

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
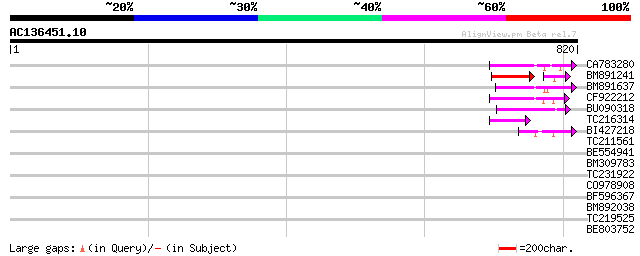
Query= AC136451.10 - phase: 0 /pseudo
(820 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polym... 97 3e-20
BM891241 82 4e-19
BM891637 90 5e-18
CF922212 77 2e-14
BU090318 weakly similar to GP|9743345|gb| F21J9.30 {Arabidopsis ... 66 7e-11
TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {A... 59 9e-09
BI427218 53 6e-07
TC211561 weakly similar to UP|O22675 (O22675) Reverse transcript... 42 0.001
BE554941 42 0.001
BM309783 35 0.003
TC231922 33 0.66
CO978908 32 1.1
BF596367 31 2.5
BM892038 30 4.3
TC219525 similar to UP|Q9LV64 (Q9LV64) Gb|AAD28645.1, partial (27%) 29 9.6
BE803752 29 9.6
>CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polymerase
homolog T24M8.8 - Arabidopsis thaliana, partial (4%)
Length = 456
Score = 97.1 bits (240), Expect = 3e-20
Identities = 51/134 (38%), Positives = 75/134 (55%), Gaps = 8/134 (5%)
Frame = +2
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRV 753
F GG D +KI WV+W VCL K GGLG++ +R FN+ LLGKW W + ++ W +V
Sbjct: 62 FLWGGEADSRKIAWVNWKTVCLPKAKGGLGIKDLRTFNTTLLGKWRWDLFYIQQEPWAKV 241
Query: 754 LVARYGEEGGRLLDGGRTA---SAWWRAIATLHSEAWFGTNVKR----SVGNGENIYFWS 806
L ++YG G R L+ G + SAWW+ + + +KR VG G+ I FW
Sbjct: 242 LQSKYG--GWRALEEGSSGSKDSAWWKDLIKT-QQLQRNIPLKRETIWKVGGGDRIKFWE 412
Query: 807 DVWVG-GVSFKERF 819
D+W +S +++F
Sbjct: 413 DLWTNTDLSLRDKF 454
>BM891241
Length = 407
Score = 82.0 bits (201), Expect(2) = 4e-19
Identities = 33/63 (52%), Positives = 45/63 (71%)
Frame = -2
Query: 697 GGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRVLVA 756
GG+ DHKKI WV W VCL K GGLG++ I +FN ALLGKW W + +++ LW R++ +
Sbjct: 406 GGDHDHKKIPWVKWEVVCLPKTKGGLGIKDISKFNIALLGKWIWALASDQQQLWARIINS 227
Query: 757 RYG 759
+YG
Sbjct: 226 KYG 218
Score = 31.6 bits (70), Expect(2) = 4e-19
Identities = 14/42 (33%), Positives = 22/42 (52%), Gaps = 3/42 (7%)
Frame = -3
Query: 773 SAWWRAIATLHSEA---WFGTNVKRSVGNGENIYFWSDVWVG 811
S WWR + + ++ F + VG G+ I FW+D W+G
Sbjct: 174 SYWWRDLRKFYHQSDHSIFHQYMSWKVGCGDKINFWTDKWLG 49
>BM891637
Length = 427
Score = 89.7 bits (221), Expect = 5e-18
Identities = 49/124 (39%), Positives = 66/124 (52%), Gaps = 7/124 (5%)
Frame = -1
Query: 703 KKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRVLVARYGEEG 762
KKI W+ W C S +VGGLG++ I+ N+ALL KW W++ + LW R+L+++Y +G
Sbjct: 418 KKIAWISWRQCCASGDVGGLGIKDIKILNNALLIKWKWLMFHQPHQLWNRILISKY--KG 245
Query: 763 GRLLDGGRTA---SAWW---RAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWV-GGVSF 815
R LD G S WW RAI S VG G+ I FW D WV G
Sbjct: 244 WRGLDQGPQKYYFSPWWADLRAINQHQSMIAASNQFCWKVGRGDQILFWEDSWVDDGTPL 65
Query: 816 KERF 819
K++F
Sbjct: 64 KDQF 53
>CF922212
Length = 445
Score = 77.4 bits (189), Expect = 2e-14
Identities = 50/124 (40%), Positives = 66/124 (52%), Gaps = 8/124 (6%)
Frame = -2
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRV 753
F GG KKI WV+W+ VCL KE GLG R +R+FN ALLGK W + + L RV
Sbjct: 396 FLWGGGLGQKKIAWVNWDSVCLPKEKEGLGNRDLRKFNYALLGKRRWNLFHHQGELGARV 217
Query: 754 LVARYGEEGGRLLDGGR---TASAWWRAIATL-HSE---AWFGTNVKRS-VGNGENIYFW 805
L ++Y + R LD R + S WW+ I+ + HS +WF K +G + FW
Sbjct: 216 LDSKY--KRWRNLDEERRVKSESFWWQEISFITHSTEDGSWFEKRTKTGRLGCRAKVRFW 43
Query: 806 SDVW 809
D W
Sbjct: 42 EDGW 31
>BU090318 weakly similar to GP|9743345|gb| F21J9.30 {Arabidopsis thaliana},
partial (1%)
Length = 422
Score = 65.9 bits (159), Expect = 7e-11
Identities = 34/109 (31%), Positives = 59/109 (53%), Gaps = 2/109 (1%)
Frame = +3
Query: 704 KITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRVLVARY--GEE 761
K+ W W +C K+ GGLG+R + N +LL K + + + L RV+ ++Y G++
Sbjct: 3 KVHWTSWKALCSPKKSGGLGLRLLSHMNKSLLMKST*NMCTKPNDL*VRVVRSKYK*GKD 182
Query: 762 GGRLLDGGRTASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWV 810
+D +T + +W+ I + N+ R +GNG +++FW DVWV
Sbjct: 183 ILP*IDPKKTGTNFWQGIVHCWEKITHD-NIARKIGNGRSVHFWKDVWV 326
>TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Arabidopsis
thaliana;} , partial (18%)
Length = 759
Score = 58.9 bits (141), Expect = 9e-09
Identities = 24/60 (40%), Positives = 33/60 (55%)
Frame = +2
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRV 753
F GG H +I WV W +C K GGLG++ + FN+AL G+W W + LW R+
Sbjct: 119 FLWGGTQHHNRIPWVKWADICNPKIDGGLGIKDLSNFNAALRGRWIWGLASNHNQLWARL 298
>BI427218
Length = 421
Score = 52.8 bits (125), Expect = 6e-07
Identities = 33/94 (35%), Positives = 43/94 (45%), Gaps = 10/94 (10%)
Frame = +1
Query: 736 GKWSWMVLVEKESLWYRVLVARYG-----EEGGRLLDGGRTASAWWRAIATLHSE----A 786
GKW W K LW R+L ++YG EE R G ASAWW+ I + +
Sbjct: 1 GKWKWSFFYNKGELWARILDSKYGDWRHLEEPTR----GNRASAWWKDINIIDPSGADGS 168
Query: 787 WFGTNVKRSVGNGENIYFWSDVW-VGGVSFKERF 819
F +K V GE FW D W V+FK ++
Sbjct: 169 LFDKGIKWKVACGEKTKFWEDGWKEDRVTFKLKY 270
>TC211561 weakly similar to UP|O22675 (O22675) Reverse transcriptase
(Fragment) , partial (59%)
Length = 1077
Score = 42.0 bits (97), Expect = 0.001
Identities = 21/49 (42%), Positives = 26/49 (52%)
Frame = +1
Query: 674 FPSSRLPQVSSPLLNLF*IIFFLGGNDDHKKITWVDWNYVCLSKEVGGL 722
F R+PQ L * F GG D KKI+W+ W +CL KE GG+
Sbjct: 763 FSFFRVPQSVVDKLVKI*RTFLWGGGPDQKKISWIRWEKLCLPKERGGI 909
Score = 40.0 bits (92), Expect = 0.004
Identities = 35/104 (33%), Positives = 51/104 (48%)
Frame = +3
Query: 425 K*GCG*CQEM*KGTSSFQSGFRKSL*FHRLDLLR*GNAKDGVLSFMAEMDKRMYRHDNSI 484
K*G * QE + FQSG K + F ++ + ++G S M +MD R+ +
Sbjct: 21 K*GDR*GQEEQQIMPRFQSGL*KGIRFGFMEFSDVYDVENGFQSEMDKMD*RVCQICIYF 200
Query: 485 GFGQWVPNRRVPFEEGFEARRPAFPFPLSLGCRRFQYFDEVNVG 528
F +W P+ VP EG ARRP F + R ++FDE + G
Sbjct: 201 CFSEW*PDS*VPTAEGS*ARRPIDTFFIQYCG*RSKWFDEESSG 332
>BE554941
Length = 383
Score = 41.6 bits (96), Expect = 0.001
Identities = 23/87 (26%), Positives = 38/87 (43%), Gaps = 4/87 (4%)
Frame = +1
Query: 723 GVRRIREFNSALLGKWSWMVLVEKESLWYRVLVARYGEEGGRLLDGGRTASAWWRAIATL 782
G++ + FN +LL K LVE +++W +L YG S WWR I +
Sbjct: 7 GIKNLSIFNISLLVKRR*HFLVETKAIWAPILKFHYGSLDLSNTGSASRTSQWWRDILRI 186
Query: 783 ----HSEAWFGTNVKRSVGNGENIYFW 805
+ W + R +G+ ++ FW
Sbjct: 187 GNCQYQPGWLTSTFCRKIGDRKDTLFW 267
>BM309783
Length = 433
Score = 35.0 bits (79), Expect(2) = 0.003
Identities = 21/64 (32%), Positives = 29/64 (44%)
Frame = +2
Query: 697 GGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRVLVA 756
G ND +K V W + + GG+G+ + N A L K W + SLW VL
Sbjct: 8 GDNDGGRKHHVVGWKAMIMP*HSGGVGMCDLTIMNQACLMKIGWAFRMGDGSLWADVLRG 187
Query: 757 RYGE 760
YG+
Sbjct: 188 *YGQ 199
Score = 24.6 bits (52), Expect(2) = 0.003
Identities = 12/30 (40%), Positives = 17/30 (56%), Gaps = 1/30 (3%)
Frame = +3
Query: 791 NVKR-SVGNGENIYFWSDVWVGGVSFKERF 819
NV+R V NG +I W D WV ++ + F
Sbjct: 285 NVERWIVNNGRSIRAW*DKWVNDITILKAF 374
>TC231922
Length = 803
Score = 32.7 bits (73), Expect = 0.66
Identities = 15/52 (28%), Positives = 30/52 (56%), Gaps = 4/52 (7%)
Frame = -3
Query: 773 SAWWRAIATLHSEAWFGT---NVKRSVGNGENIYFWSDVWVG-GVSFKERFS 820
S WW+ + L+++ F + N+ VG G+ I FW D W+ G + +++++
Sbjct: 777 SHWWKDLRRLYNQPDFHSIHQNMVWRVGCGDKIKFWQDSWLSEGCNLQQKYN 622
>CO978908
Length = 782
Score = 32.0 bits (71), Expect = 1.1
Identities = 14/43 (32%), Positives = 22/43 (50%), Gaps = 3/43 (6%)
Frame = +1
Query: 738 WSWMVLVEKESLWYRVLVARYGEEGGRLLDGGRT---ASAWWR 777
W W + +++ LW R++ ++YG G GR S WWR
Sbjct: 604 WIWALASDQQQLWARIINSKYG--GWEEFQSGRDKRGCSHWWR 726
>BF596367
Length = 419
Score = 30.8 bits (68), Expect = 2.5
Identities = 14/34 (41%), Positives = 22/34 (64%)
Frame = +1
Query: 725 RRIREFNSALLGKWSWMVLVEKESLWYRVLVARY 758
RRI+ F+ A+L K W ++ K++L VL A+Y
Sbjct: 58 RRIKAFHKAMLAKQMWRLIDNKDALVT*VLTAKY 159
>BM892038
Length = 422
Score = 30.0 bits (66), Expect = 4.3
Identities = 27/89 (30%), Positives = 37/89 (41%), Gaps = 5/89 (5%)
Frame = -2
Query: 728 REFNSALLGKWSWMVLVEKESLWYRVLVARYGEEGGRLLDGGRTASAWWRAIATLHSEA- 786
+E N+ +W V KE + V+ EEGG L+ G + W+ A + E
Sbjct: 334 QETNNTWDSRWWRKVFRRKERIHGSVI-----EEGGVLVGDGVG*GSDWKKKAKKNKEKE 170
Query: 787 ---W-FGTNVKRSVGNGENIYFWSDVWVG 811
W FG GNG+ FW WVG
Sbjct: 169 LWKW*FGGKKMHWRGNGQLGNFWFPDWVG 83
>TC219525 similar to UP|Q9LV64 (Q9LV64) Gb|AAD28645.1, partial (27%)
Length = 770
Score = 28.9 bits (63), Expect = 9.6
Identities = 13/38 (34%), Positives = 21/38 (55%)
Frame = +1
Query: 470 MAEMDKRMYRHDNSIGFGQWVPNRRVPFEEGFEARRPA 507
+A M++ Y+ + FG W N +PF + FE+ R A
Sbjct: 175 VAAMEEYYYQRHHVPAFGSWDWNDNLPFTQCFESARQA 288
>BE803752
Length = 346
Score = 28.9 bits (63), Expect = 9.6
Identities = 19/59 (32%), Positives = 30/59 (50%)
Frame = -2
Query: 731 NSALLGKWSWMVLVEKESLWYRVLVARYGEEGGRLLDGGRTASAWWRAIATLHSEAWFG 789
+S L G +M +++E L +L+ R+ R L T +AWWRA+ S W+G
Sbjct: 306 SS*LQGSK*YMRRMQRELL---LLLVRHLLSRRRRLSRK*TMAAWWRAVVLQGSRGWWG 139
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.361 0.164 0.647
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 42,645,118
Number of Sequences: 63676
Number of extensions: 725185
Number of successful extensions: 6968
Number of sequences better than 10.0: 33
Number of HSP's better than 10.0 without gapping: 3707
Number of HSP's successfully gapped in prelim test: 239
Number of HSP's that attempted gapping in prelim test: 3125
Number of HSP's gapped (non-prelim): 4167
length of query: 820
length of database: 12,639,632
effective HSP length: 105
effective length of query: 715
effective length of database: 5,953,652
effective search space: 4256861180
effective search space used: 4256861180
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC136451.10


