

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136472.12 - phase: 0 /pseudo
(346 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
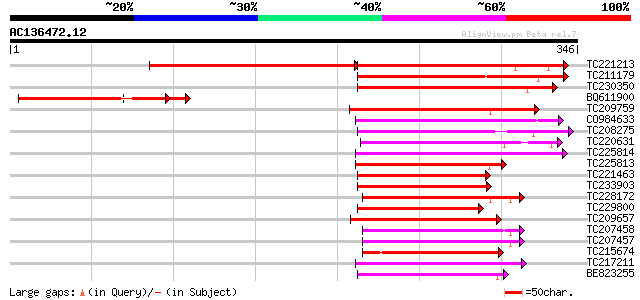
Score E
Sequences producing significant alignments: (bits) Value
TC221213 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein... 184 7e-47
TC211179 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein... 153 1e-37
TC230350 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein... 140 8e-34
BQ611900 100 1e-21
TC209759 90 2e-18
CO984633 90 2e-18
TC208275 similar to UP|Q84TV1 (Q84TV1) Nodulin-like protein, par... 89 2e-18
TC220631 weakly similar to UP|O80638 (O80638) Nodulin-like prote... 89 3e-18
TC225814 homologue to UP|Q9LQZ3 (Q9LQZ3) F10A5.28, partial (23%) 89 4e-18
TC225813 88 5e-18
TC221463 similar to UP|Q8LRB6 (Q8LRB6) Nodulin-like protein, par... 88 6e-18
TC233903 87 8e-18
TC228172 weakly similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like prote... 87 1e-17
TC229800 similar to UP|O24091 (O24091) MtN21 protein, partial (53%) 86 2e-17
TC209657 weakly similar to UP|Q94JU2 (Q94JU2) AT3g28050/MMG15_6,... 83 2e-16
TC207458 similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like protein, par... 78 5e-15
TC207457 weakly similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like prote... 77 1e-14
TC215674 weakly similar to UP|Q9SUD5 (Q9SUD5) Medicago nodulin N... 77 1e-14
TC217211 weakly similar to UP|Q8LA54 (Q8LA54) Nodulin-like prote... 77 1e-14
BE823255 similar to PIR|T00561|T005 nodulin-like protein [import... 75 4e-14
>TC221213 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein;
91922-89607, partial (37%)
Length = 941
Score = 184 bits (466), Expect = 7e-47
Identities = 94/136 (69%), Positives = 111/136 (81%), Gaps = 7/136 (5%)
Frame = +1
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
+ AY GIVVSG MV VISWCVHMRGPLFASVF+PLML+ ALAG TILNEKLHLG +IGA
Sbjct: 532 TAAYTGIVVSGEMVDVISWCVHMRGPLFASVFSPLMLVTEALAGSTILNEKLHLGCVIGA 711
Query: 273 VLIVCGLYAVVWGKSKEMKKKNQLVPSKNPHEFDP---VEIVVRSIEKDKSNLNNNNQ-- 327
VLIVCGLY V+WGKSKEMKKKNQLVP+++PH+ + VEIVVRS ++DKSN N ++
Sbjct: 712 VLIVCGLYVVLWGKSKEMKKKNQLVPAQSPHDNESNTVVEIVVRSAQEDKSNQNKTHEIV 891
Query: 328 --MVKDNEDSQTDEHE 341
+V+DN+DS D HE
Sbjct: 892 AKVVRDNDDS*XDGHE 939
Score = 139 bits (351), Expect = 1e-33
Identities = 71/128 (55%), Positives = 85/128 (65%)
Frame = +3
Query: 86 SMRFIWGIISSKLLLTSPDFNFSNICIRNGQPYTSCNLYHGCFLRYGKIEFENKSREGKD 145
S+R IWGIISSK LLTSP FNF N+CIRN +PYT NLY G LRYGKI+FEN SR+G++
Sbjct: 3 SLRLIWGIISSKFLLTSPGFNFCNVCIRNVEPYTRHNLYLGRLLRYGKIKFENSSRKGQN 182
Query: 146 IRYTNRNWWGNGIDIR*RC*N*DGIIPFKSFTSSKWCCNTHYLHC*YHFGFSMCLG**HL 205
Y N NWWGNG R*RC ++ F+ F S K T C Y GFS+ G* H
Sbjct: 183 SGYINWNWWGNGTYFR*RCAYRIRVLSFEPFASPKRYARTFSYRCSYPLGFSVRPG*WHQ 362
Query: 206 LCTLAYYS 213
LC +A++S
Sbjct: 363 LCIVAHHS 386
>TC211179 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein;
91922-89607, partial (30%)
Length = 771
Score = 153 bits (387), Expect = 1e-37
Identities = 81/134 (60%), Positives = 103/134 (76%), Gaps = 5/134 (3%)
Frame = +2
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
+VAYAGIVVSG MVAVISWCV RGPLF S+F+PLML++VA AG TIL+EKL+LGSIIG+
Sbjct: 347 TVAYAGIVVSGVMVAVISWCVRTRGPLFVSIFSPLMLVVVAFAGSTILDEKLYLGSIIGS 526
Query: 273 VLIVCGLYAVVWGKSKEMKKKNQLVPSKNPHEFDPVEIVVRSIEKDKSN-----LNNNNQ 327
+LI+CGLY V+WGKSKEM KKNQ S++ H+ D +EI+V+ +DKSN L N+
Sbjct: 527 MLIICGLYVVLWGKSKEM-KKNQSGQSESTHKSDTIEIMVKPRVEDKSNNKSNTLINSVN 703
Query: 328 MVKDNEDSQTDEHE 341
+ DN+DS + E
Sbjct: 704 VTGDNKDSWKNGRE 745
Score = 36.2 bits (82), Expect = 0.022
Identities = 26/67 (38%), Positives = 32/67 (46%), Gaps = 2/67 (2%)
Frame = +1
Query: 149 TNRNWWGNGIDIR*RC*N*DGIIPFKSFTSSKWCC--NTHYLHC*YHFGFSMCLG**HLL 206
T+ N WGN DI *R +*D + P K F K C +T Y H S LL
Sbjct: 1 TSWN*WGNATDIH*RPRS*DAVFPCKPFQPPKRPCRTSTRYFWPYDHIWCSRFCRKQRLL 180
Query: 207 CTLAYYS 213
C +AY+S
Sbjct: 181 CHVAYHS 201
>TC230350 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein;
91922-89607, partial (29%)
Length = 1057
Score = 140 bits (353), Expect = 8e-34
Identities = 69/126 (54%), Positives = 92/126 (72%), Gaps = 4/126 (3%)
Frame = +3
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
+VAY+GIV SG +V + +WC+ MRGPLFASVF PLML++VA+A +LNE L++ S++GA
Sbjct: 279 AVAYSGIVASGIVVIITAWCIQMRGPLFASVFNPLMLVLVAIASSLMLNENLYVRSVVGA 458
Query: 273 VLIVCGLYAVVWGKSKEMKKKNQLVPSKNPHEFDPVEIVVRS----IEKDKSNLNNNNQM 328
VLIVCGLY V+WGKSKEMK QLVPS+ E + +E+VV S I+ +K + NN +
Sbjct: 459 VLIVCGLYMVLWGKSKEMKNITQLVPSETIREAEAIEVVVMSISTPIDYEKCDQNNQGER 638
Query: 329 VKDNED 334
D ED
Sbjct: 639 NVDKED 656
>BQ611900
Length = 456
Score = 100 bits (248), Expect = 1e-21
Identities = 52/93 (55%), Positives = 68/93 (72%)
Frame = +2
Query: 6 NLCNVVQGLKPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFI 65
++CNVV LKP LLMV+VQ+A A VNVLYKLA+NDGM+L I+VAYR++FATAFIAP+AFI
Sbjct: 143 DMCNVVHALKPILLMVLVQVANAWVNVLYKLALNDGMNLSIIVAYRYVFATAFIAPLAFI 322
Query: 66 LERLQEEEGKDDLDNTVSVISMRFIWGIISSKL 98
+ER K T +++ F+ G+I L
Sbjct: 323 VER------KTRTKMTWTILFQAFLCGLIGGAL 403
Score = 46.2 bits (108), Expect = 2e-05
Identities = 24/41 (58%), Positives = 29/41 (70%)
Frame = +1
Query: 70 QEEEGKDDLDNTVSVISMRFIWGIISSKLLLTSPDFNFSNI 110
+E E KDDLD+TVS IS+R W I+SKL S FNFSN+
Sbjct: 328 KENEDKDDLDDTVSGISVRLNWWSIASKLKHGSDCFNFSNV 450
>TC209759
Length = 899
Score = 89.7 bits (221), Expect = 2e-18
Identities = 45/123 (36%), Positives = 79/123 (63%), Gaps = 7/123 (5%)
Frame = +1
Query: 208 TLAYYSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLG 267
T+ S YAG + + +++W + +GPL+ SVF+PL+L+++A+A +L+E+L++G
Sbjct: 313 TIRLTSALYAGTISTALAYVLLAWTIERKGPLYVSVFSPLLLVIIAVASWALLHEQLYVG 492
Query: 268 SIIGAVLIVCGLYAVVWGKSKEMKK------KNQLVPSKNPHEFDPV-EIVVRSIEKDKS 320
+ IG++LIV GLY V+WGK+KEM K + ++ + E D V ++ ++ E D S
Sbjct: 493 TAIGSLLIVLGLYFVLWGKNKEMNKIDMVEVEGTVMEAIKESEKDEVKDLELQPYEYDPS 672
Query: 321 NLN 323
N+N
Sbjct: 673 NVN 681
>CO984633
Length = 705
Score = 89.7 bits (221), Expect = 2e-18
Identities = 48/127 (37%), Positives = 72/127 (55%)
Frame = -3
Query: 212 YSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIG 271
++ AYAGIV SG + + GP+ + F PL +++V C IL+E+L LGSIIG
Sbjct: 541 FAPAYAGIVTSGVQYYIQGMVSKIMGPVIVTAFNPLRMIIVTALACIILSEQLFLGSIIG 362
Query: 272 AVLIVCGLYAVVWGKSKEMKKKNQLVPSKNPHEFDPVEIVVRSIEKDKSNLNNNNQMVKD 331
A+++V GLY VVWGK+KE + P++N D ++ V + D N NNN +
Sbjct: 361 AIVVVLGLYLVVWGKAKERRGLMTPSPAENNFPEDQRQLPVTAPRNDSIN-NNNKA*LVH 185
Query: 332 NEDSQTD 338
D ++D
Sbjct: 184 RHDEKSD 164
>TC208275 similar to UP|Q84TV1 (Q84TV1) Nodulin-like protein, partial (34%)
Length = 732
Score = 89.4 bits (220), Expect = 2e-18
Identities = 49/139 (35%), Positives = 79/139 (56%), Gaps = 7/139 (5%)
Frame = +2
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
S YAGI +G ++SW + +GPL+ SVFTPL L++ A+ +L EKL++G+ +G+
Sbjct: 164 SALYAGIFCTGLAYCLMSWTIERKGPLYVSVFTPLQLVLTAILSWALLREKLYVGTAVGS 343
Query: 273 VLIVCGLYAVVWGKSKEMKKKNQLVPSKNPHEFDPVEIVVRSIEKD-------KSNLNNN 325
+LIV GLY+V+WGKS+E+ K + + E D V+ V+ + D SN NN
Sbjct: 344 LLIVLGLYSVLWGKSEEVNKGDGI-------EEDAVKEAVKDSKNDMELQSYVPSNGNNG 502
Query: 326 NQMVKDNEDSQTDEHEQQS 344
+ + ++ D + S
Sbjct: 503 RVIA*RRDAAE*DSSQDAS 559
>TC220631 weakly similar to UP|O80638 (O80638) Nodulin-like protein
(At2g39510), partial (31%)
Length = 797
Score = 89.0 bits (219), Expect = 3e-18
Identities = 52/131 (39%), Positives = 73/131 (55%), Gaps = 8/131 (6%)
Frame = +1
Query: 215 AYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGAVL 274
AY+GIV S V + GP+F + F PL +++V C +L+EKLHLGSIIG V+
Sbjct: 325 AYSGIVTSAIQFYVQGMVIKTTGPVFVTAFNPLRMIIVTALACIVLSEKLHLGSIIGGVV 504
Query: 275 IVCGLYAVVWGKSKEMKKKNQLVPSK---NPHEFDPVEIVVRSIEKDKSNLNNNNQMV-- 329
+V GLY VVWGK+KE K P K + PV + + D +N NN Q+V
Sbjct: 505 VVIGLYLVVWGKAKEQKHLMPPSPEKVTLQRQQQLPVTVPI----SDDANDNNKAQLVII 672
Query: 330 ---KDNEDSQT 337
KD+ +++T
Sbjct: 673 GDRKDDVEART 705
>TC225814 homologue to UP|Q9LQZ3 (Q9LQZ3) F10A5.28, partial (23%)
Length = 736
Score = 88.6 bits (218), Expect = 4e-18
Identities = 48/131 (36%), Positives = 70/131 (52%), Gaps = 2/131 (1%)
Frame = +2
Query: 212 YSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIG 271
+++ YAG+V SG AV WC+ GP+F +V+ P+ +VA+ L E+ +LG IIG
Sbjct: 2 FTILYAGVVASGIAFAVQIWCIDRGGPVFVAVYQPVQTFVVAIMASIALGEEFYLGGIIG 181
Query: 272 AVLIVCGLYAVVWGKSKEMK-KKNQLVPSKNPHE-FDPVEIVVRSIEKDKSNLNNNNQMV 329
AVLIV GLY V+WGKS+E K + QL + H P S+ + + + N
Sbjct: 182 AVLIVAGLYLVLWGKSEERKFAREQLAIASTEHSIIRPASHAKASLAQPLLSSSTENV*T 361
Query: 330 KDNEDSQTDEH 340
+ QT H
Sbjct: 362 RSKFQLQTTHH 394
>TC225813
Length = 1551
Score = 88.2 bits (217), Expect = 5e-18
Identities = 43/94 (45%), Positives = 61/94 (64%), Gaps = 2/94 (2%)
Frame = +1
Query: 212 YSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIG 271
+++ YAG+V SG AV WC+ GP+F +V+ P+ L+VA+ L E+ +LG IIG
Sbjct: 892 FTILYAGVVASGIAFAVQIWCIDRGGPVFVAVYQPVQTLVVAIMASLALGEEFYLGGIIG 1071
Query: 272 AVLIVCGLYAVVWGKSKEMK--KKNQLVPSKNPH 303
AVLIV GLY V+WGKS+E K K++ + S H
Sbjct: 1072AVLIVVGLYFVLWGKSEERKFAKEHAAITSTPEH 1173
Score = 38.9 bits (89), Expect = 0.003
Identities = 18/61 (29%), Positives = 35/61 (56%)
Frame = +1
Query: 8 CNVVQGLKPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILE 67
C++ + ++ M+ +Q +AG +V+ + A+N G+S + YR I A + P A+ LE
Sbjct: 124 CSIPERVQLHAAMLALQFGYAGFHVVSRAALNMGISKLVFPVYRNIIAFLLLLPFAYFLE 303
Query: 68 R 68
+
Sbjct: 304 K 306
>TC221463 similar to UP|Q8LRB6 (Q8LRB6) Nodulin-like protein, partial (37%)
Length = 827
Score = 87.8 bits (216), Expect = 6e-18
Identities = 41/81 (50%), Positives = 59/81 (72%)
Frame = +2
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
+ AYAGI+ SG V + +GP+F + F+PLM+++VA+ G IL EK++LG +IGA
Sbjct: 263 AAAYAGIISSGITYYVQGIVMQKKGPVFVTAFSPLMMIIVAIMGTFILAEKIYLGGVIGA 442
Query: 273 VLIVCGLYAVVWGKSKEMKKK 293
+LIV GLY+V+WGK KE K+K
Sbjct: 443 ILIVMGLYSVLWGKHKENKEK 505
>TC233903
Length = 502
Score = 87.4 bits (215), Expect = 8e-18
Identities = 37/82 (45%), Positives = 58/82 (70%)
Frame = +2
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
+V Y+GIV S +SWCV RGP+F + F+PL+ +M + L+E+LHLGS++G+
Sbjct: 134 TVLYSGIVGSSVCYVGMSWCVKKRGPVFTAAFSPLVQIMSGMIDIPFLHEQLHLGSVVGS 313
Query: 273 VLIVCGLYAVVWGKSKEMKKKN 294
+L++ GLY ++WGKSK+M + N
Sbjct: 314 MLVMIGLYILLWGKSKDMMQNN 379
>TC228172 weakly similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like protein, partial
(36%)
Length = 764
Score = 87.0 bits (214), Expect = 1e-17
Identities = 48/111 (43%), Positives = 67/111 (60%), Gaps = 12/111 (10%)
Frame = +2
Query: 216 YAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGAVLI 275
Y+G++ SG V RGP+F + F+PL +++ A G +L E++HLGSI GA+LI
Sbjct: 164 YSGVICSGMAYYVQGVVTRERGPVFVTSFSPLCMIITAALGSLVLAEQVHLGSIFGAILI 343
Query: 276 VCGLYAVVWGKSKEMKK-------KNQLVPSKNPHE-----FDPVEIVVRS 314
VCGLY VVWGKSK+ K ++Q +P KN + FD +EI V S
Sbjct: 344 VCGLYTVVWGKSKDRKSTTEIEKGESQELPIKNGTKSASDIFDGIEINVPS 496
>TC229800 similar to UP|O24091 (O24091) MtN21 protein, partial (53%)
Length = 1303
Score = 86.3 bits (212), Expect = 2e-17
Identities = 40/77 (51%), Positives = 57/77 (73%)
Frame = +2
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
+ AYAGIV S V + M+GP+FA+ F+PLM+++VA+ G IL E+++LG +IGA
Sbjct: 455 AAAYAGIVTSSISYYVQGLVIKMKGPVFATAFSPLMMIIVAIMGSFILAEQIYLGGVIGA 634
Query: 273 VLIVCGLYAVVWGKSKE 289
+LIV GLY+V+WGK KE
Sbjct: 635 ILIVIGLYSVLWGKHKE 685
>TC209657 weakly similar to UP|Q94JU2 (Q94JU2) AT3g28050/MMG15_6, partial
(31%)
Length = 1014
Score = 83.2 bits (204), Expect = 2e-16
Identities = 37/92 (40%), Positives = 58/92 (62%)
Frame = +1
Query: 209 LAYYSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGS 268
L +V Y+G+ S V +I WC+H GP+F S+F PL +L+ + G L + +LGS
Sbjct: 457 LRLLAVLYSGVFGSAFQVGIICWCLHQTGPVFVSMFKPLGILISVVLGVLFLGDAFYLGS 636
Query: 269 IIGAVLIVCGLYAVVWGKSKEMKKKNQLVPSK 300
+IGA +IV G Y+V+WGK+K+++ + SK
Sbjct: 637 LIGATVIVVGFYSVLWGKAKDIEDAGLSLESK 732
>TC207458 similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like protein, partial (47%)
Length = 1006
Score = 78.2 bits (191), Expect = 5e-15
Identities = 45/112 (40%), Positives = 64/112 (56%), Gaps = 13/112 (11%)
Frame = +2
Query: 216 YAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGAVLI 275
Y+G+V SG V RGP+F + F+PL +++ A G +L E+++LGS+IGA++I
Sbjct: 428 YSGVVCSGMAYYVQGVVTRERGPVFVTSFSPLCMIITAALGSIVLAEQVYLGSVIGAIII 607
Query: 276 VCGLYAVVWGKSKEMKKKNQLVPSKNPHE-------------FDPVEIVVRS 314
V GLY VVWGKSK+ K + S+ HE FD +EI V S
Sbjct: 608 VSGLYTVVWGKSKDKLNKTKEGNSEG-HELPIKDGTKSGSDIFDSIEINVAS 760
>TC207457 weakly similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like protein, partial
(32%)
Length = 757
Score = 77.0 bits (188), Expect = 1e-14
Identities = 42/112 (37%), Positives = 61/112 (53%), Gaps = 13/112 (11%)
Frame = +1
Query: 216 YAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGAVLI 275
Y+G+V SG V RGP+F + F+PL +++ A G +L E++++GS+IGA++I
Sbjct: 133 YSGVVCSGMAYYVQGVVTRERGPVFVTSFSPLCMIITAALGSIVLAEQVYMGSVIGAIII 312
Query: 276 VCGLYAVVWGKSKEMKKKNQLVPSKNPHE-------------FDPVEIVVRS 314
V GLY VVWGKSK+ + HE FD +EI V S
Sbjct: 313 VSGLYTVVWGKSKDKLNNKTNEGNSEGHELPIKDGTKSGSDIFDSIEINVPS 468
>TC215674 weakly similar to UP|Q9SUD5 (Q9SUD5) Medicago nodulin N21-like
protein, partial (28%)
Length = 1577
Score = 77.0 bits (188), Expect = 1e-14
Identities = 34/86 (39%), Positives = 56/86 (64%)
Frame = +3
Query: 216 YAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGAVLI 275
YAGI ++ + + SWC+ RGPL+ ++F PL ++ AL T L E++++GS++GAV +
Sbjct: 963 YAGIGIAVSFF-IQSWCISERGPLYCAMFNPLATVITALISATFLEEEVYVGSLVGAVGV 1139
Query: 276 VCGLYAVVWGKSKEMKKKNQLVPSKN 301
+ GLY V+WGK+KE + P +
Sbjct: 1140 IAGLYVVLWGKAKEFAEIKPEAPQSS 1217
Score = 39.3 bits (90), Expect = 0.003
Identities = 20/55 (36%), Positives = 33/55 (59%)
Frame = +2
Query: 16 PTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILERLQ 70
P ++M+ +QI +A + + + A+ DG+S + V YR AT +API F +R Q
Sbjct: 59 PLIVMIGLQIHYAALAIFTRAALLDGLSTTVFVVYRQGIATLALAPIFFSPKRRQ 223
>TC217211 weakly similar to UP|Q8LA54 (Q8LA54) Nodulin-like protein, partial
(40%)
Length = 909
Score = 76.6 bits (187), Expect = 1e-14
Identities = 40/104 (38%), Positives = 54/104 (51%)
Frame = +2
Query: 212 YSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIG 271
Y Y GIV SG + RGP+F + F PL +++ + G + E+LHLGSIIG
Sbjct: 305 YGPLYTGIVTSGITYYAQGLILQTRGPVFLTAFNPLCMVITSALGSFLFAEQLHLGSIIG 484
Query: 272 AVLIVCGLYAVVWGKSKEMKKKNQLVPSKNPHEFDPVEIVVRSI 315
AV+I GLY+VVWGK K+ P+ E + I I
Sbjct: 485 AVIIALGLYSVVWGKGKDYSNPTPSSPTTKHTETPQLPITSSDI 616
>BE823255 similar to PIR|T00561|T005 nodulin-like protein [imported] -
Arabidopsis thaliana, partial (22%)
Length = 537
Score = 75.1 bits (183), Expect = 4e-14
Identities = 37/97 (38%), Positives = 58/97 (59%), Gaps = 5/97 (5%)
Frame = -2
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
+ Y+GIV SG + + RGP+F + F PL +++VA+ G L E ++LG ++GA
Sbjct: 509 AAVYSGIVCSGMAYYIQGAVMKDRGPVFVTTFNPLCMVIVAIMGSFFLAEIMYLGRVVGA 330
Query: 273 VLIVCGLYAVVWGKSKEMKKKNQL-----VPSKNPHE 304
++I+ GLY VVWGKS + + N + +PSK E
Sbjct: 329 IVIILGLYLVVWGKSNDYESSNSITKKHTLPSKQTVE 219
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.334 0.145 0.470
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,277,569
Number of Sequences: 63676
Number of extensions: 247916
Number of successful extensions: 2356
Number of sequences better than 10.0: 101
Number of HSP's better than 10.0 without gapping: 2338
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2353
length of query: 346
length of database: 12,639,632
effective HSP length: 98
effective length of query: 248
effective length of database: 6,399,384
effective search space: 1587047232
effective search space used: 1587047232
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC136472.12


