

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137081.4 - phase: 0
(325 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
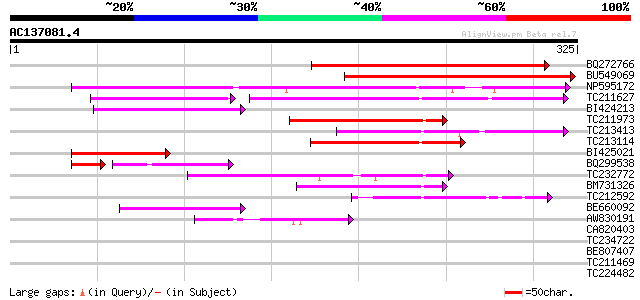
Score E
Sequences producing significant alignments: (bits) Value
BQ272766 weakly similar to GP|28558781|gb| pol protein {Cucumis ... 192 1e-49
BU549069 189 1e-48
NP595172 polyprotein [Glycine max] 154 4e-38
TC211627 84 1e-30
BI424213 77 1e-14
TC211973 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, parti... 70 1e-12
TC213413 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, parti... 67 8e-12
TC213114 weakly similar to UP|Q8W150 (Q8W150) Polyprotein, parti... 67 1e-11
BI425021 66 2e-11
BQ299538 51 2e-10
TC232772 weakly similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial ... 59 4e-09
BM731326 weakly similar to GP|21740635|em OSJNBb0043H09.2 {Oryza... 54 7e-08
TC212592 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, parti... 54 7e-08
BE660092 weakly similar to GP|9884624|dbj retroelement pol polyp... 45 3e-05
AW830191 42 5e-04
CA820403 weakly similar to GP|13273463|gb| pol protein integrase... 39 0.003
TC234722 similar to UP|Q6WAY9 (Q6WAY9) Pol (Fragment), partial (... 39 0.004
BE807407 33 0.18
TC211469 32 0.39
TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 31 0.67
>BQ272766 weakly similar to GP|28558781|gb| pol protein {Cucumis melo},
partial (9%)
Length = 410
Score = 192 bits (489), Expect = 1e-49
Identities = 92/136 (67%), Positives = 113/136 (82%)
Frame = -3
Query: 174 SYHDKRRKDLEFQEGDHVFLRVTPLTGVGRALKSRKLTPKFIGPYQISERVGTVAYRVGL 233
SYHDKRRKDLEF+ GDHVFLRVTP TGVGRALKS KLTP FIGP+QI ++V VAY++ L
Sbjct: 408 SYHDKRRKDLEFEVGDHVFLRVTP*TGVGRALKS*KLTPHFIGPFQILKKVDFVAYQIAL 229
Query: 234 PPHLSNLHDVFHVSQLRKYVADPSHVIPRDDVQVRDNLTVETMPLRIDNRKVKSLRGKEI 293
PP L++LH+VFHVSQ+ KY+ DPSH++ D VQV++NL ET+PLRI + + K LRGKEI
Sbjct: 228 PPSLTSLHNVFHVSQIHKYINDPSHMVNLDVVQVKENLAHETLPLRIKDMRTKHLRGKEI 49
Query: 294 PLVRVVWGGATGESLT 309
LV+V+WGGA+GE T
Sbjct: 48 LLVKVIWGGASGEEAT 1
>BU549069
Length = 615
Score = 189 bits (480), Expect = 1e-48
Identities = 89/132 (67%), Positives = 109/132 (82%)
Frame = -1
Query: 193 LRVTPLTGVGRALKSRKLTPKFIGPYQISERVGTVAYRVGLPPHLSNLHDVFHVSQLRKY 252
L+VTP TGVG+ALKSRKLTP FIG +QI +R VAY++ LPP LSNLH+VFHVSQLR Y
Sbjct: 615 LKVTPRTGVGQALKSRKLTPHFIGHFQILKRAXPVAYQIALPPSLSNLHNVFHVSQLRMY 436
Query: 253 VADPSHVIPRDDVQVRDNLTVETMPLRIDNRKVKSLRGKEIPLVRVVWGGATGESLTWEL 312
+ DPSHV+ DDVQV++NLT ET+PLRI++R+ K LR KE PLV+V+WGG +GE TWEL
Sbjct: 435 IHDPSHVVKLDDVQVKENLTYETLPLRIEDRRTKHLRRKENPLVKVIWGGTSGEDATWEL 256
Query: 313 ESKMRESYPELF 324
ES+MR +YP LF
Sbjct: 255 ESQMRVAYPSLF 220
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 154 bits (390), Expect = 4e-38
Identities = 104/298 (34%), Positives = 153/298 (50%), Gaps = 12/298 (4%)
Frame = +1
Query: 36 PSSIVSDRDPRFTSRFWKSLQDALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRVCVLEQ 95
P SIVSDRD FTS FW+ L G+ L +SSAYHPQ+DGQSE + LE LR E
Sbjct: 3607 PRSIVSDRDRVFTSTFWQHLFKLQGTTLAMSSAYHPQSDGQSEVLNKCLEMYLRCFTYEH 3786
Query: 96 GGASDSHLPLIEFTYNNSYHSSIGMAPFEALYGWRCRTPLCWFESGESVVLGPDLVHETT 155
LP EF YN +YH S+GM PF ALYG R P S+ ++ + T
Sbjct: 3787 PKGWVKALPWAEFWYNTAYHMSLGMTPFRALYG---REPPTLTRQACSIDDPAEVREQLT 3957
Query: 156 EK---VRMIREKMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPLTGVGRAL-KSRKLT 211
++ + ++ + +Q K DK+R D+ FQ GD V +++ P L K++KL+
Sbjct: 3958 DRDALLAKLKINLTRAQQVMKRQADKKRLDVSFQIGDEVLVKLQPYRQHSAVLRKNQKLS 4137
Query: 212 PKFIGPYQISERVGTVAYRVGLPPHLSNLHDVFHVSQLRKY---VADPSHVIPRDDVQVR 268
++ GP+++ ++G VAY++ L P + +H VFHVSQL+ + DP +P
Sbjct: 4138 MRYFGPFKVLAKIGDVAYKLEL-PSAARIHPVFHVSQLKPFNGTAQDPYLPLP------- 4293
Query: 269 DNLTVETM-----PLRIDNRKVKSLRGKEIPLVRVVWGGATGESLTWELESKMRESYP 321
LTV M P++I ++ +I + V W + TWE ++ SYP
Sbjct: 4294 --LTVTEMGPVMQPVKILASRIIIRGHNQIEQILVQWENGLQDEATWEDIEDIKASYP 4461
>TC211627
Length = 1034
Score = 84.3 bits (207), Expect(2) = 1e-30
Identities = 57/185 (30%), Positives = 89/185 (47%), Gaps = 2/185 (1%)
Frame = +3
Query: 138 FESGESVVLGPD-LVHETTEKVRMIREKMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVT 196
+ +G S V D ++H + +++ Q K D R+DL F GD V++R+
Sbjct: 276 YSAGTSTVEAVDAILHSLATIHHTLTCRLQKYQDSMKRIADSHRRDLTFNIGDWVYVRL* 455
Query: 197 PLTGVGRALKSRKLTPKFIGPYQISERVGTVAYRVGLPPHLSNLHDVFHVSQLRKYVAD- 255
P KL+ +F GPYQI RVG VAYR+ LPP S +H +FHVS L+ +
Sbjct: 456 PYRQTSIQSTYTKLSKRFYGPYQIQARVGQVAYRLQLPP-TSKIHPIFHVSLLKVHHGPI 632
Query: 256 PSHVIPRDDVQVRDNLTVETMPLRIDNRKVKSLRGKEIPLVRVVWGGATGESLTWELESK 315
P ++ ++ V+ PL+ + K+ IP V V W E TWE ++
Sbjct: 633 PPELLALPPFSTTNHPLVQ--PLQFLDWKMDESTTPPIPQVLVQWTNLAPEDTTWESWTQ 806
Query: 316 MRESY 320
+++ Y
Sbjct: 807 LKDIY 821
Score = 67.0 bits (162), Expect(2) = 1e-30
Identities = 39/83 (46%), Positives = 45/83 (53%)
Frame = +2
Query: 47 FTSRFWKSLQDALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRVCVLEQGGASDSHLPLI 106
F S W L G+KLR S+AYHPQTDGQ+E + LE LR V + L L
Sbjct: 5 FISGLWHELFHISGTKLRFSTAYHPQTDGQTEVINRILEQYLRAFVHDHPQHWFKFLSLA 184
Query: 107 EFTYNNSYHSSIGMAPFEALYGW 129
E YN S HS IG +PFE Y W
Sbjct: 185 E*CYNTSVHSGIGFSPFEC-YVW 250
>BI424213
Length = 426
Score = 77.0 bits (188), Expect = 1e-14
Identities = 38/87 (43%), Positives = 52/87 (59%)
Frame = +1
Query: 49 SRFWKSLQDALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRVCVLEQGGASDSHLPLIEF 108
S FWK+L LG+KL S+ HPQTDGQ++ +SL LLR + + D +LP +EF
Sbjct: 4 SHFWKTLWAKLGTKLLFSTTCHPQTDGQTKVVNRSLSTLLRALLKGNHKSWDEYLPHVEF 183
Query: 109 TYNNSYHSSIGMAPFEALYGWRCRTPL 135
YN H + +PFE +YG+ TPL
Sbjct: 184 AYNRGVHRTTKQSPFEVVYGFNPLTPL 264
>TC211973 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, partial (4%)
Length = 730
Score = 70.5 bits (171), Expect = 1e-12
Identities = 37/92 (40%), Positives = 59/92 (63%), Gaps = 1/92 (1%)
Frame = +1
Query: 161 IREKMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPLTGVGRALK-SRKLTPKFIGPYQ 219
+RE + S + +KRR+D+E+ GD VFL++ P A + + KL+P+F P+Q
Sbjct: 64 LRENLLKS*DIM*ANTNKRRRDIEYVVGD*VFLKMQPYRRRSLAKRINEKLSPRFYAPFQ 243
Query: 220 ISERVGTVAYRVGLPPHLSNLHDVFHVSQLRK 251
+ +VGT+AY++ LP H+ +H VFHVS L+K
Sbjct: 244 VFNKVGTIAYKLDLPSHI-KIHPVFHVSLLKK 336
>TC213413 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, partial (5%)
Length = 761
Score = 67.4 bits (163), Expect = 8e-12
Identities = 43/136 (31%), Positives = 68/136 (49%), Gaps = 3/136 (2%)
Frame = +1
Query: 188 GDHVFLRVTP-LTGVGRALKSRKLTPKFIGPYQISERVGTVAYRVGLPPHLSNLHDVFHV 246
GD V +++ P G KLT ++ GP+++ ER+G V YR+ L H S +H VFHV
Sbjct: 7 GDWVLVKLRPHRQGSASETTYSKLTKRYYGPFEVQERLGKVVYRLKLTAH-SRIHPVFHV 183
Query: 247 SQLRKYVADP--SHVIPRDDVQVRDNLTVETMPLRIDNRKVKSLRGKEIPLVRVVWGGAT 304
S L+ +V DP +H P ++ + PL + + K+ +V V W A+
Sbjct: 184 SLLKAFVGDPETTHAGPLPVMRTEE---ATNTPLTVIDSKLVPADNGPRRMVLVQWPSAS 354
Query: 305 GESLTWELESKMRESY 320
+ +WE +RE Y
Sbjct: 355 RQDASWEDWQVLRERY 402
>TC213114 weakly similar to UP|Q8W150 (Q8W150) Polyprotein, partial (7%)
Length = 810
Score = 67.0 bits (162), Expect = 1e-11
Identities = 36/90 (40%), Positives = 55/90 (61%), Gaps = 1/90 (1%)
Frame = +3
Query: 173 KSYHDKRRKDLEFQEGDHVFLRVTPLTGVGRALK-SRKLTPKFIGPYQISERVGTVAYRV 231
K+Y D+ R+ + GD V+L++ P A K + KL+P+F GPYQI +++G VA+ +
Sbjct: 6 KAYADRSRRAVTLSVGDWVYLKLQPYRLKSLAKKRNEKLSPRFYGPYQIKKQIGLVAFEL 185
Query: 232 GLPPHLSNLHDVFHVSQLRKYVADPSHVIP 261
LPP +H VFH S L+K VA ++ P
Sbjct: 186 DLPP-ARKIHPVFHASLLKKAVAATANPQP 272
>BI425021
Length = 426
Score = 65.9 bits (159), Expect = 2e-11
Identities = 33/57 (57%), Positives = 39/57 (67%)
Frame = -1
Query: 36 PSSIVSDRDPRFTSRFWKSLQDALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRVCV 92
P SIVSDRDP F S FW+ L G+ LR+SSAYHPQTDGQ+E + +E LR V
Sbjct: 216 PRSIVSDRDPLFISHFWQDLFRLSGTVLRMSSAYHPQTDGQTEVLNRVIEQYLRAFV 46
>BQ299538
Length = 426
Score = 50.8 bits (120), Expect(2) = 2e-10
Identities = 27/69 (39%), Positives = 39/69 (56%)
Frame = +3
Query: 60 GSKLRLSSAYHPQTDGQSERTIQSLEDLLRVCVLEQGGASDSHLPLIEFTYNNSYHSSIG 119
G+ L++S++YHP DGQ+ LE LR V +Q L E+ YN ++H+S G
Sbjct: 144 GTYLKMSTSYHP*IDGQTVN--HCLETFLRCFVADQPKM*VQWLSWAEYWYNTNFHASTG 317
Query: 120 MAPFEALYG 128
PFE +YG
Sbjct: 318 TTPFEVVYG 344
Score = 32.0 bits (71), Expect(2) = 2e-10
Identities = 12/20 (60%), Positives = 15/20 (75%)
Frame = +1
Query: 36 PSSIVSDRDPRFTSRFWKSL 55
P+S++SD DP F S FWK L
Sbjct: 73 PASVLSDEDPIFVSSFWKEL 132
>TC232772 weakly similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (7%)
Length = 729
Score = 58.5 bits (140), Expect = 4e-09
Identities = 42/158 (26%), Positives = 73/158 (45%), Gaps = 6/158 (3%)
Frame = +1
Query: 103 LPLIEFTYNNSYHSSIGMAPFEALYGWRCRTPLCWFESGESVVLGPDLVHETTEKVRMIR 162
LP +EF YN + HS+ PFE +Y + TPL H E ++ +
Sbjct: 7 LPHVEFAYNRAVHSTTQHFPFEVVYDFNPLTPLDSLPLSNISGFKHKDAHAKVEYIKRLH 186
Query: 163 EKMKASQSRQKSYH----DKRRKDLEFQEGDHVFLRVTPLTGVGRALKSR--KLTPKFIG 216
E+ K +++ + +K RK + + D V++ + R K R KL P+ G
Sbjct: 187 EQAKTQIAKKNESYVKQTNKNRKKVVLEPSDWVWVHMRK----ERFPKQRMSKLQPRGDG 354
Query: 217 PYQISERVGTVAYRVGLPPHLSNLHDVFHVSQLRKYVA 254
P+Q+ ER+ AY++ +P + F+++ L +VA
Sbjct: 355 PFQVLERINYNAYKIDIPGEY-EVSSSFNIADLTPFVA 465
>BM731326 weakly similar to GP|21740635|em OSJNBb0043H09.2 {Oryza sativa
(japonica cultivar-group)}, partial (3%)
Length = 424
Score = 54.3 bits (129), Expect = 7e-08
Identities = 30/88 (34%), Positives = 50/88 (56%), Gaps = 1/88 (1%)
Frame = +1
Query: 165 MKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTP-LTGVGRALKSRKLTPKFIGPYQISER 223
++ +QS + + R+ + GD V+L++ P G KLT +F GPY + +
Sbjct: 25 LERAQSLMVKHANNHRRPHDINVGDWVYLKIRPHRQGSMPPRLHPKLTARFYGPYLVMRQ 204
Query: 224 VGTVAYRVGLPPHLSNLHDVFHVSQLRK 251
VG VA+++ LP + +H VFHVSQL++
Sbjct: 205 VGAVAFQLQLPSE-ARIHPVFHVSQLKR 285
>TC212592 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, partial (4%)
Length = 664
Score = 54.3 bits (129), Expect = 7e-08
Identities = 40/115 (34%), Positives = 59/115 (50%)
Frame = +2
Query: 197 PLTGVGRALKSRKLTPKFIGPYQISERVGTVAYRVGLPPHLSNLHDVFHVSQLRKYVADP 256
P+TG KL+ +F GP+Q+ ERVG VAYR+ LP + LH VFH S L+ + +P
Sbjct: 62 PITG--------KLSKRFFGPFQVVERVGKVAYRLQLPVD-AKLHPVFHCSLLKPFQGNP 214
Query: 257 SHVIPRDDVQVRDNLTVETMPLRIDNRKVKSLRGKEIPLVRVVWGGATGESLTWE 311
+ D+ +V PL I +++ +I V V W G + + TWE
Sbjct: 215 PDTAAPLPPTLFDHQSV-IAPLVI--LATRTVNDDDIE-VLVQWQGLSPDDATWE 367
>BE660092 weakly similar to GP|9884624|dbj retroelement pol polyprotein-like
{Arabidopsis thaliana}, partial (13%)
Length = 378
Score = 45.4 bits (106), Expect = 3e-05
Identities = 21/72 (29%), Positives = 40/72 (55%)
Frame = -3
Query: 64 RLSSAYHPQTDGQSERTIQSLEDLLRVCVLEQGGASDSHLPLIEFTYNNSYHSSIGMAPF 123
R+S+ YHPQT+GQ+E + + ++ +L V + L + + +Y + IGM+P+
Sbjct: 340 RVSTPYHPQTNGQAEISNREIKRILEKIVQPSRKDWSTRLDDALWAHRTAYKAPIGMSPY 161
Query: 124 EALYGWRCRTPL 135
++G C P+
Sbjct: 160 RVVFGKACHLPV 125
>AW830191
Length = 372
Score = 41.6 bits (96), Expect = 5e-04
Identities = 28/101 (27%), Positives = 49/101 (47%), Gaps = 10/101 (9%)
Frame = +3
Query: 107 EFTYNNSYHSSIGMAPFEALYGWRCRTPLCWFESGESVVLGPDLVHETTEKVRMI---RE 163
EF +N +Y++S+ + PF+ LYG C P +L ++ T E+V ++ R+
Sbjct: 63 EFWFNTNYNNSLKLTPFKVLYG--CDPP---------HLLKGAIISSTAEEVNVMTNDRD 209
Query: 164 KM-------KASQSRQKSYHDKRRKDLEFQEGDHVFLRVTP 197
+M A Q Y D+ R+ + GD V+L++ P
Sbjct: 210 QMLHDLKGNLAKAQNQMKYADRSRRSIPLNVGDWVYLKLQP 332
>CA820403 weakly similar to GP|13273463|gb| pol protein integrase region
{Ginkgo biloba}, partial (52%)
Length = 421
Score = 38.9 bits (89), Expect = 0.003
Identities = 23/50 (46%), Positives = 29/50 (58%), Gaps = 1/50 (2%)
Frame = -3
Query: 37 SSIVSDRDPRFT-SRFWKSLQDALGSKLRLSSAYHPQTDGQSERTIQSLE 85
SSIVSD D F S FW L G+KL+ S AYHPQ D ++ + +E
Sbjct: 299 SSIVSDWDRLFLIS*FWTELFKMEGTKLKFSLAYHPQPDSHTKVVNRCIE 150
>TC234722 similar to UP|Q6WAY9 (Q6WAY9) Pol (Fragment), partial (32%)
Length = 482
Score = 38.5 bits (88), Expect = 0.004
Identities = 19/63 (30%), Positives = 35/63 (55%)
Frame = -3
Query: 69 YHPQTDGQSERTIQSLEDLLRVCVLEQGGASDSHLPLIEFTYNNSYHSSIGMAPFEALYG 128
YHPQT+GQ+E + + ++ +L V+ L + Y ++ + IG++PF+ +YG
Sbjct: 234 YHPQTNGQAEVSNKEIKRVLENIVVSSRKDWALKLDDAFWAYRIAFKTPIGLSPFQLVYG 55
Query: 129 WRC 131
C
Sbjct: 54 KAC 46
>BE807407
Length = 341
Score = 33.1 bits (74), Expect = 0.18
Identities = 13/28 (46%), Positives = 20/28 (71%)
Frame = +2
Query: 209 KLTPKFIGPYQISERVGTVAYRVGLPPH 236
KLT ++ GPY + +VG VA+++ LP H
Sbjct: 158 KLTARYYGPYLVLIKVGVVAFQLQLPKH 241
>TC211469
Length = 480
Score = 32.0 bits (71), Expect = 0.39
Identities = 25/67 (37%), Positives = 34/67 (50%)
Frame = +2
Query: 254 ADPSHVIPRDDVQVRDNLTVETMPLRIDNRKVKSLRGKEIPLVRVVWGGATGESLTWELE 313
A PS +IP + V D+ V T PL I + K+ S LV V W G E +WEL
Sbjct: 14 APPSDIIPLPPLAV-DHHPVVT-PLTILDWKLDSSVTPPRRLVLVQWMGLPPEDTSWELW 187
Query: 314 SKMRESY 320
+++SY
Sbjct: 188 DDLQQSY 208
>TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 669
Score = 31.2 bits (69), Expect = 0.67
Identities = 33/133 (24%), Positives = 50/133 (36%), Gaps = 15/133 (11%)
Frame = +1
Query: 103 LPLIEFTYNNSYHSSIGMAPFEALYGWRCRTPLCWFE--------SGESVVLGPDLVHET 154
LP Y S +S G PF +YG P FE ES + +
Sbjct: 73 LPFALHGYRTSVRTSTGATPFSLVYGMEAVLP---FEVEVPSLRILAESGLKESEWAQTR 243
Query: 155 TEKVRMIREKM-------KASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPLTGVGRALKS 207
+++ +I K + Q R KS DK+ +F EGD V +++ R
Sbjct: 244 YDQLNLIEGKRLTAMSHGRLYQQRMKSAFDKKVCLRKFHEGDLVLKKMSHAVKDHRG--- 414
Query: 208 RKLTPKFIGPYQI 220
K P + GP+ +
Sbjct: 415 -KWAPNYEGPFVV 450
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.135 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,631,264
Number of Sequences: 63676
Number of extensions: 163099
Number of successful extensions: 857
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 841
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 845
length of query: 325
length of database: 12,639,632
effective HSP length: 97
effective length of query: 228
effective length of database: 6,463,060
effective search space: 1473577680
effective search space used: 1473577680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC137081.4


