 339 SKFS 342
SK S
Sbjct: 714 SKRS 725
>BG405593 homologue to GP|10441354|gb| ARF GAP-like zinc finger-containing
protein ZiGA4 {Arabidopsis thaliana}, partial (8%)
Length = 385
Score = 37.0 bits (84), Expect = 0.016
Identities = 12/41 (29%), Positives = 22/41 (53%)
Frame = +1
Query: 12 VFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVH 52
+ R L N+ C +CN+ P + ++ F+C+ CS +H
Sbjct: 262 IIRGLMKLPPNRRCINCNSLGPQYVCTSFWTFICMTCSGIH 384
>TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;} , partial (46%)
Length = 902
Score = 37.0 bits (84), Expect = 0.016
Identities = 39/160 (24%), Positives = 64/160 (39%), Gaps = 4/160 (2%)
Frame = -3
Query: 226 APVSSSTITKNNLPSGPPLTSRFEYTEDVQSSELNSGGSNVTGHVSVPKSSSSFFSDFGM 285
A S + + + S P +S + + SS ++ G SVPKSSSS S
Sbjct: 702 ASAFSQSSSSTSSSSSSPFSSGSKSLDASSSSAVSKGRIRY----SVPKSSSSPCSSSSS 535
Query: 286 DSGFQKKSGPSSSKVQIQESDEAR----KKFSNAKSISSSQFFGDQNKANADAQATLSKF 341
S S SS + R S++ S SS + ++ + ++ S
Sbjct: 534 SSSSSSSSESSSESLSSSSGSPPRWTSPSSSSSSSSSSSPPSGASSSSSSLSSPSSSSSS 355
Query: 342 SGSSAISSADLFGDSSDNVDLAASDLINRISFQAQQDISS 381
S SS+ SSA S+D+ ++S SF + ++S
Sbjct: 354 SSSSSSSSASSTTSSTDSYSSSSSSTKTLTSFSSTCSLNS 235
Score = 37.0 bits (84), Expect = 0.016
Identities = 40/161 (24%), Positives = 66/161 (40%), Gaps = 10/161 (6%)
Frame = -3
Query: 209 TRKPSESLYEQKPEELPAPVSSSTITKNNLPSGPPLTSRFEYTEDVQSSELNSGGSNVTG 268
T S S + + L A SSS ++K + P +S + SS +S + +
Sbjct: 672 TSSSSSSPFSSGSKSLDAS-SSSAVSKGRIRYSVPKSSSSPCSSSSSSSSSSSSSESSSE 496
Query: 269 HVSV----------PKSSSSFFSDFGMDSGFQKKSGPSSSKVQIQESDEARKKFSNAKSI 318
+S P SSSS S SG + SSS + S + S++ S
Sbjct: 495 SLSSSSGSPPRWTSPSSSSSSSSSSSPPSG----ASSSSSSLSSPSSSSSSSSSSSSSSA 328
Query: 319 SSSQFFGDQNKANADAQATLSKFSGSSAISSADLFGDSSDN 359
SS+ D +++ + TL+ FS + +++S F S N
Sbjct: 327 SSTTSSTDSYSSSSSSTKTLTSFSSTCSLNSFVAFIFESRN 205
>BE556628
Length = 112
Score = 36.2 bits (82), Expect = 0.027
Identities = 13/26 (50%), Positives = 18/26 (69%)
Frame = +2
Query: 22 NKSCFDCNAKNPTWASVTYGIFLCID 47
N C +C+A +P W+S+ GI LCID
Sbjct: 35 NDKCAECSAPDPYWSSLNLGILLCID 112
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.307 0.123 0.332
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,879,072
Number of Sequences: 63676
Number of extensions: 195624
Number of successful extensions: 1133
Number of sequences better than 10.0: 88
Number of HSP's better than 10.0 without gapping: 1081
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1102
length of query: 409
length of database: 12,639,632
effective HSP length: 100
effective length of query: 309
effective length of database: 6,272,032
effective search space: 1938057888
effective search space used: 1938057888
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC137670.5


