

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137546.13 - phase: 0
(363 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
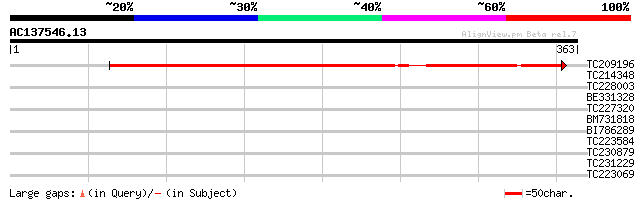
Score E
Sequences producing significant alignments: (bits) Value
TC209196 398 e-111
TC214348 UP|Q39832 (Q39832) Ribulose-1,5-bisphosphate carboxylas... 31 1.0
TC228003 similar to UP|Q94F26 (Q94F26) Lipase-like protein, part... 30 1.3
BE331328 similar to GP|3928140|emb chlorophyll a/b binding prote... 28 5.0
TC227320 similar to UP|Q93YF1 (Q93YF1) Nucleic acid binding prot... 28 5.0
BM731818 28 5.0
BI786289 28 6.5
TC223584 weakly similar to UP|RB87_DROME (P48810) Heterogeneous ... 28 6.5
TC230879 similar to GB|AAT41775.1|48310209|BT014792 At2g39000 {A... 28 6.5
TC231229 28 8.5
TC223069 similar to UP|Q71LW1 (Q71LW1) Lecithin cholesterol acyl... 28 8.5
>TC209196
Length = 1098
Score = 398 bits (1023), Expect = e-111
Identities = 199/293 (67%), Positives = 244/293 (82%), Gaps = 1/293 (0%)
Frame = +1
Query: 65 GLFSNHGPLYFVATKRFFSKMGLACHIAKVHSEASVEHNAMEIKQYIEEIYWGSGKPVML 124
GLFSNHGPLYFV+TK FSKMGLACHIAK+HSEASVE NA E+K+YIEEIYWGS K VML
Sbjct: 7 GLFSNHGPLYFVSTKVSFSKMGLACHIAKIHSEASVEKNAKELKEYIEEIYWGSNKRVML 186
Query: 125 LGHSKGGIDAAAALSLYWSDLKGKVAGLALVQSPYGGTPIASDILREGQIGD-KETRRIL 183
LGHSKGG+DAAAALSLYWSDLK KVAGLAL QSPYGGTPIASD+LREGQ+GD R++
Sbjct: 187 LGHSKGGVDAAAALSLYWSDLKDKVAGLALAQSPYGGTPIASDLLREGQLGDYVNLRKLT 366
Query: 184 ELIICKIIKGDIRALEDLTYEKRKDFIMKHKLPLDIPLISFRSEASITPSVLATMTQIAH 243
E++ICK+IKGD+RALEDLTYE+R++F+ +H LP ++P++SFR+EA I+P+VLAT++ +AH
Sbjct: 367 EILICKVIKGDMRALEDLTYERRREFLKEHHLPKEVPIVSFRTEAGISPAVLATLSHVAH 546
Query: 244 AELPRLILPKFGSKVSDQFVESGRQVPVMVPVSAAMAAFALHLQLRYGEKSDGVVTCRDA 303
AELP L+ P S R++P+++P+ AAMAA A LQ+RYGEKSDG+VTCRDA
Sbjct: 547 AELP-LVAPGGES----------RKLPLVMPLGAAMAACAQLLQVRYGEKSDGLVTCRDA 693
Query: 304 EVPGSVVVRPNMKLDHAWMVYSSNSKKKKSSEPDAREMCQAIFTLLVELGKTE 356
EVPGS+VVRP KLDHAWMVYS S SE DA ++C+A+ TLLVE+G+T+
Sbjct: 694 EVPGSIVVRPKRKLDHAWMVYS--SLNDDPSEGDASQVCEALLTLLVEIGQTK 846
>TC214348 UP|Q39832 (Q39832) Ribulose-1,5-bisphosphate carboxylase small
subunit precursor (Ribulose-1,5-bisphosphate carboxylase
small subunit rbcS1), partial (21%)
Length = 768
Score = 30.8 bits (68), Expect = 1.0
Identities = 21/76 (27%), Positives = 38/76 (49%), Gaps = 5/76 (6%)
Frame = +3
Query: 90 HIAKVHSEASVEHNAMEIKQYIEEIY-----WGSGKPVMLLGHSKGGIDAAAALSLYWSD 144
H+A + S ++E N+ I +Y E + + V+L+GHS GG+ + A+ L+
Sbjct: 405 HLASIKSRRAIELNS--ITEYFEPLMEFLLSLAEEEQVILVGHSFGGLCISVAMELF--- 569
Query: 145 LKGKVAGLALVQSPYG 160
K+A ++P G
Sbjct: 570 -PTKIAAAVFDKTPRG 614
>TC228003 similar to UP|Q94F26 (Q94F26) Lipase-like protein, partial (92%)
Length = 1444
Score = 30.4 bits (67), Expect = 1.3
Identities = 25/73 (34%), Positives = 37/73 (50%), Gaps = 3/73 (4%)
Frame = +1
Query: 97 EASVEHNAMEIKQYIEEIYWGSGKPVMLLGHSKGG---IDAAAALSLYWSDLKGKVAGLA 153
+ SVE ++ I+E+Y S ++L+GHS GG + AA SL +AGL
Sbjct: 445 DLSVETMCNDVLAVIKELYSDSPPAIILVGHSMGGSIAVHVAARRSL------SALAGLV 606
Query: 154 LVQSPYGGTPIAS 166
+V GT +AS
Sbjct: 607 VV-DVVEGTAMAS 642
>BE331328 similar to GP|3928140|emb chlorophyll a/b binding protein {Cicer
arietinum}, partial (52%)
Length = 421
Score = 28.5 bits (62), Expect = 5.0
Identities = 15/41 (36%), Positives = 26/41 (62%), Gaps = 1/41 (2%)
Frame = +2
Query: 35 DGSSRFLEL-LSDIRNGKDSIPSSFVYLLIPGLFSNHGPLY 74
D F EL +++++NG+ ++ S FVY L+ + + GPLY
Sbjct: 281 DNPYAFAELNVNELKNGRLAMSSMFVY-LVKAIVTEKGPLY 400
>TC227320 similar to UP|Q93YF1 (Q93YF1) Nucleic acid binding protein, partial
(70%)
Length = 1724
Score = 28.5 bits (62), Expect = 5.0
Identities = 15/32 (46%), Positives = 19/32 (58%)
Frame = -1
Query: 219 IPLISFRSEASITPSVLATMTQIAHAELPRLI 250
IP+IS RS +TP+ L MT ELP+ I
Sbjct: 1658 IPIISLRSSLVMTPTGLNKMTN*IPTELPQAI 1563
>BM731818
Length = 333
Score = 28.5 bits (62), Expect = 5.0
Identities = 15/32 (46%), Positives = 19/32 (58%)
Frame = -3
Query: 219 IPLISFRSEASITPSVLATMTQIAHAELPRLI 250
IP+IS RS +TP+ L MT ELP+ I
Sbjct: 244 IPIISLRSSLVMTPTGLNKMTN*IPTELPQAI 149
>BI786289
Length = 421
Score = 28.1 bits (61), Expect = 6.5
Identities = 13/31 (41%), Positives = 18/31 (57%)
Frame = +2
Query: 32 PVHDGSSRFLELLSDIRNGKDSIPSSFVYLL 62
P H+ + + L SD GKD IPSS + +L
Sbjct: 302 PFHNSLDKEMSLYSDHHCGKDQIPSSTLQIL 394
>TC223584 weakly similar to UP|RB87_DROME (P48810) Heterogeneous nuclear
ribonucleoprotein 87F (HRP36.1 protein) (P11 protein),
partial (8%)
Length = 432
Score = 28.1 bits (61), Expect = 6.5
Identities = 15/48 (31%), Positives = 26/48 (53%), Gaps = 4/48 (8%)
Frame = -3
Query: 28 PGMPPVHDGSSRFL----ELLSDIRNGKDSIPSSFVYLLIPGLFSNHG 71
PG+P +HD R+L ++L ++RN + S + + YL G + G
Sbjct: 322 PGLPVLHDNGIRYLTVFTKVLDELRNRRFSR*TPYEYLSGTGAVAGEG 179
>TC230879 similar to GB|AAT41775.1|48310209|BT014792 At2g39000 {Arabidopsis
thaliana;} , partial (74%)
Length = 832
Score = 28.1 bits (61), Expect = 6.5
Identities = 20/57 (35%), Positives = 29/57 (50%), Gaps = 2/57 (3%)
Frame = +3
Query: 64 PGLFSNHGPLYFVATKRFFS--KMGLACHIAKVHSEASVEHNAMEIKQYIEEIYWGS 118
PG FS G + +K S K+GL+CH + E S HN ++ + +IY GS
Sbjct: 432 PGKFSEKGDS*EIDSKGRVSCQKLGLSCHCIAL*LEQSCGHNVIQRTRL--QIYHGS 596
>TC231229
Length = 509
Score = 27.7 bits (60), Expect = 8.5
Identities = 23/96 (23%), Positives = 43/96 (43%), Gaps = 3/96 (3%)
Frame = +3
Query: 122 VMLLGHSKGGIDAAAALSLYWSDLKGKVAGLALVQSPYGGTPIASDILREGQIGD---KE 178
V+L G +K A+ +Y ++ K A AL+ PI D+L I + K
Sbjct: 186 VILEGDAKRRALASIKSLIYQPSVRNKKATFALICQTLKHLPIIKDVLEAASILNTKWKR 365
Query: 179 TRRILELIICKIIKGDIRALEDLTYEKRKDFIMKHK 214
R ++ +I+ I+ G +L + ++M+H+
Sbjct: 366 QRELIYIIVYDILLGQAVSL----VGDAEKYLMRHQ 461
>TC223069 similar to UP|Q71LW1 (Q71LW1) Lecithin cholesterol
acyltransferase-like protein, partial (44%)
Length = 659
Score = 27.7 bits (60), Expect = 8.5
Identities = 18/58 (31%), Positives = 27/58 (46%), Gaps = 1/58 (1%)
Frame = +2
Query: 107 IKQYIEEIYWGSG-KPVMLLGHSKGGIDAAAALSLYWSDLKGKVAGLALVQSPYGGTP 163
+K +E + SG + V L+ HS GGI + +SLY V + P+ G P
Sbjct: 467 LKSKLETAHKASGGRKVNLISHSMGGIMISCFMSLYRDVFTKYVNKWICLACPFQGAP 640
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.136 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,639,149
Number of Sequences: 63676
Number of extensions: 193038
Number of successful extensions: 901
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 894
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 898
length of query: 363
length of database: 12,639,632
effective HSP length: 98
effective length of query: 265
effective length of database: 6,399,384
effective search space: 1695836760
effective search space used: 1695836760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC137546.13


