

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
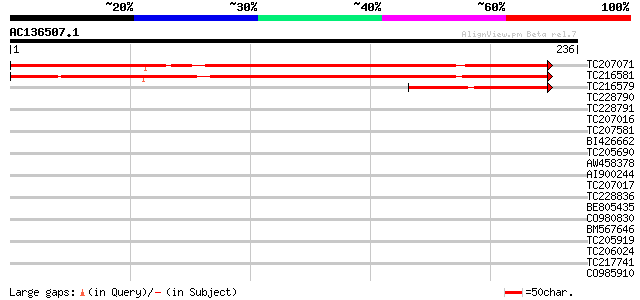
Query= AC136507.1 - phase: 0
(236 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC207071 similar to UP|Q6NLB1 (Q6NLB1) At2g26850, partial (66%) 332 9e-92
TC216581 295 1e-80
TC216579 similar to UP|Q6NLB1 (Q6NLB1) At2g26850, partial (50%) 86 1e-17
TC228790 37 0.008
TC228791 34 0.066
TC207016 33 0.15
TC207581 32 0.33
BI426662 32 0.33
TC205690 weakly similar to UP|Q6K3E9 (Q6K3E9) F-box family prote... 31 0.43
AW458378 31 0.43
AI900244 similar to GP|25082966|gb| spliceosome associated prote... 31 0.56
TC207017 30 0.96
TC228836 similar to UP|Q6SH18 (Q6SH18) Lipase/esterase, Lip3/Bch... 30 0.96
BE805435 weakly similar to GP|13491750|gb| cysteine protease {Ip... 30 1.2
CO980830 30 1.2
BM567646 28 2.8
TC205919 similar to UP|Q94F39 (Q94F39) AT3g29575/MWE13_2, partia... 28 2.8
TC206024 weakly similar to UP|Q9FLU7 (Q9FLU7) Phloem-specific le... 28 2.8
TC217741 weakly similar to UP|Q84TE4 (Q84TE4) At4g12510, partial... 28 3.6
CO985910 28 3.6
>TC207071 similar to UP|Q6NLB1 (Q6NLB1) At2g26850, partial (66%)
Length = 1609
Score = 332 bits (851), Expect = 9e-92
Identities = 168/234 (71%), Positives = 187/234 (79%), Gaps = 8/234 (3%)
Frame = +2
Query: 1 MLLYFITCVSFFLFLKPLLLLKPLPSWTNEVRLISLLFHKDFCLKSFCKLFRKTL----- 55
MLLY ITC+SF LFLKPLL LKPLPSW +E+RLIS LF+KD +KS KLFR L
Sbjct: 83 MLLYLITCISFLLFLKPLLSLKPLPSWASELRLISPLFYKDLSVKSISKLFRNRLWLGFL 262
Query: 56 ---VSRMSLVKKTLVSKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGV 112
+RMSLVKK+ SKVE +E+ Q++SVLDLPELVLECILEKLPP SLCQMAGV
Sbjct: 263 CQIPTRMSLVKKS--SKVEIIEDS-----QDMSVLDLPELVLECILEKLPPPSLCQMAGV 421
Query: 113 CHSLRERCVSDHLWERHMKQKWGRVIGSVAYREWKWHVATKKNIGSLRHGKPRSLIKFMS 172
C SLRE CVSDHLWERHMKQKWGRVIG AYREWKWH A+K N+G+LRHGK RSL++ +S
Sbjct: 422 CRSLRESCVSDHLWERHMKQKWGRVIGQAAYREWKWHAASKGNVGTLRHGKQRSLLRLVS 601
Query: 173 LYWPFSWMRMKVDAAYSNSANKQNSYLPPDSFMTWYLALETGNFSFPAQVYNRE 226
L WPFSWMR KVDA N++ KQ SYLPPDS MTWYLALETGNF FPAQVYNRE
Sbjct: 602 LSWPFSWMRTKVDA---NNSIKQQSYLPPDSLMTWYLALETGNFWFPAQVYNRE 754
>TC216581
Length = 773
Score = 295 bits (755), Expect = 1e-80
Identities = 147/234 (62%), Positives = 179/234 (75%), Gaps = 8/234 (3%)
Frame = +2
Query: 1 MLLYFITCVSFFLFLKPLLLLKPLPSWTNEVRLISLLFHKDFCLKSFCKLFRKT------ 54
MLLYFITC+SFFLFLK LLL KPLPSW +E+ L LLF ++ +++ KLFR
Sbjct: 95 MLLYFITCISFFLFLKSLLL-KPLPSWASEMWLFPLLFSQELSVRTISKLFRNCFWIGFL 271
Query: 55 --LVSRMSLVKKTLVSKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGV 112
+ MSLV+K+L S+VENVE+ D +SVLDLPEL L+CILE+LPPS+LC+MA V
Sbjct: 272 CQIPIGMSLVRKSLTSRVENVEDSHD-----MSVLDLPELALDCILERLPPSALCRMAAV 436
Query: 113 CHSLRERCVSDHLWERHMKQKWGRVIGSVAYREWKWHVATKKNIGSLRHGKPRSLIKFMS 172
C SLRERCVSDHLWERHM+QKWGRVIG AYREWKWHVA+K+N+G L+HG+ + L++FMS
Sbjct: 437 CRSLRERCVSDHLWERHMRQKWGRVIGPAAYREWKWHVASKRNVGGLKHGRQKGLMRFMS 616
Query: 173 LYWPFSWMRMKVDAAYSNSANKQNSYLPPDSFMTWYLALETGNFSFPAQVYNRE 226
L WP W+R K DA +N++ KQ S LP DS M WYLALE+GNF F AQVYNRE
Sbjct: 617 LRWPLQWIRPKADA--NNNSPKQGSSLPVDSGMNWYLALESGNFWFLAQVYNRE 772
>TC216579 similar to UP|Q6NLB1 (Q6NLB1) At2g26850, partial (50%)
Length = 1079
Score = 86.3 bits (212), Expect = 1e-17
Identities = 41/60 (68%), Positives = 47/60 (78%)
Frame = +3
Query: 167 LIKFMSLYWPFSWMRMKVDAAYSNSANKQNSYLPPDSFMTWYLALETGNFSFPAQVYNRE 226
L++FMSL+WPF W+R KVDA Y+N K S LP DS M WYLALE+GNF FPAQVYNRE
Sbjct: 3 LMRFMSLHWPFQWIRPKVDANYNNP--KLRSSLPVDSVMNWYLALESGNFWFPAQVYNRE 176
>TC228790
Length = 1215
Score = 37.0 bits (84), Expect = 0.008
Identities = 19/72 (26%), Positives = 35/72 (48%)
Frame = +2
Query: 88 DLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVIGSVAYREWK 147
D+PE + C+ L P +C +A + + R SD +WE + + + ++ V E
Sbjct: 182 DIPESCVACVFLHLTPPEICNLARLNRAFRGAASSDSVWEAKLPRNYQDLLDLVPPPERH 361
Query: 148 WHVATKKNIGSL 159
+KK+I +L
Sbjct: 362 HRSLSKKDIFAL 397
>TC228791
Length = 584
Score = 33.9 bits (76), Expect = 0.066
Identities = 13/47 (27%), Positives = 24/47 (50%)
Frame = +1
Query: 88 DLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKW 134
D+PE + C+ L P +C +A + + R SD +WE + + +
Sbjct: 265 DIPESCVACVFLHLTPPEICNLARLNRAFRGAASSDSVWEAKLPRNY 405
>TC207016
Length = 1310
Score = 32.7 bits (73), Expect = 0.15
Identities = 16/71 (22%), Positives = 32/71 (44%)
Frame = +1
Query: 71 ENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHM 130
EN + SL S+ D+PE + ++ P +C +A V + ++ +WE +
Sbjct: 262 ENDDGGTFSLSSETSLDDIPENCISSMMMNFDPQEICSLARVNKTFHRASSANFVWESKL 441
Query: 131 KQKWGRVIGSV 141
Q + ++ V
Sbjct: 442 PQNYKFLLNKV 474
>TC207581
Length = 1562
Score = 31.6 bits (70), Expect = 0.33
Identities = 12/40 (30%), Positives = 23/40 (57%)
Frame = +2
Query: 88 DLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWE 127
D+PE + +L L P +C++A + + R+ +D +WE
Sbjct: 461 DIPESCVALVLMYLDPPDICKLARLNRAFRDASSADFIWE 580
>BI426662
Length = 421
Score = 31.6 bits (70), Expect = 0.33
Identities = 17/68 (25%), Positives = 32/68 (47%)
Frame = +3
Query: 89 LPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVIGSVAYREWKW 148
LPE + I+ P C ++ V S R V+D +WER + + +I +
Sbjct: 21 LPEGCIANIVSFTTPPDACVLSLVSSSFRSASVTDFVWERFLPSDYQAIISQSSKPSTLT 200
Query: 149 HVATKKNI 156
+ ++KK++
Sbjct: 201 NYSSKKDL 224
>TC205690 weakly similar to UP|Q6K3E9 (Q6K3E9) F-box family protein-like,
partial (33%)
Length = 1360
Score = 31.2 bits (69), Expect = 0.43
Identities = 14/50 (28%), Positives = 25/50 (50%)
Frame = +2
Query: 89 LPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVI 138
LPE + IL + P+ +C+ + V R SD +W+R + + +I
Sbjct: 215 LPEGCIASILSRTTPADVCRFSVVSKIFRSAAESDAVWKRFLPSDYHSII 364
>AW458378
Length = 363
Score = 31.2 bits (69), Expect = 0.43
Identities = 18/53 (33%), Positives = 26/53 (48%), Gaps = 1/53 (1%)
Frame = +2
Query: 83 NLSVLD-LPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKW 134
++S+L LPE V I L LC + RE C SD +WE ++ +W
Sbjct: 107 SVSILSSLPEDVALKIASLLQVRDLCALGCCSRFWRELCFSDCIWESLVRNRW 265
>AI900244 similar to GP|25082966|gb| spliceosome associated protein - like
{Arabidopsis thaliana}, partial (14%)
Length = 421
Score = 30.8 bits (68), Expect = 0.56
Identities = 18/44 (40%), Positives = 25/44 (55%)
Frame = +1
Query: 152 TKKNIGSLRHGKPRSLIKFMSLYWPFSWMRMKVDAAYSNSANKQ 195
T + + L + + +S IK+MSLY P SWM K+ Y S KQ
Sbjct: 124 TSQGLKGLIYLEDKSQIKWMSLYLPESWMPWKM--FYPRSMKKQ 249
>TC207017
Length = 905
Score = 30.0 bits (66), Expect = 0.96
Identities = 16/69 (23%), Positives = 30/69 (43%)
Frame = +3
Query: 88 DLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVIGSVAYREWK 147
D+PE + + L P +C++A V + +D +WE + + + + + E
Sbjct: 186 DIPESCISSLFMNLDPPDICKLARVNRAFHRASSADFVWESKLPPSY-KFLANKVLGEEN 362
Query: 148 WHVATKKNI 156
TKK I
Sbjct: 363 IATMTKKEI 389
>TC228836 similar to UP|Q6SH18 (Q6SH18) Lipase/esterase, Lip3/BchO family,
partial (9%)
Length = 702
Score = 30.0 bits (66), Expect = 0.96
Identities = 17/49 (34%), Positives = 22/49 (44%)
Frame = +3
Query: 87 LDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWG 135
L LP+ +L I L SL M VC S +HLWE+ +G
Sbjct: 483 LPLPQDILVLIFSFLDMKSLVSMGLVCRSWNIAANDNHLWEKQYVALFG 629
>BE805435 weakly similar to GP|13491750|gb| cysteine protease {Ipomoea
batatas}, partial (22%)
Length = 388
Score = 29.6 bits (65), Expect = 1.2
Identities = 21/70 (30%), Positives = 28/70 (40%)
Frame = -2
Query: 79 SLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVI 138
S QQN L+ L C + P+S AG C SL H ++K GR
Sbjct: 297 SNQQNCLGLNCTSFFLRCCVSL*TPNSP*ISAGACPSL------------HCQRKHGRAW 154
Query: 139 GSVAYREWKW 148
S + W+W
Sbjct: 153 PSTLHAVWQW 124
>CO980830
Length = 834
Score = 29.6 bits (65), Expect = 1.2
Identities = 16/33 (48%), Positives = 19/33 (57%), Gaps = 1/33 (3%)
Frame = -1
Query: 91 ELVLECILEKLPPSSL-CQMAGVCHSLRERCVS 122
E V CILE+ PP SL + G CH +RE S
Sbjct: 306 EKVSTCILEEEPPESLKTRYGG*CHEMRESIYS 208
>BM567646
Length = 242
Score = 28.5 bits (62), Expect = 2.8
Identities = 11/35 (31%), Positives = 21/35 (59%)
Frame = -3
Query: 96 CILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHM 130
C L+KL PS+ ++ +C+ ++ H W+RH+
Sbjct: 120 CYLKKLNPSNKTKIN*LCNDGTKKQKGQHQWQRHV 16
>TC205919 similar to UP|Q94F39 (Q94F39) AT3g29575/MWE13_2, partial (51%)
Length = 1558
Score = 28.5 bits (62), Expect = 2.8
Identities = 15/44 (34%), Positives = 22/44 (49%), Gaps = 8/44 (18%)
Frame = +3
Query: 113 CHSLRERCVSDH--LWER------HMKQKWGRVIGSVAYREWKW 148
C +L E CV DH LW R +++W V S+A+ + W
Sbjct: 837 CTTLGENCVVDHTCLWTRLEWGQ*KREEEWNSVATSLAFSGFHW 968
>TC206024 weakly similar to UP|Q9FLU7 (Q9FLU7) Phloem-specific lectin-like
protein, partial (47%)
Length = 954
Score = 28.5 bits (62), Expect = 2.8
Identities = 13/44 (29%), Positives = 23/44 (51%)
Frame = +3
Query: 84 LSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWE 127
+ + DLPE + IL P +C+++ V + R SD +W+
Sbjct: 21 MELQDLPEGCIAKILSYTTPVDVCRLSLVSKAFRSAAESDTVWD 152
>TC217741 weakly similar to UP|Q84TE4 (Q84TE4) At4g12510, partial (66%)
Length = 1279
Score = 28.1 bits (61), Expect = 3.6
Identities = 16/60 (26%), Positives = 24/60 (39%)
Frame = +2
Query: 89 LPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVIGSVAYREWKW 148
+P + IL+ L L + VC L + LW R +WG + + RE W
Sbjct: 377 IPPELFHHILKFLSSEDLVSCSLVCRFLNYAASDEALWRRLYCMRWGMLPPTRKLRECPW 556
>CO985910
Length = 811
Score = 28.1 bits (61), Expect = 3.6
Identities = 14/45 (31%), Positives = 22/45 (48%)
Frame = +2
Query: 59 MSLVKKTLVSKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPP 103
MSL KT +K + ++ L L V+ L+LEC + + P
Sbjct: 599 MSLYDKTCQAKNTRTQSSIEYLIMYLHVIAFKHLILECTCKAITP 733
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.137 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,257,195
Number of Sequences: 63676
Number of extensions: 240704
Number of successful extensions: 1932
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 1907
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1926
length of query: 236
length of database: 12,639,632
effective HSP length: 94
effective length of query: 142
effective length of database: 6,654,088
effective search space: 944880496
effective search space used: 944880496
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC136507.1


