

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
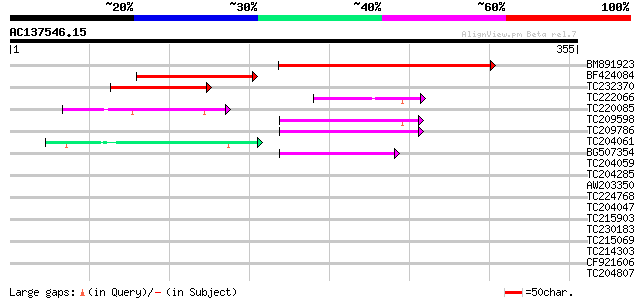
Query= AC137546.15 + phase: 0 /pseudo
(355 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BM891923 similar to GP|8778588|gb| F28C11.4 {Arabidopsis thalian... 205 3e-53
BF424084 similar to GP|8778588|gb|A F28C11.4 {Arabidopsis thalia... 99 2e-21
TC232370 weakly similar to UP|Q9LS91 (Q9LS91) Elongation factor ... 63 2e-10
TC222066 similar to GB|AAQ89616.1|37202002|BT010594 At3g01370 {A... 50 2e-06
TC220085 similar to UP|Q9FFU1 (Q9FFU1) Emb|CAA18192.1, partial (... 48 8e-06
TC209598 weakly similar to UP|Q9LS85 (Q9LS85) Emb|CAB10230.1, pa... 44 1e-04
TC209786 similar to GB|AAS49116.1|44917583|BT011753 At2g28480 {A... 43 2e-04
TC204061 homologue to UP|Q09083 (Q09083) Hydroxyproline-rich gly... 42 6e-04
BG507354 41 7e-04
TC204059 similar to UP|Q94ES3 (Q94ES3) Root nodule extensin (Fra... 40 0.001
TC204285 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 40 0.002
AW203350 39 0.005
TC224768 similar to UP|Q39835 (Q39835) Extensin, partial (33%) 37 0.010
TC204047 hydroxyproline-rich glycoprotein 37 0.014
TC215903 similar to UP|Q9M7N4 (Q9M7N4) MFP1 attachment factor 1,... 37 0.014
TC230183 homologue to UP|GLHR_ANTEL (P35409) Probable glycoprote... 37 0.018
TC215069 similar to UP|O24300 (O24300) PtxA protein precursor, p... 37 0.018
TC214303 similar to UP|Q9T0K5 (Q9T0K5) Extensin-like protein, pa... 36 0.030
CF921606 36 0.030
TC204807 weakly similar to UP|HQGT_RAUSE (Q9AR73) Hydroquinone g... 30 1.3
>BM891923 similar to GP|8778588|gb| F28C11.4 {Arabidopsis thaliana}, partial
(24%)
Length = 421
Score = 205 bits (521), Expect = 3e-53
Identities = 91/136 (66%), Positives = 113/136 (82%)
Frame = +3
Query: 169 GEPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDM 228
GEPL K EI+ LV+ + +RQ+NLGRDG HNML+ IH+HWKRRRVCKI+C+GV TVDM
Sbjct: 12 GEPLKKWEIHMLVKPMMSYNRQVNLGRDGLTHNMLELIHSHWKRRRVCKIRCLGVPTVDM 191
Query: 229 DNVCQQLEEKTGGKVIYRRGGVIYLFRGRNYNHKTRPRFPLMLWKPVPPVYPRLIQQVPE 288
DNVC +EEKTGGK+I+R GGV+YLFRGRNYN+ TRP++P+MLWKP PVYP+LIQ P
Sbjct: 192 DNVCHHIEEKTGGKIIHRVGGVVYLFRGRNYNYSTRPQYPVMLWKPAAPVYPKLIQDAPG 371
Query: 289 GLTLEEATEMRQKGRT 304
GLT +EA E+R KG++
Sbjct: 372 GLTKDEADELRMKGKS 419
>BF424084 similar to GP|8778588|gb|A F28C11.4 {Arabidopsis thaliana}, partial
(12%)
Length = 236
Score = 99.4 bits (246), Expect = 2e-21
Identities = 47/77 (61%), Positives = 56/77 (72%), Gaps = 1/77 (1%)
Frame = +3
Query: 80 FEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSK-KKLKEFDSFVLPPP 138
FEF++SY+ETPK+KP+ +REP F+PF P TMPRPWTG+ PL K KK+ FDSF PP
Sbjct: 3 FEFQFSYSETPKAKPIAIREPAFLPFAPPTMPRPWTGKAPLKGKKSKKVPLFDSFNPPPA 182
Query: 139 HKKGVKPVQSPGPFLPG 155
KGVK V PGPF G
Sbjct: 183 GTKGVKLVDMPGPFSMG 233
>TC232370 weakly similar to UP|Q9LS91 (Q9LS91) Elongation factor EF-2,
partial (3%)
Length = 455
Score = 62.8 bits (151), Expect = 2e-10
Identities = 28/63 (44%), Positives = 39/63 (61%)
Frame = -3
Query: 64 VKISEDGVSYVIEGAPFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPS 123
+ I + GV Y++ APFEF++ Y++T K KP+ + E F PF P TM R WT + PL
Sbjct: 360 IVIGDSGVLYLLPTAPFEFQFDYSKTSKVKPIAIHELSFPPFVPPTMSRSWTEKAPLKGK 181
Query: 124 KKK 126
K K
Sbjct: 180 KSK 172
>TC222066 similar to GB|AAQ89616.1|37202002|BT010594 At3g01370 {Arabidopsis
thaliana;} , partial (14%)
Length = 459
Score = 50.1 bits (118), Expect = 2e-06
Identities = 27/74 (36%), Positives = 42/74 (56%), Gaps = 4/74 (5%)
Frame = +3
Query: 191 LNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMDNVCQQLEEKTGGKVI----YR 246
L LGR G ++N+H HWK R + KI C G D ++ LE ++GG ++ R
Sbjct: 48 LLLGRRGVFDGTVENMHLHWKYRELVKIICNGSLEED-HHIALTLEAESGGILVAVERVR 224
Query: 247 RGGVIYLFRGRNYN 260
+G I ++RG+NY+
Sbjct: 225 KGFAIIVYRGKNYS 266
Score = 32.0 bits (71), Expect = 0.44
Identities = 15/49 (30%), Positives = 27/49 (54%)
Frame = +3
Query: 298 MRQKGRTLTPICKLGKNGVYYNLVNNVREAFEECELVRVNCQGLNKSDY 346
+R+ G + P LG+ GV+ V N+ ++ ELV++ C G + D+
Sbjct: 15 LRRIGLMMKPFLLLGRRGVFDGTVENMHLHWKYRELVKIICNGSLEEDH 161
>TC220085 similar to UP|Q9FFU1 (Q9FFU1) Emb|CAA18192.1, partial (13%)
Length = 464
Score = 47.8 bits (112), Expect = 8e-06
Identities = 36/112 (32%), Positives = 55/112 (48%), Gaps = 7/112 (6%)
Frame = +2
Query: 34 PKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVI--EGAPFEFKYSYTET-P 90
P P P K +K K N EN +++G + PF+F+YSY+E+ P
Sbjct: 116 PPFSPSPKPYYKSKKNHAEKKKKNNS--ENGDPNKNGAPKLPFKSNLPFDFRYSYSESDP 289
Query: 91 KSKPVQMREPP-FVPFGPVTMPRPWTG-RPPL--PPSKKKLKEFDSFVLPPP 138
P+ RE P F PFGP + R WTG P+ P +++++E + +L P
Sbjct: 290 SVGPISFRESPKFSPFGPGRIDRKWTGVSAPVQGEPDRERVEEERNRILGEP 445
>TC209598 weakly similar to UP|Q9LS85 (Q9LS85) Emb|CAB10230.1, partial (22%)
Length = 1029
Score = 43.5 bits (101), Expect = 1e-04
Identities = 28/95 (29%), Positives = 45/95 (46%), Gaps = 5/95 (5%)
Frame = +3
Query: 170 EPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMD 229
E LT EE + L L LGR ++N+H HWK R + K+ G + +
Sbjct: 477 ETLTNEERFLFRKIGLSMKPYLLLGRRDVYAGTIENMHLHWKYRELVKLIVKGRNSAQVK 656
Query: 230 NVCQQLEEKTGGKVI-----YRRGGVIYLFRGRNY 259
++ LE ++GG ++ R I ++RG+NY
Sbjct: 657 HISISLEAESGGVLVSVDKDTRGHHTIIVYRGKNY 761
>TC209786 similar to GB|AAS49116.1|44917583|BT011753 At2g28480 {Arabidopsis
thaliana;} , partial (64%)
Length = 1160
Score = 43.1 bits (100), Expect = 2e-04
Identities = 27/91 (29%), Positives = 41/91 (44%), Gaps = 1/91 (1%)
Frame = +2
Query: 170 EPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMD 229
+ LT EE L + K S L +GR G ++ N+H HWK+ K+ C +
Sbjct: 473 DELTGEERFYLKKMGQKRSNYLQIGRRGLFGGVVLNMHMHWKKHETVKVFCKPCKPGQVH 652
Query: 230 NVCQQLEEKTGGKVIYRRG-GVIYLFRGRNY 259
Q+L +GG + G I +R +NY
Sbjct: 653 EYAQELARLSGGIPLQIIGDDTIIFYRRKNY 745
>TC204061 homologue to UP|Q09083 (Q09083) Hydroxyproline-rich glycoprotein
precursor, partial (28%)
Length = 848
Score = 41.6 bits (96), Expect = 6e-04
Identities = 42/144 (29%), Positives = 52/144 (35%), Gaps = 8/144 (5%)
Frame = +1
Query: 23 PTGKPNKNPSKP----KVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGA 78
P KP K PS P K P K+ + P KP K P S Y +
Sbjct: 85 PPKKPYKYPSPPPPVYKYKSPPPPVYKYKSPPPPPKKPY-KYP-----SPPPPVYKYKSP 246
Query: 79 PFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWT-GRPPLPPSKKKLKEFDSFVL-- 135
P +KY P KP + PP + + P P+ PP PP K ++
Sbjct: 247 PPPYKYPSPPPPPKKPYKYPSPPPPVYKYKSPPPPYKYSSPPPPPYKYPSPPPPAYYYKS 426
Query: 136 -PPPHKKGVKPVQSPGPFLPGTSP 158
PPP KK K P P SP
Sbjct: 427 PPPPPKKPYKYPSPPPPHYVYASP 498
Score = 33.9 bits (76), Expect = 0.12
Identities = 26/91 (28%), Positives = 36/91 (38%), Gaps = 5/91 (5%)
Frame = +1
Query: 73 YVIEGAPFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRP-WTGRPPLPPSKKKLKEFD 131
Y + P +KY P KP + PP + + P P + + P PP KK K
Sbjct: 34 YYYKSPPPPYKYPSPPPPPKKPYKYPSPPPPVYKYKSPPPPVYKYKSPPPPPKKPYK--- 204
Query: 132 SFVLPPP----HKKGVKPVQSPGPFLPGTSP 158
+ PPP +K P + P P P P
Sbjct: 205 -YPSPPPPVYKYKSPPPPYKYPSPPPPPKKP 294
>BG507354
Length = 438
Score = 41.2 bits (95), Expect = 7e-04
Identities = 21/75 (28%), Positives = 37/75 (49%)
Frame = +1
Query: 170 EPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMD 229
E LT EE ++ LKS + +GR G ++ N+H HWK+ + K+ ++
Sbjct: 214 EILTPEEHFFFLKMGLKSKNYVPVGRRGIYQGVILNMHLHWKKHQTLKVVVKTFSAEEVK 393
Query: 230 NVCQQLEEKTGGKVI 244
+ +L +GG V+
Sbjct: 394 EIAAELARLSGGIVL 438
>TC204059 similar to UP|Q94ES3 (Q94ES3) Root nodule extensin (Fragment),
partial (96%)
Length = 853
Score = 40.4 bits (93), Expect = 0.001
Identities = 42/133 (31%), Positives = 46/133 (34%), Gaps = 4/133 (3%)
Frame = +2
Query: 23 PTGKPNKNPSKP----KVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGA 78
P KP K PS P K P HP P +K P VK
Sbjct: 131 PPKKPYKYPSPPPPVKKPYPHPHPVYHSPPPPPKKPYKYPSPPPPVKKPYPH-------- 286
Query: 79 PFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPP 138
P +S TPK PP V P +P P PP PP KK K + PPP
Sbjct: 287 PHPVYHSPPPTPKKPYKYPSPPPPVHTYPPHVPTPVYHSPPPPPYKKPYK----YKSPPP 454
Query: 139 HKKGVKPVQSPGP 151
PV SP P
Sbjct: 455 ------PVHSPPP 475
Score = 34.7 bits (78), Expect = 0.068
Identities = 41/149 (27%), Positives = 47/149 (31%), Gaps = 11/149 (7%)
Frame = +2
Query: 23 PTGKPNKNPSKP----KVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGA 78
P KP K PS P K P HP P +K P VK V
Sbjct: 38 PPKKPYKYPSPPPPVHKPYPHPHPVYHSPPPPPKKPYKYPSPPPPVKKPYPHPHPVYHSP 217
Query: 79 PFEFKYSYTETPKSKPVQMREP---PFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVL 135
P K Y PV+ P P P T +P+ P PP + V
Sbjct: 218 PPPPKKPYKYPSPPPPVKKPYPHPHPVYHSPPPTPKKPYKYPSPPPPVHTYPPHVPTPVY 397
Query: 136 ----PPPHKKGVKPVQSPGPFLPGTSPRY 160
PPP+KK K P P P Y
Sbjct: 398 HSPPPPPYKKPYKYKSPPPPVHSPPPPHY 484
Score = 32.3 bits (72), Expect = 0.34
Identities = 26/80 (32%), Positives = 29/80 (35%)
Frame = +2
Query: 79 PFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPP 138
P +S P KP + PP P P P PP PP KK K + PPP
Sbjct: 5 PHPVYHSPPPPPPKKPYKYPSPPPPVHKPYPHPHPVYHSPP-PPPKKPYK----YPSPPP 169
Query: 139 HKKGVKPVQSPGPFLPGTSP 158
K KP P P P
Sbjct: 170 PVK--KPYPHPHPVYHSPPP 223
>TC204285 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (32%)
Length = 1539
Score = 39.7 bits (91), Expect = 0.002
Identities = 42/133 (31%), Positives = 46/133 (34%), Gaps = 1/133 (0%)
Frame = +1
Query: 23 PTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFEF 82
P P K PS PKV P HP P KP K P K+
Sbjct: 370 PPHPPKKPPSPPKVKPP-HPPKPPKVEPPHPPKPPKKPPCPPKVKPP------------- 507
Query: 83 KYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPP-PHKK 141
PK V EPP P P P P PPLPP K + LPP P K+
Sbjct: 508 -----HPPKPPKV---EPPHPPKPPKKPPSPPKVVPPLPPKPPK----NPPPLPPKPPKE 651
Query: 142 GVKPVQSPGPFLP 154
P + P P P
Sbjct: 652 PPTPPKEPPPTPP 690
Score = 39.3 bits (90), Expect = 0.003
Identities = 43/140 (30%), Positives = 51/140 (35%), Gaps = 4/140 (2%)
Frame = +1
Query: 23 PTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFEF 82
P P K P PKV P HP P KP K P K V+ P
Sbjct: 457 PPKPPKKPPCPPKVKPP-HPPKPPKVEPPHPPKPPKKPPSPPK--------VVPPLP--- 600
Query: 83 KYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPP-SKKKLKEFDSFVLPPP--- 138
+ PK+ P +PP P P P P + P PP KK L PP
Sbjct: 601 ----PKPPKNPPPLPPKPPKEPPTPPKEPPPTPPKKPCPPIFKKPLPPIPKLPPLPPVPI 768
Query: 139 HKKGVKPVQSPGPFLPGTSP 158
+KK + P+ P P LP P
Sbjct: 769 YKKPLPPL-PPLPKLPPLHP 825
Score = 34.3 bits (77), Expect = 0.088
Identities = 41/147 (27%), Positives = 49/147 (32%), Gaps = 7/147 (4%)
Frame = +1
Query: 23 PTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFEF 82
P P K P P P+ P +P K PV K P + V P
Sbjct: 178 PPPVPEKKPPPPV--PEKPPPCPPKVVPPPKPPPV-KPPPKPPVKPPCPPKVEPPHP--- 339
Query: 83 KYSYTETPKSKPVQMREPPFVPFGPVTM-----PRPWTGRPPLPPSKKKLKEFDSFVLP- 136
+ PK KP PP P P + P+P PP PP K V P
Sbjct: 340 ----PKPPKVKPPHPPHPPKKPPSPPKVKPPHPPKPPKVEPPHPPKPPKKPPCPPKVKPP 507
Query: 137 -PPHKKGVKPVQSPGPFLPGTSPRYVM 162
PP V+P P P SP V+
Sbjct: 508 HPPKPPKVEPPHPPKPPKKPPSPPKVV 588
Score = 32.7 bits (73), Expect = 0.26
Identities = 24/74 (32%), Positives = 30/74 (40%), Gaps = 4/74 (5%)
Frame = +1
Query: 90 PKSKPVQMREPP----FVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKP 145
P P+ +PP P PV P P + P PP +K V+PPP VKP
Sbjct: 103 PPLPPLIFHKPPPPVYTPPPVPVYTPPPVPEKKPPPPVPEKPPPCPPKVVPPPKPPPVKP 282
Query: 146 VQSPGPFLPGTSPR 159
P P P P+
Sbjct: 283 PPKP-PVKPPCPPK 321
>AW203350
Length = 411
Score = 38.5 bits (88), Expect = 0.005
Identities = 23/75 (30%), Positives = 35/75 (46%)
Frame = +1
Query: 10 PISASNADQSSRRPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISED 69
P SA N + +S T NP+ P S + + PK+ + K + +S+D
Sbjct: 58 PFSAPNPNPNSNTAT---TPNPNTPNPSSSSFKSRSATEKPKKLPRTKPKLTPELLLSDD 228
Query: 70 GVSYVIEGAPFEFKY 84
G+ YV+ P EFKY
Sbjct: 229 GLGYVLRYFPREFKY 273
>TC224768 similar to UP|Q39835 (Q39835) Extensin, partial (33%)
Length = 483
Score = 37.4 bits (85), Expect = 0.010
Identities = 42/160 (26%), Positives = 55/160 (34%), Gaps = 2/160 (1%)
Frame = +3
Query: 3 LKLATTFPISASNADQSSRRPTGKPN--KNPSKPKVDPQSHPALKFSNIPKQKLKPVNKT 60
+ L+ ISA N SS P KP +P P P P S P +K +
Sbjct: 63 VSLSLPSQISAHNYLYSSPPPPKKPYYYHSPPPPSPPPPKKPYYYHSPPPPKKPYYYHSP 242
Query: 61 PENVKISEDGVSYVIEGAPFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPL 120
P P++ Y Y P P ++P + P +P+ P
Sbjct: 243 PPPSP-----------PPPYKKPYYYHSPPPPSPPPPKKPYYYHSPPPPPKKPYYYHSPP 389
Query: 121 PPSKKKLKEFDSFVLPPPHKKGVKPVQSPGPFLPGTSPRY 160
PPS PPP+KK V SP P P P Y
Sbjct: 390 PPSP-----------PPPYKKPYHYV-SPPPPSPSPPPPY 473
>TC204047 hydroxyproline-rich glycoprotein
Length = 887
Score = 37.0 bits (84), Expect = 0.014
Identities = 41/134 (30%), Positives = 46/134 (33%), Gaps = 5/134 (3%)
Frame = +1
Query: 23 PTGKPNKNPSKP----KVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGA 78
P KP K PS P K P HP P +K P VK V
Sbjct: 109 PPKKPYKYPSPPPPVKKPYPHPHPVYHSPPPPPKKPYKYPSPPPPVKKPYPHPHPVYHSP 288
Query: 79 PFEFKYSYTETPKSKPVQMREPPF-VPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPP 137
P P KP + PP V P +P P PP PP +K K + PP
Sbjct: 289 P---------PPPKKPYKYPSPPPPVHTYPPHVPTPVYHSPPPPPYEKPYK----YKSPP 429
Query: 138 PHKKGVKPVQSPGP 151
P PV SP P
Sbjct: 430 P------PVHSPPP 453
Score = 35.4 bits (80), Expect = 0.040
Identities = 40/136 (29%), Positives = 45/136 (32%), Gaps = 4/136 (2%)
Frame = +1
Query: 23 PTGKPNKNPSKP----KVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGA 78
P KP K PS P K P HP P +K P VK V
Sbjct: 16 PPKKPYKYPSPPPPVKKPYPHPHPVYHSPPPPPKKPYKYPSPPPPVKKPYPHPHPVYHSP 195
Query: 79 PFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPP 138
P P KP + PP P P P PP PP KK K + PPP
Sbjct: 196 P---------PPPKKPYKYPSPPPPVKKPYPHPHPVYHSPP-PPPKKPYK----YPSPPP 333
Query: 139 HKKGVKPVQSPGPFLP 154
PV + P +P
Sbjct: 334 ------PVHTYPPHVP 363
Score = 31.2 bits (69), Expect = 0.75
Identities = 25/75 (33%), Positives = 27/75 (35%)
Frame = +1
Query: 84 YSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGV 143
Y P KP + PP P P P PP PP KK K + PPP K
Sbjct: 91 YHSPPPPPKKPYKYPSPPPPVKKPYPHPHPVYHSPP-PPPKKPYK----YPSPPPPVK-- 249
Query: 144 KPVQSPGPFLPGTSP 158
KP P P P
Sbjct: 250 KPYPHPHPVYHSPPP 294
>TC215903 similar to UP|Q9M7N4 (Q9M7N4) MFP1 attachment factor 1, partial
(64%)
Length = 780
Score = 37.0 bits (84), Expect = 0.014
Identities = 37/119 (31%), Positives = 50/119 (41%)
Frame = +3
Query: 20 SRRPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAP 79
S+ PT K + P++ K + P PK P+ TP S S G P
Sbjct: 51 SKCPTRKTHPRPNRSKTPLRHRP-------PKTNPSPL--TPS----SAPPASPSAYGPP 191
Query: 80 FEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPP 138
+ + + SKP PP P VT P T RPPLPP++ + + F S PPP
Sbjct: 192 LSARATPSSLASSKPYP--PPPSSP--NVTGQCPPT-RPPLPPARSRTRPFPSPPAPPP 353
>TC230183 homologue to UP|GLHR_ANTEL (P35409) Probable glycoprotein hormone
G-protein coupled receptor precursor, partial (6%)
Length = 878
Score = 36.6 bits (83), Expect = 0.018
Identities = 25/74 (33%), Positives = 28/74 (37%), Gaps = 5/74 (6%)
Frame = -1
Query: 90 PKSKPVQMREPPFVPFGPVTM-----PRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVK 144
P KP ++ PP P P P P G+PP PP K PPP K G
Sbjct: 467 PPPKPGLLKPPPPKPGKPPPP*PGKPPPPKPGKPPPPPKPGK---------PPPPKPGEP 315
Query: 145 PVQSPGPFLPGTSP 158
P PG P P
Sbjct: 314 PPPKPGNLPPPPKP 273
Score = 34.3 bits (77), Expect = 0.088
Identities = 23/62 (37%), Positives = 27/62 (43%), Gaps = 3/62 (4%)
Frame = -1
Query: 100 PPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKPVQSPG---PFLPGT 156
PP +P P T P+P +PP P K + PPP K G P PG P PG
Sbjct: 542 PPQMP--PPTPPKPGLLKPPPKPGLLKPPPKPGLLKPPPPKPGKPPPP*PGKPPPPKPGK 369
Query: 157 SP 158
P
Sbjct: 368 PP 363
Score = 30.0 bits (66), Expect = 1.7
Identities = 26/70 (37%), Positives = 31/70 (44%), Gaps = 1/70 (1%)
Frame = -1
Query: 91 KSKPVQMREPPFVPFGPVTMPRP-WTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKPVQSP 149
K P + EPP P P +P P G+PP PP KL P P K KP + P
Sbjct: 344 KPPPPKPGEPP--PPKPGNLPPPPKPGKPPPPPKPGKLP------TPKPGKPPPKPGEPP 189
Query: 150 GPFLPGTSPR 159
P P +PR
Sbjct: 188 -PKPP*LTPR 162
>TC215069 similar to UP|O24300 (O24300) PtxA protein precursor, partial (83%)
Length = 1405
Score = 36.6 bits (83), Expect = 0.018
Identities = 40/150 (26%), Positives = 52/150 (34%), Gaps = 15/150 (10%)
Frame = +2
Query: 27 PNKNPSKPK-VDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFEFKYS 85
P P KP V P H K PK KP + P V + P
Sbjct: 125 PKPPPKKPPIVKPPVHKQPKPCPPPKSSPKPPHVHPPYVPKPPHYPKPPVHPHP-----P 289
Query: 86 YTETPKSKPVQMREPPFVPFGPVTMPR----------PWTGRPPL----PPSKKKLKEFD 131
+ P KP PP+VP PV P P+ +PP+ PP K +
Sbjct: 290 HVPKPHPKPPLHPHPPYVPKPPVVKPPYVPKPPVVNPPYVPKPPVVPVTPPYVPKPPIVN 469
Query: 132 SFVLPPPHKKGVKPVQSPGPFLPGTSPRYV 161
+P P V+P P P + +P YV
Sbjct: 470 PPYVPKPPVVPVRPPYIPKPPVVPVTPPYV 559
Score = 31.6 bits (70), Expect = 0.57
Identities = 35/134 (26%), Positives = 50/134 (37%), Gaps = 6/134 (4%)
Frame = +2
Query: 22 RPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVS------YVI 75
+P P +P P V P P K + PK PV+ P +V YV
Sbjct: 182 KPCPPPKSSPKPPHVHPPYVP--KPPHYPKP---PVHPHPPHVPKPHPKPPLHPHPPYVP 346
Query: 76 EGAPFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVL 135
+ + Y + P P + +PP VP P +P+P PP P + ++
Sbjct: 347 KPPVVKPPY-VPKPPVVNPPYVPKPPVVPVTPPYVPKPPIVNPPYVPKPPVVPVRPPYIP 523
Query: 136 PPPHKKGVKPVQSP 149
PP V PV P
Sbjct: 524 KPP----VVPVTPP 553
Score = 31.2 bits (69), Expect = 0.75
Identities = 38/140 (27%), Positives = 52/140 (37%), Gaps = 3/140 (2%)
Frame = +2
Query: 22 RPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFE 81
+P KP +P P V P +K +PK PV P K V+ P
Sbjct: 299 KPHPKPPLHPHPPYVPKP--PVVKPPYVPKP---PVVNPPYVPKPPVVPVTPPYVPKPPI 463
Query: 82 FKYSYTETPKSKPVQ---MREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPP 138
Y P PV+ + +PP VP P +P+P PP+ P+ V PP
Sbjct: 464 VNPPYVPKPPVVPVRPPYIPKPPVVPVTPPYVPKPPIVYPPVVPT-------PPIVTPPI 622
Query: 139 HKKGVKPVQSPGPFLPGTSP 158
V PV P + +P
Sbjct: 623 VYPPVVPVPPLPPIVTPPNP 682
>TC214303 similar to UP|Q9T0K5 (Q9T0K5) Extensin-like protein, partial (18%)
Length = 1014
Score = 35.8 bits (81), Expect = 0.030
Identities = 24/69 (34%), Positives = 26/69 (36%), Gaps = 1/69 (1%)
Frame = +2
Query: 84 YSYTETPKSKPVQMREPP-FVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKG 142
Y Y P PV PP + P P P P PP PP +E S P PH
Sbjct: 116 YPYLSPPPPPPVHSPPPPVYSPPPPPPSPPPCIESPPPPPPPPPCEE-HSPPPPSPHSAP 292
Query: 143 VKPVQSPGP 151
P SP P
Sbjct: 293 YHPPPSPSP 319
Score = 27.7 bits (60), Expect = 8.3
Identities = 21/59 (35%), Positives = 22/59 (36%)
Frame = +2
Query: 100 PPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKPVQSPGPFLPGTSP 158
P + P P P P PP PP L S PPP PV SP P P P
Sbjct: 44 PTYSPPPPPPSPPPPHSPPPPPPVYPYL----SPPPPPPVHSPPPPVYSPPPPPPSPPP 208
>CF921606
Length = 518
Score = 35.8 bits (81), Expect = 0.030
Identities = 24/67 (35%), Positives = 29/67 (42%), Gaps = 2/67 (2%)
Frame = +2
Query: 90 PKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPH--KKGVKPVQ 147
P K +++PP G +P+ T PP PP KKK PPPH KKG P
Sbjct: 2 PPQKKKNLKKPPHRGGGKSPLPKKPTPPPPSPPKKKK--------GPPPH*KKKGGAPPT 157
Query: 148 SPGPFLP 154
G P
Sbjct: 158 RGGETPP 178
>TC204807 weakly similar to UP|HQGT_RAUSE (Q9AR73) Hydroquinone
glucosyltransferase (Arbutin synthase) , partial (79%)
Length = 1673
Score = 30.4 bits (67), Expect = 1.3
Identities = 25/93 (26%), Positives = 35/93 (36%), Gaps = 23/93 (24%)
Frame = +1
Query: 100 PPFVPFGPVTMPRPWTGRPPLPPSKKKLKEF----------------------DSFVLPP 137
PP F P ++P P + PL ++ L VLPP
Sbjct: 514 PPLPSFPPPSIPLPLRHKHPLRRRRRPLLHRRLRRRRRIQCLPLRLLSLHCHRPLLVLPP 693
Query: 138 PHKKGVKPVQSPGPFLPGTSPR-YVMSREEVLG 169
PH + PV+ P P G PR + SR+ +G
Sbjct: 694 PHPRPAGPVRVPRPPRTGQHPRLHSSSRQGPVG 792
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.137 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,194,425
Number of Sequences: 63676
Number of extensions: 311532
Number of successful extensions: 3210
Number of sequences better than 10.0: 205
Number of HSP's better than 10.0 without gapping: 2630
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2945
length of query: 355
length of database: 12,639,632
effective HSP length: 98
effective length of query: 257
effective length of database: 6,399,384
effective search space: 1644641688
effective search space used: 1644641688
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC137546.15


