

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136451.2 - phase: 0 /pseudo
(355 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
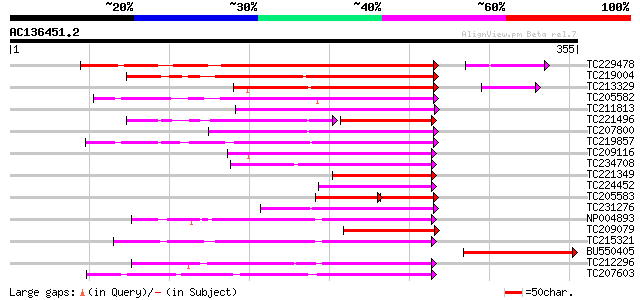
Sequences producing significant alignments: (bits) Value
TC229478 similar to UP|Q94IW5 (Q94IW5) Cytochrome P450-like prot... 193 3e-51
TC219004 homologue to UP|Q9LKH7 (Q9LKH7) Cytochrome P450, partia... 140 7e-34
TC213329 similar to UP|Q9LKH7 (Q9LKH7) Cytochrome P450, partial ... 132 2e-32
TC205582 similar to UP|Q9FH76 (Q9FH76) Cytochrome P450 monooxyge... 109 2e-24
TC211813 similar to UP|Q9FH76 (Q9FH76) Cytochrome P450 monooxyge... 94 9e-20
TC221496 similar to UP|CP85_LYCES (Q43147) Cytochrome P450 85 (... 64 3e-17
TC207800 weakly similar to UP|Q9LVY4 (Q9LVY4) Similarity to cyto... 75 3e-14
TC219857 similar to UP|Q9LVY7 (Q9LVY7) Cytochrome P450-like, par... 75 3e-14
TC209116 similar to UP|Q84ZW1 (Q84ZW1) Ent-kaurenoic acid oxidas... 75 6e-14
TC234708 weakly similar to UP|Q9LN32 (Q9LN32) F18O14.38, partial... 74 8e-14
TC221349 similar to UP|CP85_LYCES (Q43147) Cytochrome P450 85 (... 72 4e-13
TC224452 70 1e-12
TC205583 similar to UP|Q9FH76 (Q9FH76) Cytochrome P450 monooxyge... 45 1e-11
TC231276 weakly similar to GB|BAC56038.1|28071350|AP005103 5-alp... 65 5e-11
NP004893 cytochrome P-450 (CYP93A2) 63 2e-10
TC209079 59 3e-09
TC215321 UP|Q9M6D6 (Q9M6D6) Isoflavone synthase 1, complete 58 6e-09
BU550405 similar to PIR|T04602|T046 cytochrome P450 homolog F23E... 58 7e-09
TC212296 UP|C933_SOYBN (O81973) Cytochrome P450 93A3 (P450 CP5)... 57 1e-08
TC207603 UP|C773_SOYBN (O48928) Cytochrome P450 77A3 , complete 56 2e-08
>TC229478 similar to UP|Q94IW5 (Q94IW5) Cytochrome P450-like protein, partial
(55%)
Length = 1226
Score = 193 bits (491), Expect(2) = 3e-51
Identities = 105/224 (46%), Positives = 142/224 (62%)
Frame = +3
Query: 45 LLGETLDFIASGYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMKLIMDNNAED 104
LL + SG S P+ + LY+SL+AK++M+K+V RI+ +
Sbjct: 150 LLKKHFQEFISGLMSLPIKLPGTK---LYQSLQAKKKMVKLVKRIILAK---------RS 293
Query: 105 DEKVGVVNDVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKF 164
V DVVD LL D E L ++I+ NII+ MIPGE+++P MTLA K+
Sbjct: 294 SGFCKVPKDVVDVLLSDANE--------KLTDDLIADNIIDMMIPGEDSVPLLMTLATKY 449
Query: 165 LTDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRK 224
L++ P AL +L EENM+LKK + + +WTDY+SLPFTQ VI+ETLRM NI+ G+ RK
Sbjct: 450 LSECPAALQQLTEENMKLKKIQDQDGESLSWTDYLSLPFTQTVITETLRMGNIIIGVMRK 629
Query: 225 AVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWE 268
A+KDVEIKG+LIPK W V + SVH+D KNYE PY+F+PWRW+
Sbjct: 630 ALKDVEIKGHLIPKGWCVFVNFRSVHLDDKNYECPYQFNPWRWQ 761
Score = 26.6 bits (57), Expect(2) = 3e-51
Identities = 21/53 (39%), Positives = 28/53 (52%)
Frame = +1
Query: 286 KLELCLVTIALPRLGVDIDCVLG*NFPG*SSLSSFTILLLLIDGRLRGMK*YT 338
K ++ V I+L G D C L +PG SS+T L L DG L+ M+* T
Sbjct: 760 KTKIRAVAISLLLEG-DKGCALALTWPGWKLPSSYTTLSLNSDGMLKKMQ**T 915
>TC219004 homologue to UP|Q9LKH7 (Q9LKH7) Cytochrome P450, partial (95%)
Length = 1575
Score = 140 bits (354), Expect = 7e-34
Identities = 76/195 (38%), Positives = 124/195 (62%)
Frame = +2
Query: 74 KSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSSN 133
+++KA+ ++ + + +V +R K E+ EK+ +D++ ALL +S +
Sbjct: 641 RAIKARTKVAEALALVVRQRRK----EYGENKEKM---SDMLGALL---------ASGDH 772
Query: 134 LMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGY 193
L E I ++ ++ G ET T MTLA+KFLT+ PLAL++L EE+ +++ K++
Sbjct: 773 LSDEEIVDFLLALLVAGYETTSTIMTLAIKFLTETPLALAQLKEEHDQIRA-KSHPGAPL 949
Query: 194 TWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDS 253
WTDY S+ FTQ V++ETLR+ANI+ GI+R+A D+ IKGY IPK W V AS +VH++
Sbjct: 950 EWTDYKSMAFTQCVVNETLRVANIIGGIFRRATTDINIKGYTIPKGWKVFASFRAVHLNP 1129
Query: 254 KNYENPYKFDPWRWE 268
++Y++ F+PWRW+
Sbjct: 1130EHYKDARSFNPWRWQ 1174
Score = 34.7 bits (78), Expect = 0.068
Identities = 17/59 (28%), Positives = 35/59 (58%)
Frame = +2
Query: 10 SIIVLLMSLWFVKKEKNNIKLRNGIKIPKGNSGWPLLGETLDFIASGYSSCPVTFMEKR 68
++I ++L ++++ + R ++P G+ G PL+GETL I++ S P F+++R
Sbjct: 11 AVIFFTVALLYLRR----VFRRRQFRLPPGSHGLPLIGETLQLISAYKSDNPEPFIDER 175
>TC213329 similar to UP|Q9LKH7 (Q9LKH7) Cytochrome P450, partial (44%)
Length = 887
Score = 132 bits (333), Expect(2) = 2e-32
Identities = 63/133 (47%), Positives = 97/133 (72%), Gaps = 5/133 (3%)
Frame = +1
Query: 141 QNIIEFM----IPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDG-YTW 195
+ I++FM + G ET T MTLA+KFLT+ PLAL++L EE+ +++ +K+ CP+ W
Sbjct: 25 EEIVDFMLALLVAGYETTSTIMTLAIKFLTETPLALAQLKEEHDQIRAKKS-CPEAPLEW 201
Query: 196 TDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKN 255
TDY S+ FTQ V++ETLR+ANI+ I+R+A+ D+ IKGY IPK W V+AS +VH++ +
Sbjct: 202 TDYKSMAFTQCVVNETLRVANIIGAIFRRAMTDINIKGYTIPKGWRVVASFRAVHLNPDH 381
Query: 256 YENPYKFDPWRWE 268
+++ F+PWRW+
Sbjct: 382 FKDARTFNPWRWQ 420
Score = 24.3 bits (51), Expect(2) = 2e-32
Identities = 16/37 (43%), Positives = 20/37 (53%)
Frame = +2
Query: 296 LPRLGVDIDCVLG*NFPG*SSLSSFTILLLLIDGRLR 332
+P L CVLG + P SLSSF L +I G L+
Sbjct: 455 IPHLEEGHVCVLGMSLPEWCSLSSFIALSPVIVGFLQ 565
>TC205582 similar to UP|Q9FH76 (Q9FH76) Cytochrome P450 monooxygenase
(AT5g45340/K9E15_12), partial (60%)
Length = 1278
Score = 109 bits (273), Expect = 2e-24
Identities = 65/218 (29%), Positives = 114/218 (51%), Gaps = 2/218 (0%)
Frame = +2
Query: 53 IASGYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVN 112
+ GY+S P+ ++ +K++KA++ + ++V +I+ R + D +
Sbjct: 92 LEQGYNSMPINVPG---TLFHKAMKARKELAQIVAQIISSRRQRKQDFH----------K 232
Query: 113 DVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLAL 172
D++ + + +K S L E I+ N+I + +T + +T +K+L + P L
Sbjct: 233 DLLGSFMDEK---------SGLTDEQIADNVIGVIFAARDTTASVLTWIVKYLGENPSVL 385
Query: 173 SKLMEENMELKKQKTNCPD--GYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVE 230
+ EE + K K + G W D +P T VI ETLR+A+I++ +R+AV+DVE
Sbjct: 386 EAVTEEQECILKSKEERGEDKGLNWEDAKKMPITSRVIQETLRVASILSFTFREAVEDVE 565
Query: 231 IKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWE 268
+GYLIPK W V+ ++H N++ P KFDP R+E
Sbjct: 566 YQGYLIPKGWKVLPLFRNIHHSPDNFKEPEKFDPSRFE 679
>TC211813 similar to UP|Q9FH76 (Q9FH76) Cytochrome P450 monooxygenase
(AT5g45340/K9E15_12), partial (38%)
Length = 889
Score = 94.0 bits (232), Expect = 9e-20
Identities = 48/128 (37%), Positives = 73/128 (56%)
Frame = +1
Query: 142 NIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMSL 201
N+I + +T +A+T LK+L D L + +E +K + G +W D +
Sbjct: 1 NLIGVIFAAHDTTASALTWVLKYLHDNANLLEAVTKEQEGIKNKLAMENRGLSWDDTRQM 180
Query: 202 PFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYK 261
PFT VI ETLR A+I++ +R+AV DVE++GY IPK W V+ S+H + + P K
Sbjct: 181 PFTSRVIQETLRSASILSFTFREAVTDVELEGYTIPKGWKVLPLFRSIHHSADFFPQPEK 360
Query: 262 FDPWRWEI 269
FDP R+E+
Sbjct: 361 FDPSRFEV 384
>TC221496 similar to UP|CP85_LYCES (Q43147) Cytochrome P450 85 (Dwarf
protein) , partial (53%)
Length = 849
Score = 64.3 bits (155), Expect(2) = 3e-17
Identities = 27/60 (45%), Positives = 36/60 (60%)
Frame = +1
Query: 208 ISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRW 267
I ET R+A VNG+ RK D+E+ GYLIPK W + ++ D Y +P F+PWRW
Sbjct: 355 IFETSRLATTVNGVLRKTTHDMELNGYLIPKGWRIYVYTREINYDPFLYHDPLAFNPWRW 534
Score = 41.6 bits (96), Expect(2) = 3e-17
Identities = 33/132 (25%), Positives = 66/132 (50%)
Frame = +3
Query: 74 KSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSSN 133
+ L+A++ ++ ++ +++EER KL + + D++ L+ ++ +
Sbjct: 9 RGLQARKSIISILSQLLEER-KLSQEAHV----------DMLGCLM------NREENRYK 137
Query: 134 LMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGY 193
L E I II M G ET+ T +ALK+L D+P L ++ +E+ ++++K D
Sbjct: 138 LTDEEIIDLIITIMYSGYETVSTTSMMALKYLHDHPKVLEEIRKEHFAIRERK-KPEDPI 314
Query: 194 TWTDYMSLPFTQ 205
D S+ FT+
Sbjct: 315 DGNDLKSMRFTR 350
>TC207800 weakly similar to UP|Q9LVY4 (Q9LVY4) Similarity to cytochrome P450,
partial (77%)
Length = 800
Score = 75.5 bits (184), Expect = 3e-14
Identities = 43/145 (29%), Positives = 72/145 (49%), Gaps = 1/145 (0%)
Frame = +2
Query: 125 LSQSSSSSNLMVEM-ISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELK 183
L S + M EM I NI+ + G + + ++L +K+L P +++E +E+
Sbjct: 20 LVTSDPNGRFMTEMEIIDNILLLLFAGHDNSRSVLSLVMKYLGQLPQVYEHVLKEQLEIS 199
Query: 184 KQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVM 243
+ K W D + ++ NV SE +R++ V+G +R+A+KD Y IPK W +
Sbjct: 200 QGK-EAGQLLQWEDVQKMKYSWNVASEVMRLSPPVSGAYREAIKDFTYADYNIPKGWKLH 376
Query: 244 ASLTSVHMDSKNYENPYKFDPWRWE 268
+ S H D + NP FD R+E
Sbjct: 377 WNTGSSHKDPTLFSNPETFDASRFE 451
>TC219857 similar to UP|Q9LVY7 (Q9LVY7) Cytochrome P450-like, partial (36%)
Length = 1146
Score = 75.5 bits (184), Expect = 3e-14
Identities = 56/222 (25%), Positives = 104/222 (46%), Gaps = 1/222 (0%)
Frame = +2
Query: 48 ETLDFIASGYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEK 107
E L+ + +G S P+ F ++ + +KA + + + + RIV++R K+ + N +
Sbjct: 2 EPLNQVNAGIISMPINFPG---TVFNRGIKASKFIRRELLRIVKQR-KVELANGMSTPTQ 169
Query: 108 VGVVNDVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTD 167
D++ +L E Q + +++ I+ +I ET T T +K+L +
Sbjct: 170 -----DILSHMLIYCDENGQYLAEHDIV-----NKILGLLIGSHETTSTVCTFVVKYLAE 319
Query: 168 YPLAL-SKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAV 226
P + + +E M + K K + W D + ++ NV E +R+ G +R+A+
Sbjct: 320 LPQNIYENVYQEQMAIAKSKAP-GELLNWDDIQKMKYSWNVACEVIRLNPPAQGAFREAI 496
Query: 227 KDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWE 268
D G+ IPK W + S S H + + + P KFDP R+E
Sbjct: 497 NDFIFDGFSIPKGWKLYWSANSTHKNPEYFPEPEKFDPSRFE 622
>TC209116 similar to UP|Q84ZW1 (Q84ZW1) Ent-kaurenoic acid oxidase, partial
(42%)
Length = 1139
Score = 74.7 bits (182), Expect = 6e-14
Identities = 42/135 (31%), Positives = 69/135 (51%), Gaps = 4/135 (2%)
Frame = +1
Query: 137 EMISQNIIEFMI----PGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDG 192
++ ++II+ M+ G E+ A FL +P L K E E+ +++ + G
Sbjct: 37 KLSDEDIIDIMLMYLNAGHESSGHITMWATFFLQKHPEYLQKAKAEQEEIIRRRPSTQKG 216
Query: 193 YTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMD 252
T + + F VI ETLR+ ++R+A DV I GY +PK W V+ SVH+D
Sbjct: 217 LTLKEVREMDFLYKVIDETLRVITFSLVVFREAKTDVNINGYTVPKGWKVLVWFRSVHLD 396
Query: 253 SKNYENPYKFDPWRW 267
+ + +P +F+P RW
Sbjct: 397 PEIFPDPKEFNPNRW 441
>TC234708 weakly similar to UP|Q9LN32 (Q9LN32) F18O14.38, partial (43%)
Length = 540
Score = 74.3 bits (181), Expect = 8e-14
Identities = 40/130 (30%), Positives = 72/130 (54%), Gaps = 1/130 (0%)
Frame = +2
Query: 139 ISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGYTWTDY 198
I NI+ +I G++T+ AMT +KF+ + + LM+E +LK +K + Y +
Sbjct: 98 IKDNILTMIIAGQDTIANAMTWMIKFVDENRQVFNTLMKE--QLKIEKNGSRNSYLTLEA 271
Query: 199 MS-LPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYE 257
++ +P+ V+ E LR A++V + R A++D I+G+ I K W + S+H D +
Sbjct: 272 LNEMPYASKVVKEALRKASVVQWLPRVALEDCVIEGFKIKKGWNINIDARSIHHDPTVHX 451
Query: 258 NPYKFDPWRW 267
+P F+P R+
Sbjct: 452 DPMVFNPSRF 481
>TC221349 similar to UP|CP85_LYCES (Q43147) Cytochrome P450 85 (Dwarf
protein) , partial (29%)
Length = 534
Score = 72.0 bits (175), Expect = 4e-13
Identities = 31/65 (47%), Positives = 42/65 (63%)
Frame = +1
Query: 203 FTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKF 262
FT+ VI ET R+A IVNG+ RK +D+E+ GYLIPK W + ++ D Y +P F
Sbjct: 10 FTRAVIFETSRLATIVNGVLRKTTQDMELNGYLIPKGWRIYVYTREINYDPFLYPDPLTF 189
Query: 263 DPWRW 267
+PWRW
Sbjct: 190 NPWRW 204
>TC224452
Length = 810
Score = 70.1 bits (170), Expect = 1e-12
Identities = 31/74 (41%), Positives = 45/74 (59%)
Frame = +3
Query: 194 TWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDS 253
TW DY + FTQ VI ETLR+ + + R+A +DV+ + ++IPK V+ L+ VH+D
Sbjct: 27 TWQDYXXMSFTQCVIDETLRLXGMXIWLMREAKEDVQYQDFVIPKGCFVVPFLSXVHLDE 206
Query: 254 KNYENPYKFDPWRW 267
Y F+PWRW
Sbjct: 207 NVYSGALNFNPWRW 248
>TC205583 similar to UP|Q9FH76 (Q9FH76) Cytochrome P450 monooxygenase
(AT5g45340/K9E15_12), partial (31%)
Length = 985
Score = 44.7 bits (104), Expect(2) = 1e-11
Identities = 20/42 (47%), Positives = 29/42 (68%)
Frame = +2
Query: 192 GYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKG 233
G W D +P T VI ETLR+A+I++ +R+AV+DVE +G
Sbjct: 26 GLNWEDAKKMPITSRVIQETLRVASILSFTFREAVEDVEYQG 151
Score = 42.4 bits (98), Expect(2) = 1e-11
Identities = 17/36 (47%), Positives = 23/36 (63%)
Frame = +1
Query: 233 GYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWE 268
GYLIPK W V+ ++H N++ P KFDP R+E
Sbjct: 250 GYLIPKGWKVLPLFRNIHHSPDNFKEPEKFDPSRFE 357
>TC231276 weakly similar to GB|BAC56038.1|28071350|AP005103
5-alpha-taxadienol-10-beta-hydroxylase-like protein
{Oryza sativa (japonica cultivar-group);} , partial
(28%)
Length = 646
Score = 65.1 bits (157), Expect = 5e-11
Identities = 34/111 (30%), Positives = 53/111 (47%)
Frame = +1
Query: 158 MTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANI 217
+T ++ + P + +++ E+ K K + TW D + +T V ET+RM
Sbjct: 22 ITFIIRXXANEPAIYAAVLQXXEEIAKGKLS-GXXLTWEDLSKMKYTWRVAMETIRMFPP 198
Query: 218 VNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWE 268
+ G +RK D+E GY IPK W + HMD + P K DP R+E
Sbjct: 199 IFGGFRKXATDIEYDGYFIPKGWQIFWVTAMTHMDENIFPEPSKIDPSRFE 351
>NP004893 cytochrome P-450 (CYP93A2)
Length = 1509
Score = 62.8 bits (151), Expect = 2e-10
Identities = 51/194 (26%), Positives = 94/194 (48%), Gaps = 3/194 (1%)
Frame = +1
Query: 77 KAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVN---DVVDALLRDKGELSQSSSSSN 133
K + R ++ RI+++R + +++++G D++D LL D GE SS
Sbjct: 694 KTRIRFDAVLDRIIKQR-----EEERRNNKEIGGTRQFKDILDVLL-DIGE--DDSSEIK 849
Query: 134 LMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGY 193
L E I I++ + G +T M A+ L + P L K +E + +
Sbjct: 850 LTKENIKAFIMDIFVAGTDTSAATMEWAMAELINNPCVLEKARQEIDAVVGNSRIIEE-- 1023
Query: 194 TWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDS 253
+D ++LP+ Q ++ ETLR+ I R++ K V + GY IP + ++ ++ D
Sbjct: 1024--SDIVNLPYLQAIVRETLRIHPGGPLIVRESSKSVVVCGYEIPAKTRLFVNVWAIGRDP 1197
Query: 254 KNYENPYKFDPWRW 267
++ENP++F P R+
Sbjct: 1198NHWENPFEFRPERF 1239
>TC209079
Length = 659
Score = 58.9 bits (141), Expect = 3e-09
Identities = 26/60 (43%), Positives = 41/60 (68%)
Frame = +2
Query: 210 ETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWEI 269
E LRMA+I++ +R+A+ DVE KG+LIPK W M ++H + + + P KF+P R+E+
Sbjct: 5 EGLRMASIISFPFREAIADVEYKGFLIPKGWKAMPLFRNIHHNPEFFPEPQKFNPLRFEV 184
>TC215321 UP|Q9M6D6 (Q9M6D6) Isoflavone synthase 1, complete
Length = 1888
Score = 58.2 bits (139), Expect = 6e-09
Identities = 53/203 (26%), Positives = 88/203 (43%), Gaps = 1/203 (0%)
Frame = +1
Query: 66 EKR-KSILYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGE 124
EKR IL K ER++K IV R + + E GV D + D+
Sbjct: 769 EKRIDDILNKFDPVVERVIKKRREIVRRRK----NGEVVEGEASGVFLDTLLEFAEDE-- 930
Query: 125 LSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKK 184
+ + E I +++F G ++ A AL L + P L K EE +
Sbjct: 931 ----TMEIKITKEQIKGLVVDFFSAGTDSTAVATEWALAELINNPRVLQKAREEVYSVVG 1098
Query: 185 QKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMA 244
+ + D +LP+ + ++ ET RM + + RK ++ EI GY+IP+ V+
Sbjct: 1099KDRLVDE----VDTQNLPYIRAIVKETFRMHPPLPVVKRKCTEECEINGYVIPEGALVLF 1266
Query: 245 SLTSVHMDSKNYENPYKFDPWRW 267
++ V D K ++ P +F P R+
Sbjct: 1267NVWQVGRDPKYWDRPSEFRPERF 1335
>BU550405 similar to PIR|T04602|T046 cytochrome P450 homolog F23E13.220 -
Arabidopsis thaliana, partial (23%)
Length = 612
Score = 57.8 bits (138), Expect = 7e-09
Identities = 41/71 (57%), Positives = 44/71 (61%)
Frame = -3
Query: 285 RKLELCLVTIALPRLGVDIDCVLG*NFPG*SSLSSFTILLLLIDGRLRGMK*YTFQLLR* 344
R L L +TIAL L VD D L * G*SSL SF ILL L DG LR MK +TFQ R*
Sbjct: 601 RILGLGXITIALHHLVVDRDYALA*XSQG*SSLFSFIILLQLTDGLLRRMKLFTFQQ*R* 422
Query: 345 RRSSQLECNPL 355
R +SQL PL
Sbjct: 421 RGNSQLV*QPL 389
>TC212296 UP|C933_SOYBN (O81973) Cytochrome P450 93A3 (P450 CP5) , complete
Length = 1646
Score = 57.4 bits (137), Expect = 1e-08
Identities = 48/194 (24%), Positives = 92/194 (46%), Gaps = 3/194 (1%)
Frame = +1
Query: 77 KAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGV--VNDVVDALLRDKGELSQSSSSS-N 133
K E++ +++ +K + +E VG D++D L ++S+ SS
Sbjct: 718 KRLEKIRDCFDTVLDRIIKQREEERRNKNETVGKREFKDMLDVLF----DISEDESSEIK 885
Query: 134 LMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGY 193
L E I I++ +I G +T M A+ L + P L K +E M+ K+ +
Sbjct: 886 LNKENIKAFILDILIAGTDTSAVTMEWAMAELINNPGVLEKARQE-MDAVVGKSRIVEE- 1059
Query: 194 TWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDS 253
+D +LP+ Q ++ ETLR+ ++R++ + + GY IP + ++ ++ D
Sbjct: 1060--SDIANLPYLQGIVRETLRLHPAGPLLFRESSRRAVVCGYDIPAKTRLFVNVWAIGRDP 1233
Query: 254 KNYENPYKFDPWRW 267
++ENP +F P R+
Sbjct: 1234NHWENPLEFRPERF 1275
>TC207603 UP|C773_SOYBN (O48928) Cytochrome P450 77A3 , complete
Length = 1929
Score = 56.2 bits (134), Expect = 2e-08
Identities = 57/220 (25%), Positives = 93/220 (41%), Gaps = 1/220 (0%)
Frame = +2
Query: 49 TLDFIASGYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKV 108
TLD Y F K++ K+L+ + ++ + I+E+R + I + ++
Sbjct: 794 TLDPRIDDYLPILSPFFSKQRK---KALEVRREQVEFLVPIIEQRRRAIQNPGSDH---T 955
Query: 109 GVVNDVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDY 168
+D L K E +S+ S +V + S EF+ G +T TA+ + L
Sbjct: 956 ATTFSYLDTLFDLKVEGKKSAPSDAELVSLCS----EFLNGGTDTTATAVEWGIAQLIAN 1123
Query: 169 PLALSKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKD 228
P +KL EE +K D +P+ V+ E LR + + AV +
Sbjct: 1124PNVQTKLYEEIKRTVGEKK-----VDEKDVEKMPYLHAVVKELLRKHPPTHFVLTHAVTE 1288
Query: 229 -VEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRW 267
+ GY IP D V ++ D KN+ NP KFDP R+
Sbjct: 1289PTTLGGYDIPIDANVEVYTPAIAEDPKNWLNPEKFDPERF 1408
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.328 0.142 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,585,306
Number of Sequences: 63676
Number of extensions: 206574
Number of successful extensions: 1306
Number of sequences better than 10.0: 108
Number of HSP's better than 10.0 without gapping: 1292
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1300
length of query: 355
length of database: 12,639,632
effective HSP length: 98
effective length of query: 257
effective length of database: 6,399,384
effective search space: 1644641688
effective search space used: 1644641688
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC136451.2


