

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137669.5 + phase: 0
(58 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
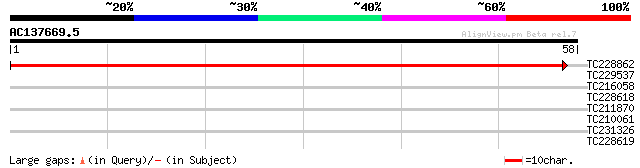
TC228862 weakly similar to UP|Q94BX2 (Q94BX2) AT3g50910/F18B3_19... 86 4e-18
TC229537 homologue to UP|Q8LD11 (Q8LD11) 26S proteasome non-ATPa... 30 0.33
TC216058 similar to UP|MCAT_ARATH (Q93XM7) Mitochondrial carniti... 26 3.7
TC228618 similar to UP|Q9SKW6 (Q9SKW6) F5J5.4, partial (29%) 25 6.3
TC211870 25 6.3
TC210061 similar to UP|Q6NQN0 (Q6NQN0) At1g53760, partial (53%) 25 8.2
TC231326 25 8.2
TC228619 similar to GB|AAO24579.1|27808598|BT003147 At1g36050 {A... 25 8.2
>TC228862 weakly similar to UP|Q94BX2 (Q94BX2) AT3g50910/F18B3_190, partial
(35%)
Length = 1187
Score = 85.9 bits (211), Expect = 4e-18
Identities = 39/57 (68%), Positives = 47/57 (82%)
Frame = +2
Query: 1 MARRERQRKSKKQKWIWGSLTTVIALGTAAVAWSYLPAGGESYSAEDHPVPKHDDAA 57
MARRERQR+S++Q+WIWGS+TT IA+GTAA+AWSYLP G S SA V +HDDAA
Sbjct: 530 MARRERQRRSRRQRWIWGSITTAIAVGTAAIAWSYLPVGRGSTSAVHDQVSEHDDAA 700
>TC229537 homologue to UP|Q8LD11 (Q8LD11) 26S proteasome non-ATPase
regulatory subunit, partial (45%)
Length = 515
Score = 29.6 bits (65), Expect = 0.33
Identities = 8/28 (28%), Positives = 18/28 (63%)
Frame = +1
Query: 9 KSKKQKWIWGSLTTVIALGTAAVAWSYL 36
K + ++W+WG +T ++ LG + W+ +
Sbjct: 376 KLEGRRWLWGGITRILDLGVGFLEWTLI 459
>TC216058 similar to UP|MCAT_ARATH (Q93XM7) Mitochondrial
carnitine/acylcarnitine carrier-like protein (A BOUT DE
SOUFFLE) (Carnitine/acylcarnitine translocase-like
protein) (CAC-like protein), partial (97%)
Length = 1441
Score = 26.2 bits (56), Expect = 3.7
Identities = 10/33 (30%), Positives = 19/33 (57%), Gaps = 1/33 (3%)
Frame = -1
Query: 4 RERQRKSKKQKWIW-GSLTTVIALGTAAVAWSY 35
RER+R+S K+ W W G+ + + + W++
Sbjct: 112 RERERESHKKVWDWNGTKRNKVCVAQTWICWNH 14
>TC228618 similar to UP|Q9SKW6 (Q9SKW6) F5J5.4, partial (29%)
Length = 730
Score = 25.4 bits (54), Expect = 6.3
Identities = 10/25 (40%), Positives = 15/25 (60%)
Frame = -1
Query: 15 WIWGSLTTVIALGTAAVAWSYLPAG 39
WI LT+V +GT + + Y+P G
Sbjct: 133 WIVCPLTSVYTVGTTLIKYWYIPLG 59
>TC211870
Length = 717
Score = 25.4 bits (54), Expect = 6.3
Identities = 9/21 (42%), Positives = 15/21 (70%)
Frame = -1
Query: 2 ARRERQRKSKKQKWIWGSLTT 22
A+RER+++ K +WI + TT
Sbjct: 171 AKREREKRKKNVEWIGAAATT 109
>TC210061 similar to UP|Q6NQN0 (Q6NQN0) At1g53760, partial (53%)
Length = 827
Score = 25.0 bits (53), Expect = 8.2
Identities = 7/28 (25%), Positives = 16/28 (57%)
Frame = -3
Query: 4 RERQRKSKKQKWIWGSLTTVIALGTAAV 31
R+ ++ + +W W S ++ + LGT +
Sbjct: 93 RKHGKRRESDRWFWRSTSSCLVLGTQTI 10
>TC231326
Length = 510
Score = 25.0 bits (53), Expect = 8.2
Identities = 10/30 (33%), Positives = 17/30 (56%)
Frame = +2
Query: 4 RERQRKSKKQKWIWGSLTTVIALGTAAVAW 33
RER ++S + K +W S+ + +A AW
Sbjct: 47 RERGKESIEVKMVWNSMVVLRVARASAEAW 136
>TC228619 similar to GB|AAO24579.1|27808598|BT003147 At1g36050 {Arabidopsis
thaliana;} , partial (70%)
Length = 1191
Score = 25.0 bits (53), Expect = 8.2
Identities = 10/25 (40%), Positives = 15/25 (60%)
Frame = -3
Query: 15 WIWGSLTTVIALGTAAVAWSYLPAG 39
WI LT+V +GT + + Y+P G
Sbjct: 559 WIVCPLTSVYTVGTTLIKY*YIPLG 485
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.313 0.126 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,775,026
Number of Sequences: 63676
Number of extensions: 26603
Number of successful extensions: 182
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 181
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 182
length of query: 58
length of database: 12,639,632
effective HSP length: 34
effective length of query: 24
effective length of database: 10,474,648
effective search space: 251391552
effective search space used: 251391552
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 52 (24.6 bits)
Medicago: description of AC137669.5


