

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136973.11 - phase: 2 /pseudo
(192 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
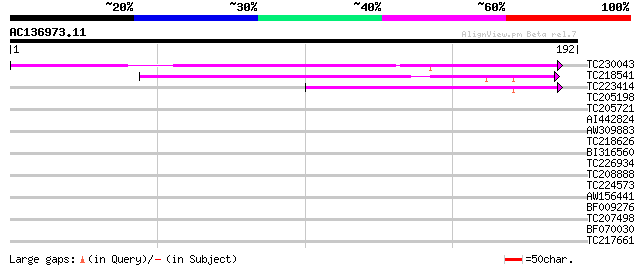
Score E
Sequences producing significant alignments: (bits) Value
TC230043 weakly similar to GB|AAL31204.1|16974602|AY060578 AT3g2... 99 9e-22
TC218541 weakly similar to UP|Q6NPD1 (Q6NPD1) At5g62960, partial... 96 1e-20
TC223414 similar to UP|Q6NPD1 (Q6NPD1) At5g62960, partial (54%) 65 3e-11
TC205198 similar to UP|Q6YBV7 (Q6YBV7) Cryptochrome 2B, partial ... 35 0.028
TC205721 similar to UP|Q9SZ53 (Q9SZ53) Protein phosphatase 2C-li... 30 0.90
AI442824 30 0.90
AW309883 similar to PIR|S33620|S3362 ADR12-2 protein - soybean, ... 29 1.2
TC218626 similar to GB|CAD40675.1|21741499|OSJN00124 OSJNBb0118P... 29 1.5
BI316560 29 1.5
TC226934 similar to UP|Q9LNV5 (Q9LNV5) F22G5.30, partial (75%) 28 2.0
TC208888 28 3.4
TC224573 homologue to UP|Q41707 (Q41707) Extensin class 1 protei... 28 3.4
AW156441 27 7.6
BF009276 27 7.6
TC207498 homologue to UP|Q8SAB7 (Q8SAB7) Adapter protein SPIKE1,... 27 7.6
BF070030 similar to GP|8954295|gb| photosystem I reaction center... 26 9.9
TC217661 similar to UP|Q9FJ10 (Q9FJ10) Gb|AAF63824.1, partial (47%) 26 9.9
>TC230043 weakly similar to GB|AAL31204.1|16974602|AY060578
AT3g27760/MGF10_16 {Arabidopsis thaliana;} , partial
(48%)
Length = 1408
Score = 99.4 bits (246), Expect = 9e-22
Identities = 62/197 (31%), Positives = 92/197 (46%), Gaps = 10/197 (5%)
Frame = +3
Query: 1 WYDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVST 60
W ++C V +SI+ + V+W ++GS +S E G +S
Sbjct: 132 WRFYLCAVCVLLSIVLSFLVIWKDKGSRKFRSGKGENQED---------------GTLSG 266
Query: 61 SQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFAL 120
+ W + +HP+ LL R+ +F SL L IH +IF+YYT+WTFTLV IYF
Sbjct: 267 DEAWKPFLKEIHPVCLLAFRVIAFSSLLASLVAKIHINGRAIFFYYTQWTFTLVTIYFGF 446
Query: 121 GTTVSAYGCWKVLNKPPPLQN----------GEMTEFLRRDLETKGSIFTFQSRYAEEEF 170
+T+SAYGC++ NK + N G FL +D + AE +
Sbjct: 447 ASTLSAYGCYR-HNKSSTINNVNVARIDVEQGPYMPFLHQDTSNLSRMEHLADPLAEIQK 623
Query: 171 EQTAGFWGYLMQITFQV 187
Q A W Y++QI FQ+
Sbjct: 624 NQVAPIWSYILQILFQM 674
>TC218541 weakly similar to UP|Q6NPD1 (Q6NPD1) At5g62960, partial (46%)
Length = 1107
Score = 95.9 bits (237), Expect = 1e-20
Identities = 57/157 (36%), Positives = 78/157 (49%), Gaps = 15/157 (9%)
Frame = +3
Query: 45 NSPSSDNRVAIGHVSTSQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFY 104
N R G + + W +C +G+HP LL R+ SF+ L LL ++ IFY
Sbjct: 12 NERRERQRETAGSLYEYEAWNTCLKGIHPAWLLAYRIISFLVLFSLLTANVVADGGGIFY 191
Query: 105 YYTEWTFTLVMIYFALGTTVSAYGCWKVLNKPPPLQNGEMTEFLRRDLETKGSIFT---- 160
+YT+WTFTLV IYF LG+ VS YGC NK + T R DL+T+ +
Sbjct: 192 FYTQWTFTLVTIYFGLGSCVSIYGCRYKHNKI------DCTTVNRADLDTEEGTYVAPTL 353
Query: 161 ---------FQSRYAEEE--FEQTAGFWGYLMQITFQ 186
+++ A +E TAG WGY+ QITFQ
Sbjct: 354 DGTPELPNLYKNSNANQEPYTRNTAGVWGYIFQITFQ 464
>TC223414 similar to UP|Q6NPD1 (Q6NPD1) At5g62960, partial (54%)
Length = 819
Score = 64.7 bits (156), Expect = 3e-11
Identities = 34/96 (35%), Positives = 50/96 (51%), Gaps = 9/96 (9%)
Frame = +3
Query: 101 SIFYYYTEWTFTLVMIYFALGTTVSAYGCWKVLNKPPPLQNGEMTEFLRRDLETKGSIFT 160
SIFYYYT+WTFT + IYF LG+ +S YGC++ NK + G + + + ++
Sbjct: 9 SIFYYYTQWTFTSITIYFGLGSLLSIYGCYQHHNKATGDKVGNVDGDAEQGMYDASALPQ 188
Query: 161 FQSRYAEEE---------FEQTAGFWGYLMQITFQV 187
+ + E+ Q AG WGY QI FQ+
Sbjct: 189 SSNPFDPEKSLGDPEEVLVRQHAGIWGYTFQIIFQI 296
>TC205198 similar to UP|Q6YBV7 (Q6YBV7) Cryptochrome 2B, partial (58%)
Length = 1040
Score = 34.7 bits (78), Expect = 0.028
Identities = 15/47 (31%), Positives = 26/47 (54%), Gaps = 8/47 (17%)
Frame = +2
Query: 20 VLWTNEGSSTSQSDSNIFVESL--------LVANSPSSDNRVAIGHV 58
+LWTNEG+S + +N+F+ ++ L N P + R +GH+
Sbjct: 437 ILWTNEGNSAGEESANLFLRAIGLREYSRYLCFNFPFTHERALLGHL 577
>TC205721 similar to UP|Q9SZ53 (Q9SZ53) Protein phosphatase 2C-like protein
(AT4g31860/F11C18_60), partial (93%)
Length = 1641
Score = 29.6 bits (65), Expect = 0.90
Identities = 25/63 (39%), Positives = 34/63 (53%), Gaps = 2/63 (3%)
Frame = -3
Query: 56 GHVSTSQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDA--SIFYYYTEWTFTL 113
GHV TS L TSCW LT R +SL +L+Y ++H+ A +I Y T + L
Sbjct: 1072 GHVITS-LTTSCWCQAPIKYFLTNRRKFILSL*LLMY-EVHKLLARHTIPYSITSYYKKL 899
Query: 114 VMI 116
V+I
Sbjct: 898 VII 890
>AI442824
Length = 437
Score = 29.6 bits (65), Expect = 0.90
Identities = 15/49 (30%), Positives = 24/49 (48%)
Frame = -2
Query: 76 LLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFALGTTV 124
LLT L + +SL + S +YYY+ W F +++I+ G V
Sbjct: 157 LLTDLLINLLSL-----ISFFRVSCSFYYYYSFWFFWVILIWVCYGLLV 26
>AW309883 similar to PIR|S33620|S3362 ADR12-2 protein - soybean, partial
(41%)
Length = 437
Score = 29.3 bits (64), Expect = 1.2
Identities = 13/40 (32%), Positives = 23/40 (57%)
Frame = +1
Query: 81 LFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFAL 120
+F +S +++YL H Y I+Y+ +F V+IY+ L
Sbjct: 313 IFLSISFVLIVYLYFHSYSIIIYYF----SFISVLIYYYL 420
>TC218626 similar to GB|CAD40675.1|21741499|OSJN00124 OSJNBb0118P14.13 {Oryza
sativa (japonica cultivar-group);} , partial (44%)
Length = 1041
Score = 28.9 bits (63), Expect = 1.5
Identities = 13/50 (26%), Positives = 25/50 (50%), Gaps = 1/50 (2%)
Frame = +3
Query: 55 IGHVSTSQLWTSCWRGVHPL-VLLTTRLFSFVSLAMLLYLDIHEYDASIF 103
+G+ S + LW W + + VL + LFS + + + +H+Y+ F
Sbjct: 225 LGYFSFTILWIRVWNQI**INVLFCSHLFSLAGIPKFILISVHDYNLPSF 374
>BI316560
Length = 400
Score = 28.9 bits (63), Expect = 1.5
Identities = 20/47 (42%), Positives = 26/47 (54%), Gaps = 6/47 (12%)
Frame = -1
Query: 69 RGVHPLVLLTTRLFS----FVSLAMLLYLDIHEYDASIFY--YYTEW 109
RGV PL +L TR S + +LL+L +EYDAS F + EW
Sbjct: 169 RGVEPLWVLQTRACSRHQTHSNSQILLHLYQYEYDASFFLRCSFREW 29
>TC226934 similar to UP|Q9LNV5 (Q9LNV5) F22G5.30, partial (75%)
Length = 1613
Score = 28.5 bits (62), Expect = 2.0
Identities = 15/55 (27%), Positives = 26/55 (47%)
Frame = +1
Query: 13 SILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVSTSQLWTSC 67
S+L ++W+ T+ ST+ + S+ + SS + V T +LWT C
Sbjct: 1402 SVLSSVWLYATSSPLSTAPTSIQCCGGSIPATSCKSSLSAFYAARVFTDRLWTGC 1566
>TC208888
Length = 548
Score = 27.7 bits (60), Expect = 3.4
Identities = 13/35 (37%), Positives = 20/35 (57%), Gaps = 5/35 (14%)
Frame = +3
Query: 91 LYLDIHEYDA-----SIFYYYTEWTFTLVMIYFAL 120
+Y +I YDA S F+++T W L+ +YF L
Sbjct: 357 IYEEIIVYDAFGFL*SFFFFFTFWYILLLSLYFGL 461
>TC224573 homologue to UP|Q41707 (Q41707) Extensin class 1 protein precursor,
partial (44%)
Length = 1112
Score = 27.7 bits (60), Expect = 3.4
Identities = 22/81 (27%), Positives = 44/81 (54%), Gaps = 2/81 (2%)
Frame = +2
Query: 25 EGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVSTSQLWTSCWRGVHPLVLLTTRLFSF 84
E S+T+Q S +++ +L + S+ + +I H +S W V PL+ T+ L
Sbjct: 605 ESSTTNQISSTTYLQVILELSRSSNKSCSSI-HCGIKS--SSIWSKVKPLICCTSSLNKS 775
Query: 85 VSLAMLLYLDIHEYD--ASIF 103
++++ L Y ++H+ + ASI+
Sbjct: 776 LNISSLSY-NLHDSNEVASIY 835
>AW156441
Length = 411
Score = 26.6 bits (57), Expect = 7.6
Identities = 16/47 (34%), Positives = 24/47 (51%)
Frame = -3
Query: 79 TRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFALGTTVS 125
TRLF VSL + L IH Y + F++ + IY +L ++S
Sbjct: 358 TRLFFLVSLPRPISLYIHTYTYTYFFF--------ISIYISLSLSLS 242
>BF009276
Length = 427
Score = 26.6 bits (57), Expect = 7.6
Identities = 10/25 (40%), Positives = 17/25 (68%)
Frame = +2
Query: 96 HEYDASIFYYYTEWTFTLVMIYFAL 120
H+ D + FYY+ + TLV+I+F +
Sbjct: 176 HQEDQNKFYYHFLYKATLVLIFFVI 250
>TC207498 homologue to UP|Q8SAB7 (Q8SAB7) Adapter protein SPIKE1, partial
(19%)
Length = 1451
Score = 26.6 bits (57), Expect = 7.6
Identities = 26/84 (30%), Positives = 40/84 (46%), Gaps = 5/84 (5%)
Frame = -3
Query: 4 FVCFAIVGVSILGALWVLWT-----NEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHV 58
F C A + IL LW L + GSS+S + + V + +A S ++R + +
Sbjct: 792 FTCTATLPCRILCRLWSLGS*SPSELRGSSSSFLKAAVRVSIIPIAFSTGENSRDS--DL 619
Query: 59 STSQLWTSCWRGVHPLVLLTTRLF 82
T+ L+T+ G P V T RLF
Sbjct: 618 ITNNLFTNA--GKEPSVRRTVRLF 553
>BF070030 similar to GP|8954295|gb| photosystem I reaction center subunit III
{Vigna radiata}, partial (44%)
Length = 463
Score = 26.2 bits (56), Expect = 9.9
Identities = 12/26 (46%), Positives = 14/26 (53%)
Frame = -2
Query: 109 WTFTLVMIYFALGTTVSAYGCWKVLN 134
WTF LV FAL + A WK L+
Sbjct: 162 WTFFLVFSTFALMASAGALPAWKSLS 85
>TC217661 similar to UP|Q9FJ10 (Q9FJ10) Gb|AAF63824.1, partial (47%)
Length = 765
Score = 26.2 bits (56), Expect = 9.9
Identities = 16/48 (33%), Positives = 19/48 (39%)
Frame = +2
Query: 23 TNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVSTSQLWTSCWRG 70
T +GS DSN + NSPSS+N T WRG
Sbjct: 188 TGKGSEIEVEDSNSNPNAPSSGNSPSSNNEQKKTCADCGTTKTPLWRG 331
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.137 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,183,491
Number of Sequences: 63676
Number of extensions: 136740
Number of successful extensions: 1190
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 1185
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1187
length of query: 192
length of database: 12,639,632
effective HSP length: 92
effective length of query: 100
effective length of database: 6,781,440
effective search space: 678144000
effective search space used: 678144000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC136973.11


