

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137553.4 - phase: 0 /pseudo
(921 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
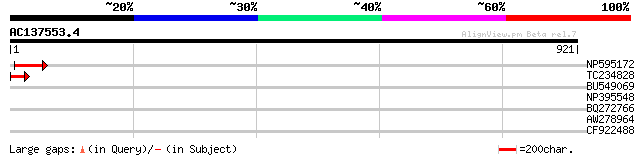
NP595172 polyprotein [Glycine max] 56 8e-08
TC234828 46 9e-05
BU549069 41 0.003
NP395548 reverse transcriptase [Glycine max] 37 0.052
BQ272766 weakly similar to GP|28558781|gb| pol protein {Cucumis ... 31 2.2
AW278964 weakly similar to SP|O82702|VAG1 Vacuolar ATP synthase ... 30 6.3
CF922488 25 9.1
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 55.8 bits (133), Expect = 8e-08
Identities = 23/54 (42%), Positives = 38/54 (69%)
Frame = +1
Query: 8 ELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNRY 61
+++K +++++ +PS SP+ P+LLVKKKDGS R DYR LN +T+K+ +
Sbjct: 1834 QIEKMIQEMLVQGIIQPSNSPFSLPILLVKKKDGSWRFCTDYRALNAITVKDSF 1995
>TC234828
Length = 857
Score = 45.8 bits (107), Expect = 9e-05
Identities = 20/32 (62%), Positives = 25/32 (77%)
Frame = +3
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAP 32
MS ELAE+K Q++DL+ +F RPS SPWGAP
Sbjct: 759 MSPVELAEVKAQVQDLLSKQFVRPSASPWGAP 854
>BU549069
Length = 615
Score = 40.8 bits (94), Expect = 0.003
Identities = 23/66 (34%), Positives = 38/66 (56%)
Frame = -3
Query: 790 ESHSCD*CWTCFEVKEVDSEIYWSVSDIGKSWNGGVSSWITTASFEFARYITCVATSKVC 849
ESHS D W+ E+ + + +Y S + KS + G+ + IT +F ++ ++CV+T V
Sbjct: 613 ESHSKDWGWSSTEIPKTHTSLYRSFPNS*KSXSCGIPNCITPITF*SSQCLSCVSTPYVY 434
Query: 850 TGSISC 855
SISC
Sbjct: 433 P*SISC 416
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 36.6 bits (83), Expect = 0.052
Identities = 25/74 (33%), Positives = 39/74 (51%), Gaps = 19/74 (25%)
Frame = +1
Query: 9 LKKQLEDLVEMKFFRP-SVSPWGAPVLLVKKKDG------------------SMRLRIDY 49
++K++ L+E+ P S S W +PVL+V KK+G S +L IDY
Sbjct: 1 VRKEVLKLLEVGLIYPISDSAWVSPVLVVSKKEGMTVIRNEKNDLIPTRTVTSWKLCIDY 180
Query: 50 RQLNKVTIKNRYLL 63
R+LN+ T K+ + L
Sbjct: 181RKLNEATRKDHFPL 222
>BQ272766 weakly similar to GP|28558781|gb| pol protein {Cucumis melo},
partial (9%)
Length = 410
Score = 31.2 bits (69), Expect = 2.2
Identities = 16/55 (29%), Positives = 30/55 (54%)
Frame = -2
Query: 788 VFESHSCD*CWTCFEVKEVDSEIYWSVSDIGKSWNGGVSSWITTASFEFARYITC 842
+ +SH D W+ FE+ + + Y S + KS G+S+ IT S+ ++ ++C
Sbjct: 355 ILKSHPIDWGWSSFEILKTHTSFYRSFPNSQKS*FCGISNCITPISY*PSQCLSC 191
>AW278964 weakly similar to SP|O82702|VAG1 Vacuolar ATP synthase subunit G 1
(EC 3.6.3.14) (V-ATPase G subunit 1), complete
Length = 328
Score = 29.6 bits (65), Expect = 6.3
Identities = 21/74 (28%), Positives = 31/74 (41%), Gaps = 6/74 (8%)
Frame = -2
Query: 788 VFESHSCD*CWTCFEVKEVDSEIYWSVSDIGKSWNGGVSS------WITTASFEFARYIT 841
+F S SC C+T V + ++ +S +W +SS W A F F T
Sbjct: 246 IFSSVSCSSCFT*APESPVMCDPFF*ISASSSAWY*AISSSACSWAWFNIAYFSFFAAFT 67
Query: 842 CVATSKVCTGSISC 855
S C+ +ISC
Sbjct: 66 IRCAS--CSAAISC 31
>CF922488
Length = 741
Score = 25.4 bits (54), Expect(2) = 9.1
Identities = 11/33 (33%), Positives = 21/33 (63%)
Frame = +3
Query: 36 VKKKDGSMRLRIDYRQLNKVTIKNRYLLRELMI 68
V K+DG + + +DYR LN + K+++ L + +
Sbjct: 3 VLKEDGKV*MCVDYRDLN*ASPKDKFPLPHINV 101
Score = 21.9 bits (45), Expect(2) = 9.1
Identities = 14/31 (45%), Positives = 18/31 (57%)
Frame = +1
Query: 86 KVITRLK*KMKICRRQLSERGMVTMNIMLCR 116
+VITR +* +I +RQLS I LCR
Sbjct: 154 QVITR*R*HQRIWKRQLSLLYGEPSAIRLCR 246
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.366 0.163 0.618
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,294,269
Number of Sequences: 63676
Number of extensions: 544499
Number of successful extensions: 7223
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 2776
Number of HSP's successfully gapped in prelim test: 214
Number of HSP's that attempted gapping in prelim test: 4355
Number of HSP's gapped (non-prelim): 3192
length of query: 921
length of database: 12,639,632
effective HSP length: 106
effective length of query: 815
effective length of database: 5,889,976
effective search space: 4800330440
effective search space used: 4800330440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC137553.4


