

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
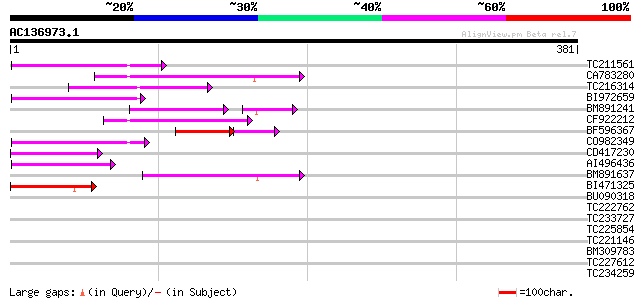
Query= AC136973.1 - phase: 0 /pseudo
(381 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC211561 weakly similar to UP|O22675 (O22675) Reverse transcript... 67 1e-11
CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polym... 63 3e-10
TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {A... 63 3e-10
BI972659 62 4e-10
BM891241 52 1e-08
CF922212 57 2e-08
BF596367 46 2e-07
CO982349 53 3e-07
CD417230 weakly similar to GP|2244915|emb| reverse transcriptase... 47 1e-05
AI496436 45 4e-05
BM891637 43 2e-04
BI471325 42 6e-04
BU090318 weakly similar to GP|9743345|gb| F21J9.30 {Arabidopsis ... 40 0.001
TC222762 39 0.003
TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR ret... 34 0.096
TC225854 similar to UP|Q6T284 (Q6T284) Predicted protein, partia... 32 0.62
TC221146 30 1.4
BM309783 30 1.8
TC227612 weakly similar to UP|Q8L7S0 (Q8L7S0) At1g09730/F21M12_1... 29 4.0
TC234259 similar to UP|Q8L7S0 (Q8L7S0) At1g09730/F21M12_12, part... 29 4.0
>TC211561 weakly similar to UP|O22675 (O22675) Reverse transcriptase
(Fragment) , partial (59%)
Length = 1077
Score = 67.0 bits (162), Expect = 1e-11
Identities = 35/105 (33%), Positives = 60/105 (56%), Gaps = 1/105 (0%)
Frame = +1
Query: 2 YLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIFR 61
YLG+P + + I + +K+S W + +S GR +IK VL +IP Y S FR
Sbjct: 598 YLGIPIGANSRRCHMWDSIIRKCERKLSKWK*RHISFGGRVTLIKSVLTSIPIYFFSFFR 777
Query: 62 LSNTLLDE-IEMMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGM 105
+ +++D+ +++ +F WG GG + ++W+ WEKL + K GG+
Sbjct: 778 VPQSVVDKLVKI*RTFLWG-GGPDQKKISWIRWEKLCLPKERGGI 909
>CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polymerase
homolog T24M8.8 - Arabidopsis thaliana, partial (4%)
Length = 456
Score = 62.8 bits (151), Expect = 3e-10
Identities = 37/147 (25%), Positives = 68/147 (46%), Gaps = 6/147 (4%)
Frame = +2
Query: 58 SIFRLSNTLLDEIE-MMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVA 116
S F L ++ ++E + F WG G + +R + W++W+ + + K GG+G KD FN
Sbjct: 5 SFFSLP*KIIAKLESLQRRFLWG-GEADSRKIAWVNWKTVCLPKAKGGLGIKDLRTFNTT 181
Query: 117 MLGKQGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSI-----LSAKVVVR 171
+LGK W L +K+ +++Y + G S W+ + L + ++
Sbjct: 182 LLGKWRWDLFYIQQEPWAKVLQSKYGGWRALEEGSSGSKDSAWWKDLIKTQQLQRNIPLK 361
Query: 172 QGARWKIGAGFDIPIISEPWIGSGSSI 198
+ WK+G G I + W + S+
Sbjct: 362 RETIWKVGGGDRIKFWEDLWTNTDLSL 442
>TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Arabidopsis
thaliana;} , partial (18%)
Length = 759
Score = 62.8 bits (151), Expect = 3e-10
Identities = 32/98 (32%), Positives = 53/98 (53%), Gaps = 1/98 (1%)
Frame = +2
Query: 40 GREVMIKYVLQAIPSYVMSIFRLSNTLLDE-IEMMNSFWWGHGGSSNRGLNWLSWEKLSV 98
GR +IK VL A+P +S F++ ++D+ + + F WG NR + W+ W +
Sbjct: 8 GRVTLIKSVLSALPIXXLSFFKIPQRIVDKLVTLQRQFLWGGTQHHNR-IPWVKWADICN 184
Query: 99 HKNDGGMGFKDFVAFNVAMLGKQGWQLQTKPDSLVSKI 136
K DGG+G KD FN A+ G+ W L + + L +++
Sbjct: 185 PKIDGGLGIKDLSNFNAALRGRWIWGLASNHNQLWARL 298
>BI972659
Length = 453
Score = 62.0 bits (149), Expect = 4e-10
Identities = 30/91 (32%), Positives = 53/91 (57%), Gaps = 1/91 (1%)
Frame = +1
Query: 2 YLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIFR 61
YLG+P V S + + + ++ K+S W+ K LS GR +IK VL A+P Y++S F+
Sbjct: 184 YLGMPIAVKASSRMVWEPLINKFKAKLSKWNQKYLSMGGRVTLIKSVLNALPIYLLSFFK 363
Query: 62 LSNTLLDE-IEMMNSFWWGHGGSSNRGLNWL 91
+ ++D+ + + +F WG NR ++W+
Sbjct: 364 IPQRIVDKLVSLQRTFMWGGNQHHNR-ISWV 453
>BM891241
Length = 407
Score = 52.4 bits (124), Expect(2) = 1e-08
Identities = 23/67 (34%), Positives = 38/67 (56%)
Frame = -2
Query: 81 GGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLGKQGWQLQTKPDSLVSKIFKAR 140
G ++ + W+ WE + + K GG+G KD FN+A+LGK W L + L ++I ++
Sbjct: 403 GDHDHKKIPWVKWEVVCLPKTKGGLGIKDISKFNIALLGKWIWALASDQQQLWARIINSK 224
Query: 141 YYPNSNF 147
Y S+F
Sbjct: 223 YGGWSDF 203
Score = 24.6 bits (52), Expect(2) = 1e-08
Identities = 12/42 (28%), Positives = 19/42 (44%), Gaps = 5/42 (11%)
Frame = -3
Query: 157 SFVWRSIL-----SAKVVVRQGARWKIGAGFDIPIISEPWIG 193
S+ WR + S + Q WK+G G I ++ W+G
Sbjct: 174 SYWWRDLRKFYHQSDHSIFHQYMSWKVGCGDKINFWTDKWLG 49
>CF922212
Length = 445
Score = 56.6 bits (135), Expect = 2e-08
Identities = 29/101 (28%), Positives = 53/101 (51%), Gaps = 1/101 (0%)
Frame = -2
Query: 64 NTLLDE-IEMMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLGKQG 122
N +LD+ + + F WG GG + + W++W+ + + K G+G +D FN A+LGK+
Sbjct: 435 NKVLDKLVRLQQWFLWG-GGLGQKKIAWVNWDSVCLPKEKEGLGNRDLRKFNYALLGKRR 259
Query: 123 WQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSI 163
W L L +++ ++Y N + + + SF W+ I
Sbjct: 258 WNLFHHQGELGARVLDSKYKRWRNLDEERRVKSESFWWQEI 136
>BF596367
Length = 419
Score = 45.8 bits (107), Expect(2) = 2e-07
Identities = 21/40 (52%), Positives = 29/40 (72%)
Frame = +1
Query: 112 AFNVAMLGKQGWQLQTKPDSLVSKIFKARYYPNSNFLDAK 151
AF+ AML KQ W+L D+LV+ + A+Y+PNS+ LDAK
Sbjct: 70 AFHKAMLAKQMWRLIDNKDALVT*VLTAKYFPNSSLLDAK 189
Score = 27.3 bits (59), Expect(2) = 2e-07
Identities = 11/31 (35%), Positives = 18/31 (57%)
Frame = +3
Query: 151 KLGHNPSFVWRSILSAKVVVRQGARWKIGAG 181
K + P++ W S+ +K + GA W+IG G
Sbjct: 189 KKAYMPNYSWSSV**SKDLAEHGAIWRIGDG 281
>CO982349
Length = 795
Score = 52.8 bits (125), Expect = 3e-07
Identities = 28/94 (29%), Positives = 46/94 (48%), Gaps = 1/94 (1%)
Frame = +3
Query: 2 YLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIFR 61
YLG+P + I + K++ W + +S GR +I +L A+P Y S FR
Sbjct: 501 YLGIPIEANPRCREI*DPIIRKCETKLARWKQRHISLGGRVTLINAILTALPIYFFSFFR 680
Query: 62 LSNTLLDE-IEMMNSFWWGHGGSSNRGLNWLSWE 94
+ ++ + + + F WG GGS R + W+ WE
Sbjct: 681 APSKVVQKLVNIQRKFLWG-GGSXQRRIAWVKWE 779
>CD417230 weakly similar to GP|2244915|emb| reverse transcriptase like
protein {Arabidopsis thaliana}, partial (7%)
Length = 688
Score = 47.4 bits (111), Expect = 1e-05
Identities = 25/62 (40%), Positives = 34/62 (54%)
Frame = -2
Query: 1 KYLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIF 60
KYLG+P TF FI D+V ++ S W S LS GR + K V+ A+PS++M
Sbjct: 246 KYLGVPIFHKNPTSHTFDFIIDKVNKRFSRWKSNLLSMVGRLTLTK*VV*ALPSHIMQFD 67
Query: 61 RL 62
L
Sbjct: 66 NL 61
>AI496436
Length = 414
Score = 45.4 bits (106), Expect = 4e-05
Identities = 21/70 (30%), Positives = 39/70 (55%)
Frame = +1
Query: 2 YLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIFR 61
YLG+P + + + +K++SW K +S GR +I VL A+P Y++S FR
Sbjct: 145 YLGIPIGANSRHSDVWEPVVRKCERKLASWKQKYVSFGGRVTLINSVLTALPIYLLSFFR 324
Query: 62 LSNTLLDEIE 71
+ + ++D+ +
Sbjct: 325 IPDKVVDKAD 354
>BM891637
Length = 427
Score = 43.1 bits (100), Expect = 2e-04
Identities = 26/114 (22%), Positives = 47/114 (40%), Gaps = 5/114 (4%)
Frame = -1
Query: 90 WLSWEKLSVHKNDGGMGFKDFVAFNVAMLGKQGWQLQTKPDSLVSKIFKARYYPNSNFLD 149
W+SW + + GG+G KD N A+L K W + +P L ++I ++Y
Sbjct: 406 WISWRQCCASGDVGGLGIKDIKILNNALLIKWKWLMFHQPHQLWNRILISKYKGWRGLDQ 227
Query: 150 AKLGHNPSFVWRSILS-----AKVVVRQGARWKIGAGFDIPIISEPWIGSGSSI 198
+ S W + + + + WK+G G I + W+ G+ +
Sbjct: 226 GPQKYYFSPWWADLRAINQHQSMIAASNQFCWKVGRGDQILFWEDSWVDDGTPL 65
>BI471325
Length = 421
Score = 41.6 bits (96), Expect = 6e-04
Identities = 23/60 (38%), Positives = 37/60 (61%), Gaps = 2/60 (3%)
Frame = +1
Query: 1 KYLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGRE--VMIKYVLQAIPSYVMS 58
+YL +PSM+ KKA F++++D + S+ + K RE ++IKYV Q+I +Y MS
Sbjct: 241 EYLDMPSMIRTKKKAIFNYLRDEFGRIYSTLVQQESFKGQREKCLIIKYVAQSITTYCMS 420
>BU090318 weakly similar to GP|9743345|gb| F21J9.30 {Arabidopsis thaliana},
partial (1%)
Length = 422
Score = 40.4 bits (93), Expect = 0.001
Identities = 27/110 (24%), Positives = 49/110 (44%), Gaps = 5/110 (4%)
Frame = +3
Query: 88 LNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLGKQGWQLQTKPDSLVSKIFKARYYPNSNF 147
++W SW+ L K GG+G + N ++L K + TKP+ L ++ +++Y +
Sbjct: 6 VHWTSWKALCSPKKSGGLGLRLLSHMNKSLLMKST*NMCTKPNDL*VRVVRSKYK*GKDI 185
Query: 148 L----DAKLGHNPSFVWRSILSA-KVVVRQGARWKIGAGFDIPIISEPWI 192
L K G N W+ I+ + + KIG G + + W+
Sbjct: 186 LP*IDPKKTGTN---FWQGIVHCWEKITHDNIARKIGNGRSVHFWKDVWV 326
>TC222762
Length = 879
Score = 39.3 bits (90), Expect = 0.003
Identities = 15/34 (44%), Positives = 25/34 (73%)
Frame = +3
Query: 140 RYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQG 173
++Y +FLD+ LG+ PS VWRSI S++ +++ G
Sbjct: 312 KHYSLGDFLDSNLGYGPSLVWRSIWSSRDLIKDG 413
>TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR retroelement
reverse transcriptase, partial (7%)
Length = 956
Score = 34.3 bits (77), Expect = 0.096
Identities = 15/59 (25%), Positives = 29/59 (48%)
Frame = -2
Query: 2 YLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIF 60
Y G+P G+ + + D++ K++SW LS GR ++ V+ + Y S++
Sbjct: 235 YFGVPIFKGKPTRRHLYCVADKIKCKLASWKGSILSIMGRIQLVNSVIHGMLLYSFSVY 59
>TC225854 similar to UP|Q6T284 (Q6T284) Predicted protein, partial (10%)
Length = 1101
Score = 31.6 bits (70), Expect = 0.62
Identities = 20/60 (33%), Positives = 29/60 (48%), Gaps = 5/60 (8%)
Frame = +1
Query: 193 GSGSSIPPVGDDMLALQPYSVGS-----N*PRAQSLE*TIGPSTFCSRNGPKHPEYSSAP 247
GSG+ IPP ++ + P+S S N P+ + P CSRN P+H + S P
Sbjct: 562 GSGAGIPPSFVSIIIVVPFSFFSYL**LNEPQVKITH-KESPELMCSRNTPRHCNFISYP 738
>TC221146
Length = 547
Score = 30.4 bits (67), Expect = 1.4
Identities = 14/28 (50%), Positives = 19/28 (67%), Gaps = 2/28 (7%)
Frame = -2
Query: 152 LGHNPSFVWRSILSAKVVVR--QGARWK 177
LGHNPS+ WRS+L ++R Q RW+
Sbjct: 273 LGHNPSYPWRSLLLMLSLMRSLQLLRWR 190
>BM309783
Length = 433
Score = 30.0 bits (66), Expect = 1.8
Identities = 15/54 (27%), Positives = 25/54 (45%)
Frame = +2
Query: 91 LSWEKLSVHKNDGGMGFKDFVAFNVAMLGKQGWQLQTKPDSLVSKIFKARYYPN 144
+ W+ + + + GG+G D N A L K GW + SL + + + Y N
Sbjct: 41 VGWKAMIMP*HSGGVGMCDLTIMNQACLMKIGWAFRMGDGSLWADVLRG*YGQN 202
>TC227612 weakly similar to UP|Q8L7S0 (Q8L7S0) At1g09730/F21M12_12, partial
(19%)
Length = 1563
Score = 28.9 bits (63), Expect = 4.0
Identities = 13/28 (46%), Positives = 16/28 (56%)
Frame = +3
Query: 262 WPLFCEKCISYMHRRSHQQ*PFAQTRVL 289
WPLF C ++ R S+Q *PF VL
Sbjct: 270 WPLFAPLCGTFSGRSSNQL*PFHDNEVL 353
>TC234259 similar to UP|Q8L7S0 (Q8L7S0) At1g09730/F21M12_12, partial (8%)
Length = 468
Score = 28.9 bits (63), Expect = 4.0
Identities = 13/28 (46%), Positives = 16/28 (56%)
Frame = +1
Query: 262 WPLFCEKCISYMHRRSHQQ*PFAQTRVL 289
WPLF C + R S+Q *PF +VL
Sbjct: 88 WPLFAPLCGRFSGRSSNQL*PFHDNKVL 171
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.339 0.148 0.518
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,211,318
Number of Sequences: 63676
Number of extensions: 309290
Number of successful extensions: 2215
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 2198
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2212
length of query: 381
length of database: 12,639,632
effective HSP length: 99
effective length of query: 282
effective length of database: 6,335,708
effective search space: 1786669656
effective search space used: 1786669656
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC136973.1


