

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136472.16 - phase: 0
(212 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
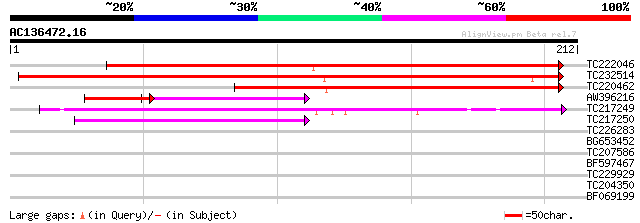
Score E
Sequences producing significant alignments: (bits) Value
TC222046 similar to UP|Q9M2C0 (Q9M2C0) Nodulin-like protein (At3... 236 6e-63
TC232514 UP|NO21_SOYBN (P16313) Nodulin 21 (N-21), complete 202 9e-53
TC220462 similar to UP|Q9M2C3 (Q9M2C3) Nodulin-like protein, par... 160 4e-40
AW396216 homologue to PIR|F96796|F967 nodulin-like protein 6611... 43 1e-09
TC217249 weakly similar to UP|CCC1_YEAST (P47818) CCC1 protein, ... 55 3e-08
TC217250 similar to UP|CCC1_YEAST (P47818) CCC1 protein, partial... 43 9e-05
TC226283 similar to UP|Q6H7H9 (Q6H7H9) Carboxyl-terminal protein... 30 0.81
BG653452 28 2.4
TC207586 similar to GB|BAD10504.1|42409240|AP005544 auxin-regula... 28 3.1
BF597467 28 3.1
TC229929 UP|Q39819 (Q39819) Hsp22.3, complete 28 4.0
TC204350 similar to UP|Q9SP48 (Q9SP48) Homeodomain-leucine zippe... 27 5.2
BF069199 similar to GP|7546729|gb|A MFP1 attachment factor 1 {Gl... 27 5.2
>TC222046 similar to UP|Q9M2C0 (Q9M2C0) Nodulin-like protein (At3g43660),
partial (86%)
Length = 754
Score = 236 bits (602), Expect = 6e-63
Identities = 117/180 (65%), Positives = 145/180 (80%), Gaps = 9/180 (5%)
Frame = +3
Query: 37 YTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGIAGLVAGACSMAIGEFV 96
Y+QRAQWLRAAVLGANDGL+S ASLMMGVGAV KD+ M+L G AGLVAGACSMAIGEFV
Sbjct: 9 YSQRAQWLRAAVLGANDGLVSVASLMMGVGAVKKDISAMLLAGFAGLVAGACSMAIGEFV 188
Query: 97 SVYSQYDIEFAQMKRQ---------GNISQKDKLPNPYYAAFASAIAFAVGAFVPLLGAA 147
SVY+QYDIE Q+KR+ +Q++KLPNP+ AA ASA+AF+VGA VPL+ A
Sbjct: 189 SVYTQYDIEMTQIKREREAYNNRGVNEETQREKLPNPFQAALASALAFSVGALVPLIAAV 368
Query: 148 FVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAMSLTFGLTKLV 207
F++++K+R+GVV VSLAL FG + AVLGK P+ +S LRVL+GGW+AM++TFGLTKL+
Sbjct: 369 FIRNHKIRMGVVAAAVSLALLVFGGVGAVLGKTPVTRSCLRVLVGGWMAMAITFGLTKLI 548
>TC232514 UP|NO21_SOYBN (P16313) Nodulin 21 (N-21), complete
Length = 819
Score = 202 bits (514), Expect = 9e-53
Identities = 110/216 (50%), Positives = 144/216 (65%), Gaps = 12/216 (5%)
Frame = +3
Query: 4 IQHHHAATLARSESKVYNMDVEKKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTASLMM 63
+ H+H + + + + E++ DY QRAQWLRAA+LGANDGL+S ASLMM
Sbjct: 60 VPHNHVGAVLLTIPTIKIDGKQTLATEDHTSIDYLQRAQWLRAAILGANDGLVSVASLMM 239
Query: 64 GVGAVTKDVKTMILTGIAGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQGNIS------- 116
GVGAV +D K M+L G AGLVAGAC MAIGEFV+VY+QY++E QMKR N+S
Sbjct: 240 GVGAVKRDAKAMLLAGFAGLVAGACGMAIGEFVAVYTQYEVEVGQMKRDMNMSVGGERDL 419
Query: 117 ----QKDKLPNPYYAAFASAIAFAVGAFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGL 172
++ LPNP A ASA+ F++GA VPLL AAF+++Y+ R+ VVV + LAL FG
Sbjct: 420 EMEMERRTLPNPLQATLASALCFSIGALVPLLSAAFIENYRTRIIVVVAMSCLALVVFGW 599
Query: 173 LSAVLGKAPLVKSSLRVLIGGW-LAMSLTFGLTKLV 207
+ A LGK P KS +R L+G +AMS+TFG TKL+
Sbjct: 600 VGAKLGKTPK*KSCVRFLLGRCIIAMSITFGSTKLL 707
>TC220462 similar to UP|Q9M2C3 (Q9M2C3) Nodulin-like protein, partial (60%)
Length = 644
Score = 160 bits (405), Expect = 4e-40
Identities = 84/136 (61%), Positives = 102/136 (74%), Gaps = 13/136 (9%)
Frame = +1
Query: 85 AGACSMAIGEFVSVYSQYDIEFAQMKRQGNISQ-------------KDKLPNPYYAAFAS 131
AGACSMAIGEFVSVYSQ DIE AQ KR+ Q K+ LPNP AA AS
Sbjct: 1 AGACSMAIGEFVSVYSQLDIEVAQRKREKERGQRRVRDPEEDTNEEKESLPNPLQAAAAS 180
Query: 132 AIAFAVGAFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLI 191
A+AF+VGA VPLL A+F+++YKVRLGV+V V+ AL FG L AVLGKAP+++S+LRVL
Sbjct: 181 ALAFSVGAMVPLLAASFIREYKVRLGVIVAAVTFALVVFGWLGAVLGKAPVLRSALRVLF 360
Query: 192 GGWLAMSLTFGLTKLV 207
GGW+AM++TFGLTKL+
Sbjct: 361 GGWMAMAITFGLTKLI 408
>AW396216 homologue to PIR|F96796|F967 nodulin-like protein 66117-66707
[imported] - Arabidopsis thaliana, partial (26%)
Length = 388
Score = 42.7 bits (99), Expect(2) = 1e-09
Identities = 18/26 (69%), Positives = 22/26 (84%)
Frame = +3
Query: 29 EEENDEKDYTQRAQWLRAAVLGANDG 54
E + D DY++R+QWLRAAVLGANDG
Sbjct: 138 ESDEDTFDYSKRSQWLRAAVLGANDG 215
Score = 36.2 bits (82), Expect(2) = 1e-09
Identities = 24/63 (38%), Positives = 32/63 (50%)
Frame = +2
Query: 50 GANDGLLSTASLMMGVGAVTKDVKTMILTGIAGLVAGACSMAIGEFVSVYSQYDIEFAQM 109
G+ ++STAS+MMGVG + + G+ G G M IG YSQ D AQ
Sbjct: 200 GSQ*WVVSTASIMMGVGQ*SMTSRP*SYQGLQGWWQGHVXMPIG*VCVCYSQLDHWVAQR 379
Query: 110 KRQ 112
KR+
Sbjct: 380 KRE 388
>TC217249 weakly similar to UP|CCC1_YEAST (P47818) CCC1 protein, partial
(29%)
Length = 1036
Score = 54.7 bits (130), Expect = 3e-08
Identities = 57/249 (22%), Positives = 105/249 (41%), Gaps = 52/249 (20%)
Frame = +3
Query: 12 LARSESKVYNMDVEKKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKD 71
+A +E ++ +++ EKK + + + + +R ++G +DGL +L G+
Sbjct: 21 MAATEHRL-SLEPEKKNLLRHHTEKHFTAGEIVRDIIIGVSDGLTVPFALAAGLSGANAT 197
Query: 72 VKTMILTGIAGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQG------------------ 113
++ GIA + AGA SM +G +++ S+ D ++KR+
Sbjct: 198 SSIVLTAGIAEVAAGAISMGLGGYLAAKSETDHYARELKREQEEIIAVPDTEAAEVAETL 377
Query: 114 --------------NISQKD---------------KLPNP---YYAAFASAIAFAVGAFV 141
N +K+ + P+P ++A AIA+ +G V
Sbjct: 378 AQYGIEAHEYAPVVNALRKNPQAWLDFMMKFELGLEKPDPRRALHSAMTIAIAYILGGVV 557
Query: 142 PLLGAAFVKD--YKVRLGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLRVLIGGWLAMSL 199
PL+ F++ V VVV +V+L +FG+ G P +S+L + G +A +
Sbjct: 558 PLVPYMFIQKAAEAVVFSVVVTLVALLIFGYA-KGHFTGNKPF-RSALETALIGAIASAA 731
Query: 200 TFGLTKLVN 208
FGL K N
Sbjct: 732 AFGLAKAFN 758
>TC217250 similar to UP|CCC1_YEAST (P47818) CCC1 protein, partial (18%)
Length = 422
Score = 43.1 bits (100), Expect = 9e-05
Identities = 23/88 (26%), Positives = 44/88 (49%)
Frame = +1
Query: 25 EKKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGIAGLV 84
EKK + + + + +R ++G +DGL +L G+ ++ GIA +
Sbjct: 13 EKKSLLRHHSEKHFTAGEIVRDIIIGVSDGLTVPFALAAGLSGANATSSIVLTAGIAEVA 192
Query: 85 AGACSMAIGEFVSVYSQYDIEFAQMKRQ 112
AGA SM +G +++ S+ D ++KR+
Sbjct: 193 AGAISMGLGGYLAAKSESDHYARELKRE 276
>TC226283 similar to UP|Q6H7H9 (Q6H7H9) Carboxyl-terminal proteinase-like,
partial (90%)
Length = 1047
Score = 30.0 bits (66), Expect = 0.81
Identities = 14/49 (28%), Positives = 29/49 (58%)
Frame = +1
Query: 25 EKKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVK 73
E++ ++EN++KDY+ + +++ V A T L+ +G +T +VK
Sbjct: 256 EERLQQENEKKDYSSKVYFMKQTVGNA----CGTIGLLHALGNITSEVK 390
>BG653452
Length = 413
Score = 28.5 bits (62), Expect = 2.4
Identities = 14/49 (28%), Positives = 28/49 (56%)
Frame = +2
Query: 25 EKKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVK 73
E++ ++EN++K+Y R +++ V A T L+ +G +T +VK
Sbjct: 194 EERLQQENEKKEYNSRVYFMKQTVGNA----CGTIGLLHALGNLTSEVK 328
>TC207586 similar to GB|BAD10504.1|42409240|AP005544 auxin-regulated
protein-like {Oryza sativa (japonica cultivar-group);} ,
partial (40%)
Length = 1981
Score = 28.1 bits (61), Expect = 3.1
Identities = 16/50 (32%), Positives = 23/50 (46%)
Frame = +3
Query: 9 AATLARSESKVYNMDVEKKGEEENDEKDYTQRAQWLRAAVLGANDGLLST 58
A +LA +E K+Y D + +EKD RAQ + DG L +
Sbjct: 744 ALSLALTEYKIYKTDGLADASTQTEEKDNRSRAQKTCTRGVSTEDGSLES 893
>BF597467
Length = 413
Score = 28.1 bits (61), Expect = 3.1
Identities = 13/33 (39%), Positives = 19/33 (57%)
Frame = +2
Query: 156 LGVVVGVVSLALFGFGLLSAVLGKAPLVKSSLR 188
L +V+G+ L L G L ++LG PLV +R
Sbjct: 191 LCIVIGIKHLKLLLMGALGSILGLPPLVSCKIR 289
>TC229929 UP|Q39819 (Q39819) Hsp22.3, complete
Length = 925
Score = 27.7 bits (60), Expect = 4.0
Identities = 13/31 (41%), Positives = 21/31 (66%)
Frame = +3
Query: 16 ESKVYNMDVEKKGEEENDEKDYTQRAQWLRA 46
ES+V + E+KGEEE +E++ + +W RA
Sbjct: 339 ESRVLRISGERKGEEEEEEEE-VEGEKWHRA 428
>TC204350 similar to UP|Q9SP48 (Q9SP48) Homeodomain-leucine zipper protein
56, partial (44%)
Length = 1197
Score = 27.3 bits (59), Expect = 5.2
Identities = 17/38 (44%), Positives = 22/38 (57%)
Frame = -2
Query: 140 FVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAVL 177
FV L+ A VK + +G ++SL LFGF SAVL
Sbjct: 152 FVVLIEGAKVKLKSIIIGFEDTIISLQLFGFPSGSAVL 39
>BF069199 similar to GP|7546729|gb|A MFP1 attachment factor 1 {Glycine max},
partial (51%)
Length = 259
Score = 27.3 bits (59), Expect = 5.2
Identities = 18/52 (34%), Positives = 28/52 (53%), Gaps = 3/52 (5%)
Frame = -1
Query: 46 AAVLGANDGLLSTASLMMGVGAVTKDVKTMILT---GIAGLVAGACSMAIGE 94
A +LG GL +++G G D + ++L G GLV G C++A+GE
Sbjct: 244 ADILGV--GLEDLDPVIVGGGGADDDGEGLVLDMAGGGGGLVGGHCTVALGE 95
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.135 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,373,360
Number of Sequences: 63676
Number of extensions: 73497
Number of successful extensions: 382
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 377
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 378
length of query: 212
length of database: 12,639,632
effective HSP length: 93
effective length of query: 119
effective length of database: 6,717,764
effective search space: 799413916
effective search space used: 799413916
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC136472.16


