

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137079.13 + phase: 0
(357 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
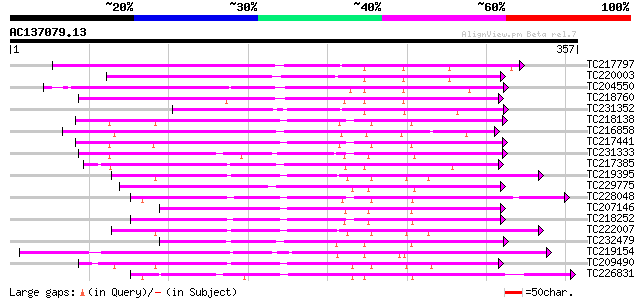
Score E
Sequences producing significant alignments: (bits) Value
TC217797 similar to UP|Q9LVI6 (Q9LVI6) Probable receptor-like pr... 185 2e-47
TC220003 similar to UP|Q6Y2W9 (Q6Y2W9) Atypical receptor-like ki... 159 2e-39
TC204550 UP|Q8L3Y5 (Q8L3Y5) Receptor-like kinase RHG1, complete 152 3e-37
TC218760 similar to UP|O04098 (O04098) Receptor-kinase isolog, 5... 151 5e-37
TC231352 similar to GB|AAN64529.1|24797034|BT001138 At5g58299/At... 145 4e-35
TC218138 homologue to UP|Q8GRK2 (Q8GRK2) Somatic embryogenesis r... 132 3e-31
TC216858 homologue to UP|Q76FZ8 (Q76FZ8) Brassinosteroid recepto... 130 7e-31
TC217441 homologue to GB|AAK68074.1|14573459|AF384970 somatic em... 128 5e-30
TC231333 similar to UP|Q8L741 (Q8L741) AT3g13690/MMM17_12, parti... 126 1e-29
TC217385 similar to UP|Q6ZGC7 (Q6ZGC7) Receptor protein kinase P... 125 2e-29
TC219395 UP|Q9LKZ4 (Q9LKZ4) Receptor-like protein kinase 3, part... 125 3e-29
TC229775 weakly similar to UP|KPEL_DROME (Q05652) Probable serin... 125 3e-29
TC228048 homologue to UP|Q8LA44 (Q8LA44) Receptor protein kinase... 124 5e-29
TC207146 weakly similar to UP|Q9H4D1 (Q9H4D1) Protein kinase, pa... 124 5e-29
TC218252 weakly similar to UP|Q03407 (Q03407) Ydr490cp, partial ... 124 7e-29
TC222007 UP|Q9LKZ5 (Q9LKZ5) Receptor-like protein kinase 2, comp... 123 1e-28
TC232479 similar to UP|Q9FL51 (Q9FL51) Receptor protein kinase-l... 123 1e-28
TC219154 similar to PIR|T05754|T05754 S-receptor kinase M4I22.1... 122 3e-28
TC209490 UP|Q8GSN9 (Q8GSN9) LRR receptor-like kinase (Fragment),... 120 9e-28
TC226831 similar to PIR|T52285|T52285 serine/threonine-specific ... 120 1e-27
>TC217797 similar to UP|Q9LVI6 (Q9LVI6) Probable receptor-like protein kinase
protein (AT3g17840/MEB5_6), partial (44%)
Length = 1612
Score = 185 bits (470), Expect = 2e-47
Identities = 112/313 (35%), Positives = 182/313 (57%), Gaps = 16/313 (5%)
Frame = +2
Query: 28 DNNNSELVFFVEDHERFKLEDLLRATADLRSENFWSSLFKVKFENNVEYAVKRLKNLQVS 87
+ N +LVFF F LEDLLRA+A++ + + + +K E AVKRLK++ +S
Sbjct: 194 EGNAKKLVFFGNAARAFDLEDLLRASAEVLGKGTFGTAYKAVLEAGPVVAVKRLKDVTIS 373
Query: 88 CDEFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLN-DYIARRKDFP 146
EF+E ++ + + H++++ L Y +++EKL++Y Y GS+ LL+ + A R
Sbjct: 374 EKEFKEKIEAVGAMDHESLVPLRAYYFSRDEKLLVYDYMPMGSLSALLHGNKGAGRTPLN 553
Query: 147 WKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFE 206
W++R IA G ARG+ +++ + ++ HGN+K SNILL +A +S+ GL+
Sbjct: 554 WEVRSGIALGAARGIEYLHSR-----GPNVSHGNIKSSNILLTKSYDARVSDFGLAHLVS 718
Query: 207 PDRGTFFSSHGYTAPE----KSLTEKGDVYSFGVILLELLTGQS-----IEVSRIDLVRW 257
P T GY APE + +++K DVYSFGV+LLELLTG++ + +DL RW
Sbjct: 719 PS-STPNRVAGYRAPEVTDPRKVSQKADVYSFGVLLLELLTGKAPTHALLNEEGVDLPRW 895
Query: 258 VRSMVREEWTGEVFDKEV--RENDHQGAFSLLNIALMCVSRSQENRPNFGEILETIEGV- 314
V+S+VREEWT EVFD E+ +N + LL +A+ C ++ + RP+ E++ I+ +
Sbjct: 896 VQSVVREEWTSEVFDLELLRYQNVEEEMVQLLQLAVDCAAQYPDMRPSMSEVVRRIQELR 1075
Query: 315 ---MNAHDQQQME 324
+ DQ Q++
Sbjct: 1076RSSLKEEDQDQIQ 1114
>TC220003 similar to UP|Q6Y2W9 (Q6Y2W9) Atypical receptor-like kinase MARK,
partial (36%)
Length = 1046
Score = 159 bits (401), Expect = 2e-39
Identities = 96/263 (36%), Positives = 155/263 (58%), Gaps = 12/263 (4%)
Frame = +1
Query: 62 WSSLFKVKFENNVEYAVKRLKNLQVSCDEFREILKQISKVKHQNILSLVGYRSTKEEKLI 121
+ + +K E+ AVKRLK++ VS EF+E + + + H+N++ L Y +++EKL+
Sbjct: 10 FGTTYKAVMEDGPVVAVKRLKDVTVSEKEFKEKIDVVGVMDHENLVPLRAYYYSRDEKLL 189
Query: 122 IYKYQSNGSVLNLLN-DYIARRKDFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGN 180
++ Y GS+ +L+ + A R W++R +IA G ARG+ +++ + S+ HGN
Sbjct: 190 VHDYMPMGSLSAILHGNKGAGRTPLNWEMRSSIALGAARGIEYLH-----SQGPSVSHGN 354
Query: 181 LKLSNILLDDKNEALISEHGLSKFFEPDRGTFFSSHGYTAPE----KSLTEKGDVYSFGV 236
+K SNILL +A +S+ GL+ T GY APE + +++K DVYSFGV
Sbjct: 355 IKSSNILLTKSYDARVSDFGLTHLV-GSSSTPNRVAGYRAPEVTDPRKVSQKADVYSFGV 531
Query: 237 ILLELLTGQS-----IEVSRIDLVRWVRSMVREEWTGEVFDKEV--RENDHQGAFSLLNI 289
+LLELLTG++ + +DL RWV+S+VREEW+ EVFD E+ +N + LL +
Sbjct: 532 LLLELLTGKAPTHALLNEEGVDLPRWVQSVVREEWSSEVFDIELLRYQNSEEEMVQLLQL 711
Query: 290 ALMCVSRSQENRPNFGEILETIE 312
A+ CV +NRP+ ++ + IE
Sbjct: 712 AVDCVVPYPDNRPSMSQVRQRIE 780
>TC204550 UP|Q8L3Y5 (Q8L3Y5) Receptor-like kinase RHG1, complete
Length = 2659
Score = 152 bits (383), Expect = 3e-37
Identities = 107/308 (34%), Positives = 164/308 (52%), Gaps = 15/308 (4%)
Frame = +2
Query: 22 GRLKGGDNNNSELVFFVEDHERFKLEDLLRATADLRSENFWSSLFKVKFENNVEYAVKRL 81
GRL+G LV F + F +DLL ATA++ ++ +++K E+ + AVKRL
Sbjct: 1730 GRLEGN------LVHF-DGPMAFTADDLLCATAEIMGKSTNGTVYKAILEDGSQVAVKRL 1888
Query: 82 KN-LQVSCDEFREILKQISKVKHQNILSLVGYR-STKEEKLIIYKYQSNGSVLNLLNDYI 139
+ + EF + + K++H N+L+L Y K EKL+++ Y S GS+ + L+
Sbjct: 1889 REKIAKGHREFESEVSVLGKIRHPNVLALRAYYLGPKGEKLLVFDYMSKGSLASFLHGG- 2065
Query: 140 ARRKDFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEH 199
W R+ IA +ARGL ++ + +I HGNL SN+LLD+ A I++
Sbjct: 2066 GTETFIDWPTRMKIAQDLARGLFCLHSQ------ENIIHGNLTSSNVLLDENTNAKIADF 2227
Query: 200 GLSKFFEPDRGTFF----SSHGYTAPE----KSLTEKGDVYSFGVILLELLTGQS--IEV 249
GLS+ + + GY APE K K D+YS GVILLELLT +S + +
Sbjct: 2228 GLSRLMSTTANSNVIATAGALGYRAPELSKLKKANTKTDIYSLGVILLELLTRKSPGVSM 2407
Query: 250 SRIDLVRWVRSMVREEWTGEVFDKEVRENDHQGAFSLLN---IALMCVSRSQENRPNFGE 306
+ +DL +WV S+V+EEWT EVFD ++ + LLN +AL CV S RP +
Sbjct: 2408 NGLDLPQWVASVVKEEWTNEVFDADLMRDASTVGDELLNTLKLALHCVDPSPSARPEVHQ 2587
Query: 307 ILETIEGV 314
+L+ +E +
Sbjct: 2588 VLQQLEEI 2611
>TC218760 similar to UP|O04098 (O04098) Receptor-kinase isolog, 5' partial;
115640-113643 (Fragment), partial (50%)
Length = 1236
Score = 151 bits (381), Expect = 5e-37
Identities = 100/283 (35%), Positives = 151/283 (53%), Gaps = 15/283 (5%)
Frame = +2
Query: 44 FKLEDLLRATADLRSENFWSSLFKVKFENNVEYAVKRLKNLQV-SCDEFREILKQISKVK 102
+ LEDLL+A+A+ S +K E+ VKRLK+ + + +EFR ++ + +
Sbjct: 83 YSLEDLLKASAETLGRGIMGSTYKAVMESGFIVTVKRLKDARYPALEEFRAHIQVLGSLT 262
Query: 103 HQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLL--NDYIARRKDFPWKLRLNIACGIARG 160
H N++ L Y KEE+L++Y Y NGS+ +L+ + K W L IA +A G
Sbjct: 263 HPNLVPLRAYFQAKEERLLVYDYFPNGSLFSLIHGSKTSGGGKPLHWTSCLKIAEDLATG 442
Query: 161 LAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDR--GTFFSSHGY 218
+ +I++ + HGNLK SN+LL E+ ++++GL+ F PD +S Y
Sbjct: 443 MLYIHQN------PGLTHGNLKSSNVLLGSDFESCLTDYGLTVFLNPDSMDEPSATSLFY 604
Query: 219 TAPE-----KSLTEKGDVYSFGVILLELLTGQS-----IEVSRIDLVRWVRSMVREEWTG 268
APE +S T+ DVYSFGV+LLELLTG++ ++ D+ RWVRS+ EE
Sbjct: 605 RAPECRNFQRSQTQPADVYSFGVLLLELLTGKTPFQDLVQTYGSDIPRWVRSVREEETES 784
Query: 269 EVFDKEVRENDHQGAFSLLNIALMCVSRSQENRPNFGEILETI 311
E + +LLNIA+ CVS ENRP E+L+ I
Sbjct: 785 GDDPASGNEASEEKLQALLNIAMACVSLVPENRPTMREVLKMI 913
>TC231352 similar to GB|AAN64529.1|24797034|BT001138 At5g58299/At5g58299
{Arabidopsis thaliana;} , partial (33%)
Length = 900
Score = 145 bits (365), Expect = 4e-35
Identities = 86/224 (38%), Positives = 129/224 (57%), Gaps = 12/224 (5%)
Frame = +3
Query: 103 HQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLN-DYIARRKDFPWKLRLNIACGIARGL 161
H N++ L Y +K+EKL++Y Y GS+ LL+ + A R W R+ I G A+G+
Sbjct: 36 HPNVMPLRAYYYSKDEKLLVYNYMPGGSLFFLLHGNRGAGRTPLDWDSRVKILLGAAKGI 215
Query: 162 AFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDRGTFFSSHGYTAP 221
AFI+ E G HGN+K +N+L++ + + IS+ GL T ++GY AP
Sbjct: 216 AFIHS--EGGP--KFAHGNIKSTNVLINQELDGCISDVGLPPLMNTP-ATMSRANGYRAP 380
Query: 222 E----KSLTEKGDVYSFGVILLELLTGQSI-----EVSRIDLVRWVRSMVREEWTGEVFD 272
E K +T K DVYSFGV+LLE+LTG++ +DL RWVRS+VREEWT EVFD
Sbjct: 381 EVTDSKKITHKSDVYSFGVLLLEMLTGKTPLRYPGYEDVVDLPRWVRSVVREEWTAEVFD 560
Query: 273 KEVRENDH--QGAFSLLNIALMCVSRSQENRPNFGEILETIEGV 314
+E+ + + +L IAL CV++ + RP +++ +E +
Sbjct: 561 EELLRGQYVEEEMVQMLQIALACVAKGPDQRPRMDQVVRMLEEI 692
>TC218138 homologue to UP|Q8GRK2 (Q8GRK2) Somatic embryogenesis receptor
kinase 1, partial (60%)
Length = 1444
Score = 132 bits (331), Expect = 3e-31
Identities = 91/299 (30%), Positives = 160/299 (53%), Gaps = 27/299 (9%)
Frame = +1
Query: 42 ERFKLEDLLRATADLRSENF-----WSSLFKVKFENNVEYAVKRLKNLQVSCDE--FREI 94
+RF L +L AT ++N + ++K + + AVKRLK + E F+
Sbjct: 115 KRFSLRELQVATDSFSNKNILGRGGFGKVYKGRLADGSLVAVKRLKEERTPGGELQFQTE 294
Query: 95 LKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIA 154
++ IS H+N+L L G+ T E+L++Y Y +NGSV + L + ++ W R +A
Sbjct: 295 VEMISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERPPYQEPLDWPTRKRVA 474
Query: 155 CGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFE-------- 206
G ARGL++++ + I H ++K +NILLD++ EA++ + GL+K +
Sbjct: 475 LGSARGLSYLHDHCDP----KIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVTT 642
Query: 207 PDRGTFFSSHGYTAPEKSLT----EKGDVYSFGVILLELLTGQ-SIEVSR------IDLV 255
RGT G+ APE T EK DV+ +G++LLEL+TGQ + +++R + L+
Sbjct: 643 AVRGTI----GHIAPEYLSTGKSSEKTDVFGYGIMLLELITGQRAFDLARLANDDDVMLL 810
Query: 256 RWVRSMVREEWTGEVFDKEVRENDHQGAF-SLLNIALMCVSRSQENRPNFGEILETIEG 313
WV+ +++E+ + D +++ N + L+ +AL+C S +RP E++ +EG
Sbjct: 811 DWVKGLLKEKKLEMLVDPDLQTNYIETEVEQLIQVALLCTQGSPMDRPKMSEVVRMLEG 987
>TC216858 homologue to UP|Q76FZ8 (Q76FZ8) Brassinosteroid receptor, partial
(29%)
Length = 1320
Score = 130 bits (328), Expect = 7e-31
Identities = 91/297 (30%), Positives = 152/297 (50%), Gaps = 22/297 (7%)
Frame = +1
Query: 34 LVFFVEDHERFKLEDLLRATADLRSENFWSS-----LFKVKFENNVEYAVKRLKNLQVSC 88
L F + + DLL AT +++ S ++K + ++ A+K+L ++
Sbjct: 16 LATFEKPLRKLTFADLLDATNGFHNDSLIGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQG 195
Query: 89 D-EFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPW 147
D EF ++ I K+KH+N++ L+GY EE+L++Y+Y GS+ ++L+D W
Sbjct: 196 DREFTAEMETIGKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDQKKAGIKLNW 375
Query: 148 KLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEP 207
+R IA G ARGLAF++ + I H ++K SN+LLD+ EA +S+ G+++
Sbjct: 376 AIRRKIAIGAARGLAFLHHNC----IPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSA 543
Query: 208 -----DRGTFFSSHGYTAPEK----SLTEKGDVYSFGVILLELLTGQ----SIEVSRIDL 254
T + GY PE + KGDVYS+GV+LLELLTG+ S + +L
Sbjct: 544 MDTHLSVSTLAGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKRPTDSADFGDNNL 723
Query: 255 VRWVRSMVREEWTGEVFDKEVRENDHQGAFSL---LNIALMCVSRSQENRPNFGEIL 308
V WV+ + + ++FD E+ + D L L IA+ C+ RP +++
Sbjct: 724 VGWVKQHAKLK-ISDIFDPELMKEDPNLEMELLQHLKIAVSCLDDRPWRRPTMIQVM 891
>TC217441 homologue to GB|AAK68074.1|14573459|AF384970 somatic embryogenesis
receptor-like kinase 3 {Arabidopsis thaliana;} , partial
(54%)
Length = 1008
Score = 128 bits (321), Expect = 5e-30
Identities = 90/299 (30%), Positives = 159/299 (53%), Gaps = 27/299 (9%)
Frame = +2
Query: 42 ERFKLEDLLRATADLRSENF-----WSSLFKVKFENNVEYAVKRLKNLQVSCD--EFREI 94
+RF L +L AT + +++ + ++K + + AVKRLK + +F+
Sbjct: 113 KRFSLRELQVATDNFSNKHILGRGGFGKVYKGRLADGSLVAVKRLKEERTQGG*LQFQTE 292
Query: 95 LKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIA 154
++ IS H+N+L L G+ T E+L++Y Y +NGSV + L + + W R IA
Sbjct: 293 VEMISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERQESQPPLGWPERKRIA 472
Query: 155 CGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFE-------- 206
G ARGLA+++ + I H ++K +NILLD++ EA++ + GL+K +
Sbjct: 473 LGSARGLAYLHDHCDP----KIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVTT 640
Query: 207 PDRGTFFSSHGYTAPEKSLT----EKGDVYSFGVILLELLTGQ-SIEVSR------IDLV 255
RGT G+ APE T EK DV+ +GV+LLEL+TGQ + +++R + L+
Sbjct: 641 AVRGTI----GHIAPEYLSTGKSSEKTDVFGYGVMLLELITGQRAFDLARLANDDDVMLL 808
Query: 256 RWVRSMVREEWTGEVFDKEVREN-DHQGAFSLLNIALMCVSRSQENRPNFGEILETIEG 313
WV+ ++++ + D +++ + + + L+ +AL+C S RP E++ +EG
Sbjct: 809 DWVKGLLKDRKLETLVDADLQGSYNDEEVEQLIQVALLCTQGSPMERPKMSEVVRMLEG 985
>TC231333 similar to UP|Q8L741 (Q8L741) AT3g13690/MMM17_12, partial (38%)
Length = 1009
Score = 126 bits (317), Expect = 1e-29
Identities = 94/297 (31%), Positives = 154/297 (51%), Gaps = 27/297 (9%)
Frame = +3
Query: 44 FKLEDLLRATADLRSENF-----WSSLFKVKFENNVEYAVKRLKNLQVSCD-EFREILKQ 97
F +L AT NF + S+ + + AVK+ K D EF ++
Sbjct: 6 FTFSELQLATGGFSQANFLAEGGFGSVHRGVLPDGQVIAVKQYKLASTQGDKEFCSEVEV 185
Query: 98 ISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKD--FPWKLRLNIAC 155
+S +H+N++ L+G+ +L++Y+Y NGS L+ +I RRK W R IA
Sbjct: 186 LSCAQHRNVVMLIGFCVDDGRRLLVYEYICNGS----LDSHIYRRKQNVLEWSARQKIAV 353
Query: 156 GIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDR------ 209
G ARGL +++ EE V I H +++ +NILL EAL+ + GL+++ +PD
Sbjct: 354 GAARGLRYLH---EECRVGCIVHRDMRPNNILLTHDFEALVGDFGLARW-QPDGDMGVET 521
Query: 210 ---GTFFSSHGYTAPEKS----LTEKGDVYSFGVILLELLTG-QSIEVSRID----LVRW 257
GTF GY APE + +TEK DVYSFG++LLEL+TG ++++++R L W
Sbjct: 522 RVIGTF----GYLAPEYAQSGQITEKADVYSFGIVLLELVTGRKAVDINRPKGQQCLSEW 689
Query: 258 VRSMVREEWTGEVFDKEVRE-NDHQGAFSLLNIALMCVSRSQENRPNFGEILETIEG 313
R ++ ++ T ++ D +R Q + +L + +C+ R RP ++L +EG
Sbjct: 690 ARPLLEKQATYKLIDPSLRNCYVDQEVYRMLKCSSLCIGRDPHLRPRMSQVLRMLEG 860
>TC217385 similar to UP|Q6ZGC7 (Q6ZGC7) Receptor protein kinase PERK1-like,
partial (80%)
Length = 1604
Score = 125 bits (315), Expect = 2e-29
Identities = 89/282 (31%), Positives = 142/282 (49%), Gaps = 17/282 (6%)
Frame = +1
Query: 47 EDLLRATADLRSENFW------SSLFKVKFENNVEYAVKRLKNLQVSC-DEFREILKQIS 99
ED++R T +L SE + S+++K +N A+KR+ + C EF L+ +
Sbjct: 352 EDIMRMTENL-SEKYIIGYGASSTVYKCVLKNCKPVAIKRIYSHYPQCIKEFETELETVG 528
Query: 100 KVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIACGIAR 159
+KH+N++SL GY + L+ Y Y NGS+ +LL+ ++K W+LRL IA G A+
Sbjct: 529 SIKHRNLVSLQGYSLSPYGHLLFYDYMENGSLWDLLHG-PTKKKKLDWELRLKIALGAAQ 705
Query: 160 GLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDRG----TFFSS 215
GLA+++ I H ++K SNILLD E +++ G++K P + +
Sbjct: 706 GLAYLHHDC----CPRIIHRDVKSSNILLDADFEPHLTDFGIAKSLCPSKSHTSTYIMGT 873
Query: 216 HGYTAPE----KSLTEKGDVYSFGVILLELLTGQSIEVSRIDLVRWVRSMVREEWTGEVF 271
GY PE LTEK DVYS+G++LLELLTG+ + +L + S E
Sbjct: 874 IGYIDPEYARTSHLTEKSDVYSYGIVLLELLTGRKAVDNESNLHHLILSKAATNAVMETV 1053
Query: 272 DKEVRE--NDHQGAFSLLNIALMCVSRSQENRPNFGEILETI 311
D ++ D + +AL+C R +RP E+ +
Sbjct: 1054DPDITATCKDLGAVKKVYQLALLCTKRQPADRPTMHEVTRVL 1179
>TC219395 UP|Q9LKZ4 (Q9LKZ4) Receptor-like protein kinase 3, partial (92%)
Length = 3037
Score = 125 bits (314), Expect = 3e-29
Identities = 86/290 (29%), Positives = 146/290 (49%), Gaps = 18/290 (6%)
Frame = +3
Query: 65 LFKVKFENNVEYAVKRLKNLQVSCDE---FREILKQISKVKHQNILSLVGYRSTKEEKLI 121
++K N AVKRL + F ++ + +++H++I+ L+G+ S E L+
Sbjct: 1878 VYKGAMPNGDHVAVKRLPAMSRGSSHDHGFNAEIQTLGRIRHRHIVRLLGFCSNHETNLL 2057
Query: 122 IYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNL 181
+Y+Y NGS+ +L+ + W R IA A+GL +++ I H ++
Sbjct: 2058 VYEYMPNGSLGEVLHG--KKGGHLHWDTRYKIAVEAAKGLCYLHHDCSP----LIVHRDV 2219
Query: 182 KLSNILLDDKNEALISEHGLSKFFEPDRGT------FFSSHGYTAPEKSLT----EKGDV 231
K +NILLD +EA +++ GL+KF + D GT S+GY APE + T EK DV
Sbjct: 2220 KSNNILLDSNHEAHVADFGLAKFLQ-DSGTSECMSAIAGSYGYIAPEYAYTLKVDEKSDV 2396
Query: 232 YSFGVILLELLTGQSIE---VSRIDLVRWVRSMV--REEWTGEVFDKEVRENDHQGAFSL 286
YSFGV+LLEL+TG+ +D+V+WVR M +E +V D + +
Sbjct: 2397 YSFGVVLLELITGRKPVGEFGDGVDIVQWVRKMTDSNKEGVLKVLDPRLPSVPLHEVMHV 2576
Query: 287 LNIALMCVSRSQENRPNFGEILETIEGVMNAHDQQQMELSASKCCSNGSN 336
+A++CV RP E+++ + + D ++ L+ ++ + SN
Sbjct: 2577 FYVAMLCVEEQAVERPTMREVVQILTELPKPPDSKEGNLTITESSLSSSN 2726
>TC229775 weakly similar to UP|KPEL_DROME (Q05652) Probable
serine/threonine-protein kinase pelle , partial (8%)
Length = 1250
Score = 125 bits (314), Expect = 3e-29
Identities = 86/257 (33%), Positives = 140/257 (54%), Gaps = 14/257 (5%)
Frame = +1
Query: 70 FENNVEYAVKRLKNLQV-SCDEFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSN 128
F ++ + AVK L V ++F +K + +V H+N+ SLVGY + + +IY+Y +N
Sbjct: 133 FIDDTQVAVKMLSPSAVRGYEQFLAEVKLLMRVHHRNLTSLVGYCNEENNIGLIYEYMAN 312
Query: 129 GSVLNLLNDYIARRKDFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILL 188
G++ +++ +R K W+ RL IA A+GL + L G I H ++K +NILL
Sbjct: 313 GNLDEIVSGKSSRAKFLTWEDRLQIALDAAQGLEY----LHNGCKPPIIHRDVKCANILL 480
Query: 189 DDKNEALISEHGLSKFFEPDRGTFFS-----SHGYTAPEKS----LTEKGDVYSFGVILL 239
++ +A +++ GLSK F D G++ S + GY PE S LTEK DVYSFGV+LL
Sbjct: 481 NENFQAKLADFGLSKSFPTDGGSYMSTVVAGTPGYLDPEYSISSRLTEKSDVYSFGVVLL 660
Query: 240 ELLTGQSIEVSRID---LVRWVRSMVREEWTGEVFDKEVREN-DHQGAFSLLNIALMCVS 295
E++TGQ D + +WV+SM+ + D +E+ D + ++ I + VS
Sbjct: 661 EMVTGQPAIAKTPDKTHISQWVKSMLSNGDIKNIADSRFKEDFDTSSVWRIVEIGMASVS 840
Query: 296 RSQENRPNFGEILETIE 312
S RP+ I+ ++
Sbjct: 841 ISPFKRPSMSYIVNELK 891
>TC228048 homologue to UP|Q8LA44 (Q8LA44) Receptor protein kinase-like
protein, partial (46%)
Length = 1208
Score = 124 bits (312), Expect = 5e-29
Identities = 99/297 (33%), Positives = 153/297 (51%), Gaps = 21/297 (7%)
Frame = +2
Query: 77 AVKRLK--NLQVSCDEFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNL 134
AVKRLK N +F+ ++ IS H+N+L L G+ T E+L++Y Y SNGSV +
Sbjct: 29 AVKRLKDGNAIGGDIQFQTEVEMISLAVHRNLLKLYGFCMTPTERLLVYPYMSNGSVASR 208
Query: 135 LNDYIARRKDFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEA 194
L + W R IA G ARGL +++++ + I H ++K +NILLDD EA
Sbjct: 209 LKG----KPVLDWGTRKQIALGAARGLLYLHEQCDP----KIIHRDVKAANILLDDYCEA 364
Query: 195 LISEHGLSKFFEPD--------RGTFFSSHGYTAPEKSLT----EKGDVYSFGVILLELL 242
++ + GL+K + RGT G+ APE T EK DV+ FG++LLEL+
Sbjct: 365 VVGDFGLAKLLDHQDSHVTTAVRGTV----GHIAPEYLSTGQSSEKTDVFGFGILLLELI 532
Query: 243 TGQ-SIEVSRI-----DLVRWVRSMVREEWTGEVFDKEVREN-DHQGAFSLLNIALMCVS 295
TGQ ++E + ++ WVR + +E+ + DK+++ N D ++ +AL+C
Sbjct: 533 TGQRALEFGKAANQKGAMLDWVRKLHQEKKLELLVDKDLKTNYDRIELEEIVQVALLCTQ 712
Query: 296 RSQENRPNFGEILETIEGVMNAHDQQQMELSASKCCSNGSNQECCSLHQIIPDTWDS 352
+RP E++ +EG A ++ E S S SN QE S + T DS
Sbjct: 713 YLPGHRPKMSEVVRMLEGDGLA---EKWEASQSADTSNCKPQELSSSDRYSDLTDDS 874
>TC207146 weakly similar to UP|Q9H4D1 (Q9H4D1) Protein kinase, partial (4%)
Length = 1166
Score = 124 bits (312), Expect = 5e-29
Identities = 72/232 (31%), Positives = 128/232 (55%), Gaps = 14/232 (6%)
Frame = +1
Query: 95 LKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIA 154
++ I +V+HQ+++ L+GY + ++++Y+Y NG++ L+ + W++R+NI
Sbjct: 4 VEAIGRVRHQSLVRLLGYCAEGAHRMLVYEYVDNGNLEQWLHGDVGPCSPLTWEIRMNII 183
Query: 155 CGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDRG---- 210
G A+GL ++++ LE + H ++K SNILL K A +S+ GL+K D
Sbjct: 184 LGTAKGLTYLHEGLEP----KVVHRDIKSSNILLSKKWNAKVSDFGLAKLLGSDSSYITT 351
Query: 211 TFFSSHGYTAPEKS----LTEKGDVYSFGVILLELLTGQS-IEVSR----IDLVRWVRSM 261
+ GY APE + L E+ DVYSFG++++EL+TG++ ++ SR ++LV W++ M
Sbjct: 352 RVMGTFGYVAPEYASTGMLNERSDVYSFGILIMELITGRNPVDYSRPPEEVNLVDWLKKM 531
Query: 262 VREEWTGEVFDKEVRENDHQGAFS-LLNIALMCVSRSQENRPNFGEILETIE 312
V V D ++ E A L +AL C + + RP G ++ +E
Sbjct: 532 VSNRNPEGVLDPKLPEKPTSRALKRALLVALRCTDPNAQKRPKMGHVIHMLE 687
>TC218252 weakly similar to UP|Q03407 (Q03407) Ydr490cp, partial (5%)
Length = 1279
Score = 124 bits (311), Expect = 7e-29
Identities = 86/254 (33%), Positives = 132/254 (51%), Gaps = 18/254 (7%)
Frame = +1
Query: 77 AVKRLKNLQVSCDEFREI-LKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLL 135
A+KR+ L D F E L+ + +KH+ +++L GY ++ KL+IY Y GS+ L
Sbjct: 1 ALKRIVKLNEGFDRFFERELEILGSIKHRYLVNLRGYCNSPTSKLLIYDYLPGGSLDEAL 180
Query: 136 NDYIARRKDFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEAL 195
++ R W RLNI G A+GLA+++ I H ++K SNILLD EA
Sbjct: 181 HE---RADQLDWDSRLNIIMGAAKGLAYLHHDCSP----RIIHRDIKSSNILLDGNLEAR 339
Query: 196 ISEHGLSKFFEPDR--------GTFFSSHGYTAPEKS----LTEKGDVYSFGVILLELLT 243
+S+ GL+K E + GTF GY APE TEK DVYSFGV+ LE+L+
Sbjct: 340 VSDFGLAKLLEDEESHITTIVAGTF----GYLAPEYMQSGRATEKSDVYSFGVLTLEVLS 507
Query: 244 GQSIEVSRI-----DLVRWVRSMVREEWTGEVFDKEVRENDHQGAFSLLNIALMCVSRSQ 298
G+ + ++V W+ ++ E E+ D + +LL++A+ CVS S
Sbjct: 508 GKRPTDAAFIEKGPNIVGWLNFLITENRPREIVDPLCEGVQMESLDALLSVAIQCVSSSP 687
Query: 299 ENRPNFGEILETIE 312
E+RP +++ +E
Sbjct: 688 EDRPTMHRVVQLLE 729
>TC222007 UP|Q9LKZ5 (Q9LKZ5) Receptor-like protein kinase 2, complete
Length = 3192
Score = 123 bits (309), Expect = 1e-28
Identities = 85/290 (29%), Positives = 146/290 (50%), Gaps = 18/290 (6%)
Frame = +3
Query: 65 LFKVKFENNVEYAVKRLKNLQVSCDE---FREILKQISKVKHQNILSLVGYRSTKEEKLI 121
++K N AVKRL + F ++ + +++H++I+ L+G+ S E L+
Sbjct: 2127 VYKGAMPNGDHVAVKRLPAMSRGSSHDHGFNAEIQTLGRIRHRHIVRLLGFCSNHETNLL 2306
Query: 122 IYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNL 181
+Y+Y NGS+ +L+ + W R IA A+GL +++ I H ++
Sbjct: 2307 VYEYMPNGSLGEVLHG--KKGGHLHWDTRYKIAVEAAKGLCYLHHDCSP----LIVHRDV 2468
Query: 182 KLSNILLDDKNEALISEHGLSKFFEPDRGT------FFSSHGYTAPEKSLT----EKGDV 231
K +NILLD +EA +++ GL+KF + D GT S+GY APE + T EK DV
Sbjct: 2469 KSNNILLDSNHEAHVADFGLAKFLQ-DSGTSECMSAIAGSYGYIAPEYAYTLKVDEKSDV 2645
Query: 232 YSFGVILLELLTGQSIE---VSRIDLVRWVRSMV--REEWTGEVFDKEVRENDHQGAFSL 286
YSFGV+LLEL+TG+ +D+V+WVR M +E +V D + +
Sbjct: 2646 YSFGVVLLELITGRKPVGEFGDGVDIVQWVRKMTDSNKEGVLKVLDPRLPSVPLHEVMHV 2825
Query: 287 LNIALMCVSRSQENRPNFGEILETIEGVMNAHDQQQMELSASKCCSNGSN 336
+A++CV RP E+++ + + ++ +L+ ++ + SN
Sbjct: 2826 FYVAMLCVEEQAVERPTMREVVQILTELPKPPGSKEGDLTITESSLSSSN 2975
>TC232479 similar to UP|Q9FL51 (Q9FL51) Receptor protein kinase-like protein,
partial (17%)
Length = 984
Score = 123 bits (309), Expect = 1e-28
Identities = 78/234 (33%), Positives = 129/234 (54%), Gaps = 14/234 (5%)
Frame = +1
Query: 95 LKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIA 154
+K ++K++H+N++ ++G+ + E +IY+Y GS+ +L++ + W +RL IA
Sbjct: 1 VKTLAKIRHKNVVKILGFCHSDESVFLIYEYLHGGSLEDLIS---SPNFQLQWGIRLRIA 171
Query: 155 CGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKF-----FEPDR 209
G+A+GLA+++K V + H N+K SNILLD E +++ L + F+
Sbjct: 172 IGVAQGLAYLHKDY----VPHLLHRNVKSSNILLDANFEPKLTDFALDRVVGEAAFQSVL 339
Query: 210 GTFFSSHGYTAPE----KSLTEKGDVYSFGVILLELLTGQSIEVSR----IDLVRWVRSM 261
+ +S Y APE K TE+ DVYSFGV+LLEL++G+ E + +D+V+WVR
Sbjct: 340 NSEAASSCYIAPENGYTKKATEQLDVYSFGVVLLELVSGRQAEQTESNDSLDIVKWVRRK 519
Query: 262 VR-EEWTGEVFDKEVRENDHQGAFSLLNIALMCVSRSQENRPNFGEILETIEGV 314
V +V D ++ HQ L+IAL C S E RP+ E+L + +
Sbjct: 520 VNITNGVQQVLDPKISHTCHQEMIGALDIALHCTSVVPEKRPSMVEVLRGLHSL 681
>TC219154 similar to PIR|T05754|T05754 S-receptor kinase M4I22.110 precursor
- Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (31%)
Length = 1363
Score = 122 bits (305), Expect = 3e-28
Identities = 93/351 (26%), Positives = 181/351 (51%), Gaps = 16/351 (4%)
Frame = +1
Query: 7 KKNNLDSPMKKATSEGRLKGGDNNNSELVFFVEDHERFKLEDLLRATADLRSENFWSSLF 66
++N D K + + +L+ D +++ + F L + + E + ++
Sbjct: 238 RRNIADKSKTKKSIDRQLQDVDVPLFDMLTITAATDNFLLNNKI-------GEGGFGPVY 396
Query: 67 KVKFENNVEYAVKRLKNLQ-VSCDEFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKY 125
K K E AVKRL +L EF +K I+K++H+N++ L+G +EKL++Y+Y
Sbjct: 397 KGKLVGGQEIAVKRLSSLSGQGITEFITEVKLIAKLQHRNLVKLLGCCIKGQEKLLVYEY 576
Query: 126 QSNGSVLNLLNDYIARRKDFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSN 185
NGS+ + + D I + K W R NI GIARGL +++ ++ + I H +LK SN
Sbjct: 577 VVNGSLNSFIFDQI-KSKLLDWPRRFNIILGIARGLLYLH---QDSRLRII-HRDLKASN 741
Query: 186 ILLDDKNEALISEHGLSKFF-----EPDRGTFFSSHGYTAPE----KSLTEKGDVYSFGV 236
+LLD+K IS+ G+++ F E + ++GY APE + + K DV+SFG+
Sbjct: 742 VLLDEKLNPKISDFGMARAFGGDQTEGNTNRVVGTYGYMAPEYAFDGNFSIKSDVFSFGI 921
Query: 237 ILLELLTG---QSI--EVSRIDLVRWVRSMVREEWTGEVFDKEVREN-DHQGAFSLLNIA 290
+LLE++ G +S+ E ++LV + ++ +E+ ++ D ++++ ++++
Sbjct: 922 LLLEIVCGIKNKSLCHENQTLNLVGYAWALWKEQNALQLIDSGIKDSCVIPEVLRCIHVS 1101
Query: 291 LMCVSRSQENRPNFGEILETIEGVMNAHDQQQMELSASKCCSNGSNQECCS 341
L+CV + E+RP +++ + M+ + ++ + G+ +E S
Sbjct: 1102LLCVQQYPEDRPTMTSVIQMLGSEMDMVEPKEPGFFPRRILKEGNLKEMTS 1254
>TC209490 UP|Q8GSN9 (Q8GSN9) LRR receptor-like kinase (Fragment), complete
Length = 3298
Score = 120 bits (301), Expect = 9e-28
Identities = 91/294 (30%), Positives = 144/294 (48%), Gaps = 26/294 (8%)
Frame = +1
Query: 44 FKLEDLLRATADLRSENFWSS-----LFKVKFENNVEYAVKRLKNLQVSCDE--FREILK 96
FK ED++ L+ EN +++ N + A+KRL ++ F+ ++
Sbjct: 2149 FKAEDVVEC---LKEENIIGKGGAGIVYRGSMPNGTDVAIKRLVGAGSGRNDYGFKAEIE 2319
Query: 97 QISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIACG 156
+ K++H+NI+ L+GY S KE L++Y+Y NGS+ L+ A+ W++R IA
Sbjct: 2320 TLGKIRHRNIMRLLGYVSNKETNLLLYEYMPNGSLGEWLHG--AKGGHLKWEMRYKIAVE 2493
Query: 157 IARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKF-FEPDRGTFFS- 214
A+GL +++ I H ++K +NILLD EA +++ GL+KF ++P S
Sbjct: 2494 AAKGLCYLHHDCSP----LIIHRDVKSNNILLDGDLEAHVADFGLAKFLYDPGASQSMSS 2661
Query: 215 ---SHGYTAPEKSLT----EKGDVYSFGVILLELLTGQSIE---VSRIDLVRWVRSMVRE 264
S+GY APE + T EK DVYSFGV+LLEL+ G+ +D+V WV E
Sbjct: 2662 IAGSYGYIAPEYAYTLKVDEKSDVYSFGVVLLELIIGRKPVGEFGDGVDIVGWVNKTRLE 2841
Query: 265 -------EWTGEVFDKEVRENDHQGAFSLLNIALMCVSRSQENRPNFGEILETI 311
V D + + NIA+MCV RP E++ +
Sbjct: 2842 LAQPSDAALVLAVVDPRLSGYPLTSVIYMFNIAMMCVKEMGPARPTMREVVHML 3003
>TC226831 similar to PIR|T52285|T52285 serine/threonine-specific protein
kinase APK2a [imported] - Arabidopsis thaliana
{Arabidopsis thaliana;} , partial (82%)
Length = 1693
Score = 120 bits (300), Expect = 1e-27
Identities = 93/300 (31%), Positives = 158/300 (52%), Gaps = 20/300 (6%)
Frame = +1
Query: 77 AVKRLK--NLQVSCDEFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNL 134
AVK+LK LQ + E+ + ++ HQN++ L+GY + E +L++Y++ S GS
Sbjct: 583 AVKKLKPEGLQGHKEWLTEV-DYLGQLHHQNLVKLIGYCADGENRLLVYEFMSKGS---- 747
Query: 135 LNDYIARRKDFP--WKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKN 192
L +++ RR P W +R+ +A G ARGL+F++ + + + + K SNILLD +
Sbjct: 748 LENHLFRRGPQPLSWSVRMKVAIGAARGLSFLHNAKSQ-----VIYRDFKASNILLDAEF 912
Query: 193 EALISEHGLSKFFEPDRGTFFS-----SHGYTAPE----KSLTEKGDVYSFGVILLELLT 243
A +S+ GL+K T S + GY APE LT K DVYSFGV+LLELL+
Sbjct: 913 NAKLSDFGLAKAGPTGDRTHVSTQVMGTQGYAAPEYVATGRLTAKSDVYSFGVVLLELLS 1092
Query: 244 G-QSIEVSRI----DLVRWVRSMVREE-WTGEVFDKEV-RENDHQGAFSLLNIALMCVSR 296
G ++++ S+ +LV W + + ++ + D ++ + +GA+ +AL C++R
Sbjct: 1093GRRAVDRSKAGVEQNLVEWAKPYLGDKRRLFRIMDTKLGGQYPQKGAYMAATLALKCLNR 1272
Query: 297 SQENRPNFGEILETIEGVMNAHDQQQMELSASKCCSNGSNQECCSLHQIIPDTWDSPGSN 356
+ RP E+L+T+E +++ASK S E +H + + GS+
Sbjct: 1273EAKGRPPITEVLQTLE-----------QIAASKTAGRNSQLEQKRVHAPVRKSSVQKGSH 1419
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.135 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,257,601
Number of Sequences: 63676
Number of extensions: 195385
Number of successful extensions: 2262
Number of sequences better than 10.0: 714
Number of HSP's better than 10.0 without gapping: 1694
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1714
length of query: 357
length of database: 12,639,632
effective HSP length: 98
effective length of query: 259
effective length of database: 6,399,384
effective search space: 1657440456
effective search space used: 1657440456
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC137079.13


